Multimodal biosensing platforms for sensor-computing integration towards intelligent healthcare
Abstract
Integrated multimodal biosensing platforms are transforming the landscape of bioanalytical technologies by enabling real-time, high-resolution, and multifunctional detection of physiological and environmental biomarkers. This review summarizes the evolution from traditional single-modal biosensors to advanced multimodal systems that unify diverse sensing modalities with computation and storage functionalities. The introduction on the transduction mechanisms of different biosensors and the representative biomarkers was first provided, highlighting the advantages of multimodal sensing with enhanced sensitivity, specificity, and robustness. The advances in fabrication techniques were then discussed, with particular emphasis on printable strategies that facilitate heterogeneous material integration and micro/nanoscale patterning. Moreover, artificial intelligence-driven data processing for on-device decision-making was discussed. Representative applications were then presented in healthcare monitoring, environment detection, and food safety tracking. Finally, current challenges related to material compatibility, data heterogeneity, device scalability, and regulatory barriers were proposed with possible strategies toward fully autonomous and intelligent biosensing systems.
Keywords
INTRODUCTION
Recent advances in biosensing technology have significantly expanded the possibilities for real-time health monitoring, disease diagnosis, and environmental analysis. Among these developments, integrated biosensing platforms capable of detecting multiple biomarkers simultaneously have drawn considerable attention due to their ability to provide comprehensive, high-fidelity data for complex biological systems[1-5]. Traditional biosensors typically rely on single-mode detection strategies, which, although effective in controlled settings, often fail to capture the multifactorial nature of physiological and pathological processes. In contrast, multimodal biosensors, which integrate diverse sensing mechanisms such as electrochemical, microwave, optical, and piezoelectric transduction, offer more robust and nuanced data acquisition, enabling cross-validation and compensation among modalities[6-8].
The convergence of multimodal biosensing with advanced materials, microfabrication techniques, and artificial intelligence (AI) has paved the way for next-generation platforms that go beyond simple detection[9,10]. These platforms are increasingly incorporating functionalities such as local signal processing, adaptive data fusion, and intelligent response behaviors, forming the basis of so-called “sensor-computing” systems[11-15]. In this review, we conceptually define sensor-computing integration as a hierarchical architecture spanning three distinct levels: On-sensor processing (Level 1), where functional materials perform signal pre-conditioning directly at the interface; on-device intelligence (Level 2), leveraging edge algorithms for real-time inference without continuous connectivity; and system-level fusion (Level 3), orchestrating cloud-based analytics for comprehensive health profiling. Recent literature illustrates the early stages of this trend. For example, a multimodal biosensor array integrating electrochemical and optical modalities has been engineered to detect glucose, lactate, and urea concurrently in sweat[16-18]. Ardalan et al. developed a cellulose-based wearable fluorescent patch with integrated microfluidic channels for simultaneous monitoring of glucose, lactate, pH, chloride, and sweat volume[19]. Similarly, a paper-based electrochemical platform enabled concurrent detection of glucose and lactate by employing a three-dimensional (3D) sweat diffusion pathway to improve sample collection efficiency and signal stability[20]. More recently, a soft epidermal optofluidic Raman system achieved real-time detection of glucose, lactate, and urea in sweat with high precision and long-term repeatability, illustrating the potential of optical-microfluidic integration for robust, noninvasive biochemical sensing[21]. Unlike existing reviews that predominantly focus on isolated domains ranging from flexible materials to algorithmic frameworks, this article centers on the synergy between multimodal sensing mechanisms and computational architectures. This paradigm shift mirrors the broader trend toward distributed and edge-level intelligence, where biosensors not only acquire data but also analyze and interpret it in situ. Such capabilities are particularly valuable for wearable and implantable systems, where latency, power constraints, and privacy concerns limit the feasibility of cloud-based analysis.
Guided by this hierarchical framework, this review provides a comprehensive overview of recent progress in integrated multimodal biosensing platforms, with particular emphasis on their material design, fabrication techniques, and functional deployment. We first introduce the fundamental categories of biosensors and representative classes of biomarkers, while highlighting the differences in capability and complexity between single-modal and multimodal systems. The discussion then turns to advances in device manufacturing, focusing on novel printing technologies that support spatially controlled, heterogeneous material integration. Particular attention is given to the convergence of diverse signal transduction mechanisms and the incorporation of machine learning frameworks, which together enhance the capacity for real-time data acquisition, fusion, and interpretation required for on-device computing hardware. Finally, we examine the remaining challenges in this field, including interfacial material compatibility, multimodal data harmonization, and translational barriers related to device reliability and regulation. The review concludes with a perspective on how future developments in materials science, AI, and system-level engineering may lead to the next generation of intelligent, scalable biosensing platforms with broad biomedical and environmental relevance.
BIOSENSING TECHNOLOGIES FOR HEALTHCARE
Biosensor technologies represent a cornerstone of modern analytical science, enabling specific, sensitive, and real-time detection of biological or chemical analytes across diverse fields including healthcare, environmental monitoring, and food safety. At their core, biosensors leverage the specificity of biological recognition processes combined with the precision of physical or chemical transduction to convert biological interactions into quantifiable signals[22]. As defined by the International Union of Pure and Applied Chemistry, a biosensor is a self-contained, integrated analytical device[23]. It comprises a biological recognition element, which includes biochemical receptors such as enzymes, antibodies, antigens, peptides, DNA, aptamers, or living cells. This element is in direct spatial contact with a transduction element, which may be electrochemical, optical, or mechanical in nature. The biological recognition element selectively interacts with target analytes, while the transduction element converts these interactions into measurable electrical, optical, or mechanical signals. The fundamental principle of a biosensor lies in its ability to translate specific biorecognition events. These events, such as interactions between enzymes and substrates, antibodies and antigens, or aptamers and their targets, are converted into measurable signals by the transduction element. This conversion enables quantitative or qualitative analysis of the target analyte, forming the basis for applications ranging from point-of-care diagnostics to on-site environmental monitoring[24].
This chapter aims to systematically elaborate on the core technologies underlying biosensors, with a focus on common types, their working mechanisms, and key applications. By examining the diverse array of biosensing platforms, we seek to provide a comprehensive understanding of their operational principles and practical implications. In the context of the hierarchical sensor-computing integration defined earlier, these transduction mechanisms constitute Level 1: on-sensor processing. By optimizing the material interfaces and signal conversion physics at this fundamental level, we lay the groundwork for high-fidelity data acquisition, which is a prerequisite for the subsequent algorithmic processing and system-level fusion discussed in later sections.
Representative biosensors for wearables
Biosensors for electrochemical biosensing
As one of the most widely used and mature biosensing technologies, electrochemical biosensors play a pivotal role in fields ranging from clinical diagnostics to environmental monitoring. Their core advantage lies in the ability to translate specific biorecognition events into quantifiable electrical signals. These events include interactions between enzymes, antibodies, aptamers, or nucleic acids and their respective targets. This transformation is mediated by redox reactions occurring at a functionalized working electrode, where the transfer of electrons during biochemical reactions generates measurable currents, potentials, or impedance changes.
To provide a comprehensive understanding of their operational mechanisms and design principles, this section first outlines the three primary electrochemical transduction modes that underpin these sensors. It then explores the common electrode materials and device architectures that enable their sensitivity, selectivity, and practical deployment in diverse applications.
(1) Amperometric biosensors
Amperometric biosensors are among the most widely utilized electrochemical sensing platforms, particularly for detecting small-molecule metabolites such as glucose, lactate, and uric acid (UA)[25,26]. As shown in Figure 1A, these sensors operate by applying a constant potential to the working electrode (WE), where the target molecule undergoes a redox reaction, typically catalyzed by an immobilized, target-specific oxidase enzyme [e.g., glucose oxidase (GOx), lactate oxidase, uricase][27-31]. The generated faradaic current is directly proportional to the analyte concentration and serves as the quantifiable output.
Figure 1. Electrochemical biosensor technologies based on different sensing principles. (A) Amperometric detection of metabolites using enzyme-functionalized electrodes; (B) Voltammetric sensing of proteins or nucleic acids via electrodes modified with antibodies or oligonucleotides. Detection strategies include multistep sandwich formats, binding-induced conformational switches, or proximity-based affinity recognition; (C) Representative architectures of ion-selective electrodes for potentiometric biosensing; (D) Conceptual illustration of conductometric sensing, where changes in electrical conductivity upon target interaction are used for analyte quantification. Reproduced with permission[44]. Copyright 2022, MDPI; (E) Impedimetric biosensing of analytes at varying concentrations, with Nyquist plots illustrating the system’s electrochemical behavior. The inset shows the randles equivalent circuit, comprising Rs, Rct, Cdl, and W. Reproduced with permission[45]. Copyright 2014, American Society for Microbiology. WE: Working electrode; AC: alternating current; Rs: solution resistance; Rct: charge-transfer resistance; Cdl: double-layer capacitance; W: Warburg impedance.
The amperometric sensing mechanism can be classified into three generations based on the mode of electron transfer between the enzyme and the electrode. First-generation sensors rely on endogenous cofactors such as dissolved oxygen as electron acceptors; the enzymatic reaction produces hydrogen peroxide (H2O2), which is subsequently oxidized or reduced at the WE to generate a current signal. Second-generation designs introduce redox mediators (e.g., ferricyanide, ferrocene derivatives, Prussian blue), which shuttle electrons between the enzyme’s active site and the electrode surface, improving electron transfer efficiency and reducing oxygen dependence[32]. Third-generation biosensors eliminate mediators entirely by enabling direct electron transfer (DET) between the redox center of the enzyme and the electrode, often facilitated by conductive nanomaterials or engineered enzyme-electrode interfaces[33]. Standard amperometric biosensors typically adopt a three-electrode configuration, comprising a WE functionalized with the biorecognition element, a counter electrode (CE) to complete the circuit, and a reference electrode (RE) to maintain a stable potential. This system enables rapid, sensitive, and selective detection under low power operation, making amperometric biosensors particularly suitable for wearable and portable diagnostic devices. Their relatively simple fabrication process, compatibility with flexible substrates, and strong electrochemical response have made them a foundational component in non-invasive health monitoring systems.
(2) Voltammetric biosensors
Voltammetric biosensors convert the selective binding of a target molecule into an electrical signal by applying a varying potential to the working electrode and recording the resulting current. In a typical format for protein or nucleic acid detection [Figure 1B], the electrode surface is functionalized with capture probes, for example antibodies, aptamers or complementary DNA strands, that bind the analyte of interest[34]. Because these biorecognition elements are not themselves electroactive, an electroactive label such as horseradish peroxidase, methylene blue or metal nanoparticle tags is introduced to generate a signal. When a potential sweep is carried out by cyclic voltammetry[35,36], scanning forward and reverse potentials, by differential pulse voltammetry[37,38], overlaying voltage pulses on a staircase waveform, or by square wave voltammetry[39,40], superimposing symmetrical pulses, the tagged complexes undergo oxidation or reduction at their characteristic potentials. Quantitation is achieved by measuring the height of the resulting current peaks or shifts in their position relative to a control. Modern implementations often employ screen-printed electrodes integrated with microfluidic flow injection systems to automate sample handling and allow a single-step, multiplexed assay. These configurations, and support the miniaturization needed for point-of-care applications, enhancing throughput, minimizing reagent use, and facilitating the miniaturization required for wearable diagnostics.
(3) Potentiometric biosensors
As depicted in Figure 1C, potentiometric biosensors quantify target analytes by recording the open-circuit potential difference between a selective sensing electrode and a stable reference electrode, with no net current flow[41]. The measured voltage adheres to the Nernst equation, shifting predictably as the activity of specific ions at the sensing membrane changes. Conventional ion-selective electrodes employ glass, crystalline or polymeric membranes to discriminate H+, K+, Na+, Ca2+ , and other biologically relevant ions, while solid-state variants replace internal electrolytes with conducting polymers or nanomaterial layers to enable miniaturization[42]. By tailoring the membrane composition and incorporating appropriate ionophores or recognition elements, potentiometric sensors achieve high sensitivity (tens of millivolts per decade change in concentration), rapid response, and ultralow power consumption[43]. These attributes make them well-suited for real-time monitoring of electrolytes in sweat, blood, or environmental samples, as well as for integration into portable and wearable diagnostic platforms.
(4) Conductometric biosensors
As illustrated in Figure 1D, conductometric biosensors detect analytes by measuring variations in the electrical conductance of the sensing element when a small alternating voltage is applied across two electrodes[44]. When target molecules bind to the sensor surface or undergo a reaction, the overall resistance changes and can be monitored directly or by using impedance spectroscopy. Because this method does not require a reference electrode and operates under low excitation voltages, conductometric sensors can be fabricated as thin-film devices using standard manufacturing processes and scaled down easily.
These sensors generally exhibit lower specificity and sensitivity than faradaic or potentiometric systems because the measured signal integrates multiple conductive pathways. Nonetheless, conductometric devices are widely used in environmental applications, where shifts in water conductivity provide a straightforward indicator of ionic composition and pollutant levels.
(5) Impedimetric biosensors
Impedimetric biosensors measure the frequency-dependent impedance of the electrode-electrolyte interface to detect molecular recognition events without the need for labels[45]. In practice, a small alternating potential is applied between the working and counter electrodes across a spectrum of frequencies while the resulting current is recorded [Figure 1E]. The system is commonly represented by the Randles equivalent circuit, which comprises the solution resistance (Rs), attributable to the bulk electrolyte; the double-layer capacitance (Cdl) at the electrode surface; the charge-transfer resistance (Rct) associated with interfacial redox reactions; and the Warburg impedance (W), reflecting mass-transport limitations[46]. When plotted as a Nyquist diagram, the high-frequency regime appears as a semicircular arc whose diameter directly corresponds to Rct, whereas the low-frequency regime rises at approximately 45 degrees, illustrating the influence of diffusion processes. Binding of a target analyte to a recognition layer on the electrode modifies local charge density and introduces steric hindrance, thereby impeding access of redox probes such as ferrocyanide or ferricyanide and increasing both Rct and overall impedance[47]. The addition of a redox mediator may shift these impedance features by altering electron-transfer kinetics. Although this label-free approach yields detailed mechanistic information and is amenable to miniaturization for point-of-care applications, nonspecific adsorption and complex sample matrices can compromise sensitivity and selectivity, necessitating careful surface engineering and assay validation[48].
Biosensors for microwave biosensing
Microwave biosensor technology offers promising solutions for continuous, non-invasive glucose monitoring. Microwave sensors harness the power of low-energy, safe electromagnetic waves that can penetrate through skin, fat, and muscle tissues to reach blood vessels and interstitial fluids[49,50]. The fundamental principle is that variations in glucose concentration alter the dielectric properties of tissues, including how water molecules and other components interact with microwave energy[51,52]. By detecting these subtle permittivity shifts, microwave sensors can generate real-time glucose readings[53-55]. Microwave-based monitoring offers several advantages, including deep tissue penetration on the millimeter-to-centimeter scale, rapid response times of less than one second, inherent safety, and compatibility with wearable integration[56-59]. External factors such as temperature and hydration may also affect dielectric responses, underscoring the need for robust calibration in practical applications[60-63].
Recent developments in microwave biosensing have focused on optimizing resonant structures and transmission lines to enhance sensitivity and miniaturization. Among these, metamaterial-inspired elements such as split-ring resonator (SRR) and complementary split-ring resonator (CSRR) are particularly attractive due to their strong electromagnetic field localization and compact size[64,65]. These resonators exhibit frequency shifts in reflection (S11) or transmission (S21) parameters when interacting with biological tissues, enabling detection of subtle changes in permittivity[66-68]. The CSRR, in particular, enhances field confinement by etching resonant rings onto the ground plane of a transmission line, which intensifies the local electric field and improves sensitivity[55,69]. Figure 2A presents a representative CSRR-based biosensor design featuring a portable radar-driven configuration capable of non-invasive fingertip glucose detection[55]. Although SRR and CSRR devices exhibit excellent sensitivity, their performance can be affected by fabrication tolerances and environmental conditions, which necessitate precise calibration and multi-cell configurations to ensure measurement stability[70,71].
Figure 2. Representative designs and working principles of microwave biosensors for non-invasive and wearable monitoring. (A) Conceptual illustration and fabricated prototype of a portable radar-driven sensor employing a four-cell CSRR configuration for non-invasive blood glucose measurement from the fingertip. Reproduced with permission[55]. Copyright 2020, Springer; (B) Fabrication process and structural schematic of a gated graphene microwave CPW sensor. Reproduced with permission[63]. Copyright 2024, Wiley; (C) Design of a tunable, non-invasive, wearable electromagnetic multi-sensing system inspired by vascular anatomy, showing slot antenna and band-reject filter prototypes adapted for hand and arm glucose monitoring. Reproduced with permission[76]. Copyright 2020, AAAS; (D) Schematic of a wearable, flexible, body-matched electromagnetic sensing system employing multi-sensing and multi-location detection to correlate reflected EM waves with blood glucose variations. Reproduced with permission[77]. Copyright 2022, Springer. SMA: SubMiniature version A; PCB: printed circuit board; Rx: Receiver; Tx: Transmitter; CSRR: complementary split-ring resonator; PDMS: polydimethylsiloxane; CPW: coplanar waveguide; FR4: flame retardant 4; EM: electromagnetic; VNA: vector network analyzer.
Another important class of microwave biosensors is based on graphene-integrated coplanar waveguide (CPW), which combine the superior electrical conductivity of graphene with the high sensitivity of planar microwave transmission lines[63]. A CPW consists of a central conductor flanked by ground planes on the same substrate surface, forming a confined electromagnetic region that interacts with biological media placed above[72-75]. As shown in Figure 2B, embedding graphene within the CPW enhances field confinement and amplifies small dielectric variations caused by glucose concentration changes[63]. These sensors are compact and sensitive, though challenges remain in impedance matching and maintaining long-term operational stability under physiological conditions.
Building upon these principles, flexible and wearable microwave devices have emerged as a new frontier for biomedical sensing. The incorporation of soft, conformal materials allows these systems to maintain stable skin contact and continuously monitor multiple physiological parameters in real time. As illustrated in Figure 2C, vascular-inspired sensor systems replicate the geometry of veins and arteries to achieve depth-selective glucose detection. These architectures employ slot antennas and band-reject filters to optimize coupling with subsurface blood vessels[76]. Another innovative design approach, as shown in Figure 2D, involves body-matched electromagnetic sensors that can simultaneously measure signals at different on-body locations while compensating for environmental factors such as temperature, humidity, and motion[77].
In summary, microwave biosensors that utilize metamaterial resonators, graphene-based CPWs, and flexible vascular-inspired architectures are advancing toward practical, non-invasive glucose monitoring. Future work should emphasize improved calibration, miniaturization, and multimodal data fusion to enhance sensitivity, stability, and reliability in clinical and wearable healthcare applications.
Biosensors for optical biosensing
Optical biosensors convert light-matter interactions in biological media into measurable signals. Depending on the underlying phenomenon, they exploit reflection, refraction, scattering, absorption, or plasmonic resonance to deliver high sensitivity and selectivity[78-80]. These sensors are generally categorized into several main types, including surface plasmon resonance (SPR) sensors, localized surface plasmon resonance (LSPR) sensors, fluorescence-based biosensors, microfluidic photonic platforms, and smartphone-integrated optical systems. Together, these diverse modalities demonstrate the versatility of optical transduction in biomedical applications[81-83].
In the realm of plasmonic sensing, a critical distinction exists between propagating SPR on planar films and LSPR on nanostructures. While conventional SPR offers high sensitivity for interface kinetics, it typically requires bulky prism coupling. In contrast, LSPR arises from electron oscillations confined within metallic nanoparticles, eliminating the need for prisms and enabling flexible device integration. As illustrated in Figure 3A, a stretchable and highly sensitive LSPR-based optical strain sensor is utilized for human-activity monitoring[84]. This sensor embeds high-density gold nanoparticles (AuNPs) within a soft elastomeric matrix. Unlike geometric waveguides, the sensing mechanism relies on the modulation of LSPR coupling. When the sensor is stretched or bent, the inter-particle distance between AuNPs changes, significantly altering the optical absorption or scattering intensity. This nanostructure-enhanced sensitivity enables the precise detection of minute physiological signals, ranging from respiration to pulse beats, while maintaining the mechanical compliance required for wearable skins.
Figure 3. Representative optical biosensor configurations for biochemical sensing. (A) Schematic of a stretchable optical strain sensor based on LSPR for human-activity monitoring. Reproduced with permission[84]. Copyright 2019, American Chemical Society; (B) Plasmonic fluorescence biosensor based on PCR amplification for rapid and sensitive biomolecular analysis. Reproduced with permission[85]. Copyright 2021, Royal Society of Chemistry; (C) Schematic illustration of a magnetic nanochain-integrated microfluidic biochip platform for multiplexed biosensing. Reproduced with permission[87]. Copyright 2018, Springer; (D) Ultra-compact dual-channel integrated CO2 infrared gas sensor based on on-chip NDIR technology. Reproduced with permission[88]. Copyright 2024, Springer. GNP: Gold nanoparticle; PDMS: polydimethylsiloxane; IR: infrared; LED: light-emitting diode; PPT: plasmonic photothermal; PCR: polymerase chain reaction; PMN: plasmonic magnetic nanoparticle; SERS: surface-enhanced Raman scattering; MEMS: PCB: printed circuit board; LSPR: localized surface plasmon resonance; NDIR: non-dispersive infrared.
Complementing index-based sensing, emissive readouts exploit fluorescence intensity or lifetime to provide molecular specificity and ultralow detection limits. Figure 3B presents a fluorescence biosensor employing plasmonic enhancement in conjunction with polymerase chain reaction (PCR) amplification[85]. Fluorescence-based biosensors operate by detecting the light emitted from fluorescent molecules following optical excitation, and the signal intensity or lifetime is directly related to analyte concentration. In this example, metallic nanostructures amplify the local electromagnetic field, which significantly increases fluorophore excitation and emission efficiency. When coupled with PCR amplification, this effect enables highly sensitive and rapid detection of nucleic acid biomarkers. Such designs demonstrate how combining photonic field enhancement with biochemical amplification mechanisms can substantially improve analytical performance in biosensing[86].
To manage transport phenomena and enable scalable automation, microfluidic optics structure both light paths and analyte motion within confined volumes. The integration of multiple detection principles is further demonstrated in Figure 3C, which shows a magnetic nanochain-assisted microfluidic biochip[87]. Microfluidic optical biosensors typically direct light through transparent channels containing the analyte, where variations in refractive index or absorption modulate the transmitted or scattered light signal. In this particular system, magnetic nanochains are used to align and concentrate target molecules within microchannels, thereby improving the interaction efficiency between light and biological components. The result is faster response time, higher sensitivity, and improved reproducibility. This microfluidic configuration exemplifies how optical and magnetic control strategies can be combined to achieve compact, automated, and high-throughput sensing platforms[87].
For environmental and physiological gas analysis, miniaturization is essential for portable deployment. Figure 3D shows an ultra-compact dual-channel integrated CO2 infrared gas sensor. As depicted in the device architecture, this platform leverages the non-dispersive infrared (NDIR) absorption principle on a photonic integrated circuit[88]. The innovative “dual-channel” design incorporates a reference waveguide alongside the sensing arm to allow for differential measurement, effectively compensating for environmental drifts and light source fluctuations. By confining light within micro-scale structures to interact with the target gas, this on-chip sensor achieves high selectivity and stability in a minimal footprint, exemplifying the potential of integrated photonics for robust environmental and breath gas diagnostics.
Overall, optical biosensors represent a rapidly evolving class of analytical tools that translate light-matter interactions into biochemical information. Different approaches provide complementary advantages by aligning sensing physics with deployment needs, where SPR sensors enable real-time label-free detection, LSPR sensors facilitate flexible motion tracking, fluorescence biosensors offer high sensitivity and selectivity, microfluidic optical systems deliver scalability and automation, and smartphone-based devices increase accessibility for field applications[84,85,88-90]. Ongoing advances in materials, optical design, and signal processing are expected to drive the integration of these modalities, paving the way for hybrid, intelligent, and personalized biosensing platforms suitable for both clinical and on-site use[91-94].
Biosensors for gas detection
Gas biosensors are emerging analytical platforms that integrate biological recognition elements (e.g., enzymes, antibodies, aptamers/peptides/DNA, living cells, and olfactory receptors) with physical transducers to detect volatile organic compounds (VOCs) and gaseous biomarkers from breath, skin emissions, indoor air, and environmental samples. By operating within a thin, hydrated microenvironment that sustains biorecognition activity under humid, complex matrices, these systems enable continuous or near-real-time measurements suited to breathomics, environmental surveillance, food safety, and occupational exposure assessment. Compared with conventional chemoresistive gas sensors, they offer lock-and-key selectivity, biological signal amplification via catalytic or cellular pathways, and robust performance in high-humidity conditions, while supporting point-of-care deployment through miniaturization, rapid response, and electronic readouts[95,96].
The choice of biorecognition play a critical role for sensing performance. Enzymatic systems continue to be the backbone of continuous gas sensing, where immobilized oxidases and dehydrogenases quantify targets through processes such as oxygen consumption, hydrogen peroxide generation, or the conversion between NADH and NAD+. These systems provide selective and humidity-resilient detection for analytes like ethanol, formaldehyde, and acetone across ppb to sub-ppm ranges[97-99]. Advanced implementations amplify signals through multi-enzyme cascades, oxygen/NAD-linked amperometry, or triphasic gas-liquid-solid reactors that enhance mass transfer[100]. Optical “bio-sniffers” extend enzyme assays into fluorometric formats, using O2-quenched probes or NADH autofluorescence, to deliver linear, real-time readouts within clinically relevant breath windows with low coefficients of variation[101,102]. As depicted in Figure 4A, whole-cell biosensors offer integrative, systems-level responses: recombinant lux reporters quantify gas toxicity (for example, benzene) with dose dependence and long shelf-life, and immobilized promoter arrays resolve VOC fingerprints from contaminated foods; coupled with machine learning, such arrays can achieve near-perfect discrimination of pre-mold states and commodity identity[103,104]. Olfactory-receptor (OR) biosensors, built by co-expressing insect or mammalian ORs with olfactory receptor co-receptor (Orco) and calcium ion (Ca2+) indicators, emulate biological olfaction to reach ppb sensitivity and quantify mixtures using relative-comparison protocols that mitigate drift and nonlinearity; co-expressing multiple ORs in a single cell line broadens dynamic range under cross-sensitivity[105]. As illustrated in Figure 4B and C, biofunctional nanocomposites enhance selectivity by integrating bioligands with two-dimensional conductors, such as peptide- or DNA-modified MXenes or graphene[106,107]. These composites achieve ppb-level detection limits, provide enhanced responses compared to pristine materials, and demonstrate robust breath classification for conditions like cancer and other pathophysiologies when analyzed with neural networks[108].
Figure 4. Representative designs and working principles of gas biosensors. (A) Schematic illustration of metal-based nanosystems with improved physico-chemical attributes through different strategies for enhanced detection of LC breath biomarkers in humans with smart and point-of-care modules. Reproduced with permission. Reproduced with permission[103]. Copyright 2024, American Chemical Society; (B) Schematic diagram of the human olfactory system and the sensor array system for gas identification; (C) Schematic diagram of the instant testing platform for cancer recognition with the assistance of machine learning (ML). (B and C) Reproduced with permission[107]. Copyright 2025, American Chemical Society.
Transduction modalities and device architectures determine how signals are captured and delivered. Electrochemical formats (amperometric, potentiometric, impedimetric) convert enzymatic redox or binding events into current, potential, or impedance changes; NAD-dependent dehydrogenase schemes expand target scope, and gas-liquid-solid triphasic photoelectrochemical reactors markedly increase oxygen utilization and mass transfer, enabling ultralow limits with rapid recovery in complex matrices[32,100]. Optical biosensors implement enzyme-coupled fluorescence or chemiluminescence with fiber optics for remote readouts and microdomain mapping of VOC release on skin and in breath[102]. Acoustic biosensors (quartz crystal microbalance; shear-horizontal surface acoustic wave, SH-SAW) track mass loading with millisecond kinetics; dual-channel SH-SAW chips with in-situ references and inkjet-printed capture coatings have demonstrated combined 100% sensitivity and 100% specificity for dual HIV antibody biomarkers in a pilot clinical study using microliter samples and sub-minute electronic outputs, highlighting compact, connected diagnostics for point-of-care screening[109]. Field-effect transistor (FET) biosensors couple catalytic or affinity layers (e.g., metal-phthalocyanine-graphene composites) to high-mobility channels for label-free, real-time gas measurements with picomolar sensitivity to species such as H2S and strong anti-interference characteristics, enabling in situ monitoring of cellular fluxes and inhibitor pharmacology[110]. Chemiresistive nanoarrays functionalized with peptides or DNA on 2D materials deliver low-power, room-temperature operation, programmable selectivity, and scalable array integration, supporting nose-on-chip platforms that, when paired with machine learning, resolve multi-class odor patterns and disease cohorts with high accuracy[107].
Biosensors for piezoelectric biosensing
Piezoelectric biosensors convert mechanical stress into an electrical signal by exploiting the piezoelectric effect of certain crystals. When an alternating voltage causes the crystal to vibrate at its resonance frequency, any additional mass on the surface slows the oscillation and lowers the measured frequency. This frequency shift is proportional to the mass of biomolecules bound to the surface.
Expanding the scope from surface mass sensing to deep-tissue imaging, Figure 5A illustrates a wearable cardiac ultrasound imager. This platform utilizes a matrix of piezoelectric transducers embedded in a stretchable substrate to conform to the curvature of the human chest[111]. The transducers operate by transmitting ultrasonic waves into the body and receiving the echoes reflected from internal organs. By converting these mechanical echoes back into electrical signals, the device constructs real-time images of the cardiac structure. This application highlights the capability of piezoelectric sensors to perform continuous, non-invasive hemodynamic monitoring, capturing critical parameters such as stroke volume and cardiac output without the need for bulky hospital equipment.
Figure 5. Representative architectures and working principles of piezoelectric biosensors. (A) Schematic of a wearable cardiac ultrasound imager based on stretchable piezoelectric transducer arrays for real-time heart monitoring. Reproduced with permission[111]. Copyright 2023, Springer; (B) Configuration of a SAW biosensor for E. coil detection, illustrating antibody immobilization on the delay line and bacterial capture. Reproduced with permission[112]. Copyright 2024, Royal Society of Chemistry; (C) Real-time monitoring of S. aureus lysis on a QCM sensor, showing the experimental setup for tracking mass and dissipation changes during phage/enzyme treatment. Reproduced with permission[114]. Copyright 2025, Springer. IDT: Interdigital transducer; E. coli: Escherichia coli; S. aureus: Staphylococcus aureus; QCM: quartz crystal microbalance.
For molecular-level detection, surface acoustic wave (SAW) devices offer high sensitivity for pathogen screening. Figure 5B details the configuration of a SAW biosensor designed for foodborne pathogen detection[112]. In this architecture, an input interdigital transducer (IDT) generates acoustic waves that travel across the piezoelectric substrate’s delay line. A gold film deposited on this delay line is functionalized with antibodies specific to the target pathogen (Escherichia coli, E. coli) via self-assembled monolayers (SAMs). Upon exposure to a contaminated sample, the bacteria bind specifically to the antibodies. This binding event introduces a significant mass load and viscoelastic change to the sensor surface, which perturbs the propagating acoustic wave. The resulting shift in wave velocity or amplitude allows for the rapid, label-free quantification of bacterial contamination directly from complex matrices.
Complementing SAW technology, the quartz crystal microbalance (QCM) remains a powerful tool for monitoring dynamic biological processes. In a typical QCM sensor, a thin quartz wafer with metal electrodes on each side is driven by an oscillator circuit to vibrate in a shear mode. Surface immobilization of antibodies or other recognition elements does not significantly alter the baseline frequency[113]. Once the target analyte binds, the added mass produces a measurable decrease in resonance frequency according to the Sauerbrey equation. Figure 5C demonstrates the real-time monitoring of bacterial lysis on a QCM platform[114]. The approach simulates phage antibiotic synergy by examining bacterial degradation induced by either lysostaphin or phage P68. Staphylococcus aureus (S. aureus) cells were immobilized on a poly-L-lysine-coated QCM surface. Changes in resonance frequency and energy dissipation were recorded as indicators of bacterial membrane disruption and biofilm breakdown[114]. The system enables tracking of the mass variations caused by cell lysis and release of intracellular contents. Additionally, a flow-through configuration permitted simultaneous nutrient delivery and metabolite removal, allowing comprehensive assessment of the dynamic interactions between bacteriophages and their bacterial hosts.
Biosensors for thermal sensing
Thermal and thermo-responsive interfaces play a dual role in multimodal biosensing platforms. They serve as diagnostic tools for monitoring physiological temperature variations and as energy harvesting units to power wearable electronics. While traditional calorimetric biosensors focus on detecting heat from enzymatic reactions[115,116], recent advances have pivoted toward thermoelectric generators (TEGs) that exploit the Seebeck effect[117,118]. These devices convert the temperature gradient between the human body and the environment into electrical energy. This mechanism enables self-powered operation while simultaneously providing precise temperature mapping, which is a critical capability for calibrating temperature-sensitive biochemical sensors in a multimodal system.
To address the power constraints of wearable systems, Figure 6A illustrates a self-powered electronic skin with multisensory functions based on thermoelectric conversion[119]. In this architecture, flexible thermoelectric materials are integrated into a skin-conformable patch. The device operates by capturing body heat to generate a voltage potential. This potential serves as both a sensing signal for continuous skin temperature monitoring and a renewable power source for the integrated circuit. This dual-functionality allows the platform to detect subtle physiological thermal fluctuations, including those indicative of inflammation or metabolic changes, without the need for external batteries. Consequently, this design significantly reduces the device footprint and maintenance burden for long-term health monitoring.
Figure 6. Thermoelectric platforms for self-powered thermal sensing and energy harvesting. (A) Schematic of a self-powered electronic skin based on thermoelectric conversion, enabling dual functions of physiological temperature monitoring and energy harvesting from body heat. Reproduced with permission[119]. Copyright 2020, Wiley; (B) Structural design and optical characterization of a stretchable temperature electronic skin, featuring high-performance Bi-Te micro-thermoelectric generators connected via an island-bridge architecture for distributed mini-region sensing. Reproduced with permission[120]. Copyright 2023, Wiley. F-TEG: Flexible thermoelectric generator; PDMS: polydimethylsiloxane; STES: self-powered temperature electronic skin; Bi-Te: Bismuth Telluride.
However, integrating rigid high-performance thermoelectric materials into soft wearables presents mechanical challenges. Addressing the conflict between flexibility and energy efficiency, Figure 6B presents a self-powered temperature electronic skin based on an “island-bridge” structure and Bismuth Telluride (Bi-Te) micro-thermoelectric generators[120]. As shown in the detailed optical images, rigid Bi-Te pillars act as the islands and are interconnected by stretchable conductive wiring that functions as the bridges. This design mechanically isolates the active thermoelectric elements from strain, ensuring that the device maintains stable performance even under stretching or bending. The array configuration enables distributed mini-region sensing, allowing for high-resolution spatiotemporal mapping of skin temperature. Such precise thermal profiling is essential not only for fever detection but also for decoupling temperature interference from concurrent electrochemical or mechanical signal recordings in a multimodal setup.
While each sensing modality discussed above possesses unique transduction characteristics, their practical implementation in “sensor-computing” systems requires a careful trade-off between performance and integration constraints. Table 1 provides a comparative analysis of these key biosensing modalities, contrasting their mechanisms, sensitivity, response times, and power consumption. Crucially, the table evaluates their suitability for sensor-computing integration, highlighting how specific limitations can be addressed through hierarchical computing architectures (Level 2/3 processing).
Comparative analysis of key biosensing modalities for wearable and integrated platforms
| Sensing modality | Key mechanism | Sensitivity & range | Response time | Power consumption | Suitability for sensor-computing integration | Key challenges |
| Electrochemical | Redox reactions (Amperometric), Charge transfer (Impedimetric), Ion activity (Potentiometric) | High (nM-µM) Broad dynamic range | Fast (Seconds) | Low (µW range) Ideal for continuous edge computing | High Simple analog front-end; signal is easily digitized for on-device ML (Level 2) | Susceptible to electrode fouling and drift; requires reference electrodes[32,48] |
| Microwave | Dielectric permittivity shifts / Resonance frequency changes (SRR, CSRR) | Moderate (mg/dL) Depth-selective detection | Very fast (< 1 s) | Moderate (mW range) Depends on RF circuit design | Medium Non-invasive data is noisy; relies heavily on system-level fusion (Level 3) for accuracy | Environmental interference (temp/hydration); sensitive to motion artifacts[60-63] |
| Optical | Light-matter interaction (SPR, Fluorescence, Raman) | Very High (fM-aM) High specificity | Fast (Real-time) | High (mW-W) Requires light source & detector | Medium Data is complex (spectral/image); requires robust processing (e.g., CNNs) at Level 2/3 | High integration cost; sensitive to optical alignment and ambient light [87] |
| Gas | Chemiresistive (FET), Electrochemical, or Acoustic transduction of VOCs | High (ppb-ppm) Specific to breath/skin VOCs | Moderate (Seconds to mins) | Low to Moderate Low for FET/chemiresistive types | High Pattern recognition algorithms (e.g., electronic nose) are essential for selectivity | Humidity interference; selectivity in complex gas mixtures[99,105] |
| Piezoelectric | Mass-induced frequency shift (QCM, SAW) | High (ng-pg mass) Label-free detection | Fast (Seconds) | Moderate Requires oscillator circuits | Medium Frequency shift is a direct digital-friendly signal suitable for edge processing | Sensitive to mechanical vibrations and viscosity changes[121] |
| Thermal | Calorimetric (Heat reaction) or Thermoelectric effect | Moderate (µJ heat change) Self-powered potential | Slow to Moderate (Mins) | Very low / Negative Can harvest energy (Self-powered) | High Unique potential for self-powered “sensor-computing” nodes without batteries | Requires strict temperature control/calibration; slower response[122] |
Representative biomarkers in different body fluids
Blood biomarkers
Blood serves as a rich reservoir of biomarkers that play a crucial role in assessing systemic health. Glucose, one of the most extensively monitored biomarkers, is a core metabolic indicator. With a normal fasting range of 3.9 - 6.1 mmol/L, glucose is of utmost importance in diabetes management. Deviations from this range, such as hyperglycemia or hypoglycemia, are indicative of metabolic disorders[123]. Biosensors for glucose commonly utilize glucose oxidase to catalyze redox reactions, effectively converting glucose concentration into electrical signals. Advanced designs incorporate nanomaterials, enabling micromolar level sensitivity for continuous blood glucose monitoring[124].
In addition to glucose, blood contains a plethora of other vital biomarkers. Lipids, including cholesterol [total, low-density lipoprotein (LDL), high-density lipoprotein (HDL)] and triglycerides, are key indicators. Enzymatic or immunological biosensors are employed to monitor these lipids, providing valuable insights into cardiovascular risk. Hormones like insulin, which regulates glucose metabolism, and thyroid hormones such as triiodothyronine (T3) and thyroxine (T4), are essential for diagnosing endocrine disorders[125]. Proteins such as C-reactive protein (CRP), an inflammatory marker, and troponin, which indicates myocardial infarction, are detected through highly specific immunosensors[126]. Electrolytes like K+, Na+, and Ca2+ are also significant. Imbalances in their levels can signal renal or metabolic problems, and they are measured using potentiometric ion-selective electrodes[127].
Sweat biomarkers
Sweat, secreted by eccrine glands on the skin surface, serves as a rich source of non-invasively accessible physiological information, making it an attractive medium for wearable monitoring devices. The key analytes present in sweat offer valuable insights into multiple physiological processes.
Lactate, a byproduct of anaerobic metabolism, is a crucial biomarker. During physical activity, muscle cells switch to anaerobic respiration when oxygen supply is insufficient, leading to lactate production. Sweat lactate concentrations typically range from 1 to 10 mM, and these levels are directly proportional to muscle exertion[128]. Biosensors that utilize lactate oxidase can catalyze the oxidation of lactate, generating current signals. These sensors have been shown to accurately reflect changes in exercise intensity, providing athletes and trainers with immediate feedback on energy expenditure and muscle fatigue[129].
Electrolytes, including sodium ions (Na+, with a concentration range of 5-40 mM) and potassium ions (K+, 3-15 mM) in sweat, are vital for maintaining proper physiological function. The levels of these electrolytes mirror the body’s hydration status and electrolyte balance. Potentiometric sensors equipped with ion-selective membranes, such as sodium-selective glass membranes for Na⁺ detection, play a fundamental role in preventing dehydration[130]. By precisely measuring electrolyte concentrations in sweat, these sensors can alert individuals to the need for fluid and electrolyte replacement[131].
Metabolites in sweat, such as urea and creatinine, also contribute to the assessment of overall metabolic health. Urea is an end product of protein metabolism and serves as an indicator of renal function. Creatinine, produced from muscle metabolism, provides additional information about metabolic activity[132]. Enzymatic reactions are commonly employed to detect these metabolites. For instance, urease can be used to break down urea, and the resulting products can be measured to determine urea concentration[133]. The detection of these metabolites in sweat, offers a non-invasive means to monitor renal function and overall metabolic status over time[134].
Tumor biomarkers
Tumor markers, a class of biomolecules aberrantly overexpressed in cancer cells or tissues, play a pivotal role in facilitating early cancer detection, evaluating disease prognosis, and monitoring treatment responses. These biomarkers, which include proteins, glycoproteins, and nucleic acids, exhibit altered expression profiles in pathological states, making them valuable indicators for clinical oncology. Key tumor markers and their clinical associations are as follows:
Carcinoembryonic antigen (CEA): A glycoprotein exhibits elevated serum levels in various malignancies such as colorectal, lung, and breast cancers. In healthy populations, normal serum CEA concentrations are generally maintained below 5 ng/mL[135], with elevations beyond this range potentially indicating the need for further diagnostic evaluation. This biomarker’s association with multiple cancer types underscores its utility in clinical settings for aiding in the identification of pathological processes related to malignant growths[136,137].
Prostate-specific antigen (PSA): A serine protease predominantly secreted by prostate epithelial cells, serving as a specific marker for prostate cancer. Clinically, serum PSA levels exceeding 4 ng/mL are considered indicative of potential malignancy, though this threshold may vary based on age and individual risk factors[138].
Alpha-fetoprotein (AFP): A fetal glycoprotein re-expressed in hepatocellular carcinoma (HCC). Elevated serum AFP levels (> 20 ng/mL) are recognized as a critical diagnostic indicator for HCC, particularly when combined with imaging modalities[139].
CA125: A mucin-like glycoprotein widely used as a biomarker for ovarian cancer. Beyond aiding in initial diagnosis, CA125 levels are clinically valuable for monitoring treatment efficacy and detecting disease recurrence[140].
In summary, tumor markers, as biomolecules aberrantly overexpressed in cancer cells or tissues, play a crucial role in early cancer detection, prognostic evaluation, and monitoring of treatment responses. Key markers such as CEA, PSA, AFP, and CA125 are each associated with specific types of cancer, providing valuable indicative information for clinical oncology. Biosensors, relying on high-specificity recognition mechanisms like antibody-antigen interactions and nucleic acid hybridization, have emerged as powerful tools for detecting these tumor markers[141,142]. Their significance in improving patient survival rates through timely intervention is paramount.
Food safety-related analytes
Biosensors designed for food safety applications primarily target harmful substances, serving as a critical barrier to prevent contamination-induced illnesses. Their utility spans the detection of diverse hazardous agents, with key focus areas as follows:
Pesticides: Organophosphates and carbamates, two major classes of pesticides, are commonly monitored using enzyme inhibition assays. A typical example is the inhibition of acetylcholinesterase, whose activity is quantitatively suppressed in the presence of these compounds. This mechanism enables sensitive quantification of pesticide residues in matrices such as fruits, vegetables, and aqueous samples, with detection limits ranging from nanograms per milliliter to micrograms per milliliter[143,144].
Pathogens: Microbial contaminants, including pathogenic bacteria (e.g., E. coli O157:H7, Salmonella spp.) and viruses, are detected through two primary strategies. Nucleic acid-based biosensors, often integrated with polymerase chain reaction (PCR) amplification, enable highly specific identification of genetic signatures. Alternatively, antibody-based biosensors leverage specific recognition of surface antigens, facilitating rapid screening of intact pathogens[145].
Toxins: Mycotoxins (e.g., aflatoxin B1) and bacterial toxins (e.g., botulinum toxin) represent a class of low-molecular-weight hazards requiring ultra-sensitive detection. Immunosensors, which rely on high-affinity antibody-toxin interactions, and aptasensors, utilizing target-specific nucleic acid aptamers, are particularly effective in this context. These platforms achieve trace-level quantification with exceptional specificity, ensuring the accurate identification of even minute toxin concentrations[146-148].
Environmental pollutants and toxins
Environmental biosensors serve as pivotal tools for monitoring pollutants, playing a crucial role in evaluating risks to ecosystems and human health through the specific detection of hazardous substances. Their application encompasses the monitoring of diverse environmental pollutants, with key focuses outlined as follows:
Heavy metals: Toxic heavy metal ions such as lead (Pb2+), mercury (Hg2+), and cadmium (Cd2+) are primary targets of environmental biosensors. These sensors typically employ recognition elements like metal-binding proteins (e.g., metallothioneins) or DNA enzymes, which exhibit high specificity toward heavy metal ions[149,150]. The remarkable sensitivity of such systems allows for detection limits as low as parts per billion (ppb) to parts per trillion (ppt), enabling the identification of even trace-level heavy metal contamination in environmental matrices such as water, soil, and sediment.
Organic pollutants: A wide range of organic pollutants, including polycyclic aromatic hydrocarbons (PAHs), pharmaceuticals, and industrial chemicals, are monitored using specialized biosensors. Microbial biosensors, which utilize pollutant-degrading bacteria engineered to produce detectable signals upon exposure to specific organic compounds, are widely applied for this purpose. Alternatively, molecularly imprinted polymers (MIPs) serve as effective recognition elements in biosensors for organic pollutant detection, leveraging their high selectivity for target analytes. These approaches facilitate accurate quantification of organic pollutants in complex environmental samples[151,152].
Neurotransmitters and neuromodulators
Neurotransmitters, as key mediators of neural signaling, play a pivotal role in maintaining normal nervous system function. Imbalances in their concentrations are closely associated with the pathogenesis of various neurological and neuropsychiatric disorders, making their accurate detection critical for both disease mechanism research and clinical diagnosis.
Dopamine: This neurotransmitter is primarily involved in the regulation of reward pathways and motor control. A deficiency in dopamine levels is a hallmark of Parkinson’s disease, characterized by motor dysfunction, while excessive dopamine activity has been implicated in the pathophysiology of schizophrenia. The biosensors capitalize on the electrochemical oxidation of dopamine, generating measurable currents that correlate with its concentration. Such systems achieve sub-micromolar sensitivity, enabling precise quantification in complex biological matrices such as brain tissue extracts or cerebrospinal fluid[153,154].
Serotonin (5-hydroxytryptamine, 5-HT): Functionally, serotonin modulates mood, sleep-wake cycles, and emotional regulation, with reduced levels strongly linked to depressive disorders. Optical biosensors have been developed for serotonin detection, leveraging either fluorescent probes that undergo specific spectral changes upon binding to 5-HT or aptamers with high affinity for serotonin[155,156]. The application of specialized biosensors in detecting these neurotransmitters not only enables precise monitoring of their dynamics in neural interfaces but also contributes to advancing our understanding of neurological disorders, thereby laying the groundwork for the development of novel diagnostic and therapeutic interventions.
To systematically guide the design of multimodal platforms, it is essential to match the biochemical nature of these analytes with the most appropriate transduction interfaces. Table 2 classifies representative biomarkers into distinct categories spanning from simple ions to complex macromolecules, mapping them to their preferred sensing modalities and recognition elements. This classification underscores the rationale for multimodal integration: by combining sensors that target distinct biochemical classes, integrated platforms can achieve a more holistic and accurate assessment of physiological states.
Classification of biomarkers by biochemical nature and their preferred sensing modalities for integrated platforms
| Biomarker class | Representative analytes | Primary body fluid | Preferred sensing modality & Recognition element | Rationale for multimodal integration |
| Ions (electrolytes) | Na+, K+, Ca2+, pH | Sweat, urine, ISF | Potentiometric (ion-selective membrane) | Calibration: real-time pH or ion data is essential to correct signal drift in enzymatic sensors (e.g., glucose sensors are pH-sensitive) |
| Small molecules (metabolites) | Glucose, lactate, UA, creatinine | ISF, sweat, blood | Electrochemical (enzymatic) (e.g., glucose oxidase, uricase) | Metabolic profiling: these fluctuate rapidly; combining them with physical signals allows for context-aware health monitoring (e.g., exercise fatigue) |
| Proteins & Hormones | Insulin, cytokines (CRP), tumor markers (CEA, PSA), troponin | Blood, serum, saliva | Immunosensor/Aptasensor (e.g., antibody, aptamer) | Disease diagnosis: High specificity is required. Multimodal fusion (e.g., combining CRP with temperature/heart rate) improves diagnostic accuracy for infection or CVD |
| Nucleic acids | DNA, RNA, miRNA | Blood, saliva | Optical/Piezoelectric (genosensor) (e.g., DNA hybridization, PCR) | Pathogen screening: often requires amplification. Integrating microfluidics with optical readouts enables “Lab-on-a-Chip” genetic analysis |
| Gaseous molecules | VOCs, ethanol, NOx | Breath, skin emissions | Chemiresistive / Optical (Bio-sniffer) (e.g., metal oxides, olfactory receptors) | Non-invasive breathomics: breath analysis complements blood/sweat analysis, offering a truly holistic view of metabolic status |
Advantages and challenges of multimodal biosensors
Advantages of multimodal biosensors
Multimodal biosensors offer a range of significant advantages over single-mode systems by leveraging the complementary strengths of different sensing technologies. A key benefit is their ability to capture orthogonal biochemical and physiological signals, creating a multidimensional data layer that enriches diagnostic insights. For example, Al-Rawhani et al. reported a multimodal integrated sensor platform which combines optical and electrochemical sensing modalities to achieve comprehensive biomarker analysis[157]. This dual-mode system integrates optical detection (e.g., surface-enhanced spectroscopy) with electrochemical transduction, enabling simultaneous quantification of multiple analytes in complex biological samples. Such integration allows for cross-validation of results. Optical signals provide high specificity for molecular recognition events while electrochemical readouts offer quantitative precision and real-time response. This addresses the limitations of single-mode systems that would otherwise capture only partial biochemical information. Similarly, Zhao et al. developed a multimodal sensing platform integrating surface-enhanced Raman scattering (SERS) and colorimetric sensing for sweat biomarker detection[158]. The SERS modality provided high specificity for molecular recognition events, while the colorimetric readout offered rapid qualitative validation. This integration effectively reduced false positives caused by non-specific adsorption in complex sweat matrices, achieving lower detection limits (down to micromolar levels for glucose and lactate) compared to either single modality alone. Such results demonstrate how combining optical sensing techniques can synergistically enhance overall detection performance, highlighting the advantage of multimodal designs in addressing the limitations of individual sensing methods. These examples demonstrate how multimodal architectures mitigate the inherent limitations of individual techniques. Such limitations include background interference in optical sensing and limited specificity in single-mode detection. By synergizing complementary strengths, multimodal architectures enhance overall detection performance.
Another advantage is the enhanced robustness in dynamic biological environments, where temperature fluctuations, pH shifts, and matrix effects can perturb single-mode sensor readings. Multimodal platforms address this by incorporating dedicated reference channels. A typical configuration might include a thermoresistive sensor to normalize electrochemical signals against temperature drifts, combined with a potentiometric pH probe to correct for environmental acidity[159]. This self-calibrating capability is particularly valuable for wearable devices, where skin temperature variations (32 to 36 °C) can alter enzyme activity in electrochemical sensors by up to 20% per °C. By contrast, single-mode devices often require frequent recalibration or suffer from cumulative drift in real-world use[160,161].
Moreover, multimodal sensing platforms overcome the limitations of single-analyte detection by enabling concurrent measurement of physiologically correlated biomarkers across multiple scales. For instance, the biosensing system develop by Wang et al. employs genetically encoded fluorescent probes to achieve spatiotemporally resolved lactate monitoring from subcellular compartments to whole-body dynamics in diabetic models, integrating responses to metabolic perturbations (glucose deprivation) and pharmacological interventions with ≤ 5-s temporal resolution[162]. This contrasts sharply with conventional single-mode setups that require separate devices for each analyte. In clinical diagnostics, such multimodal synergy enhances early disease detection through cross-validated biomarker networks. A representative implementation couples an ultrasensitive electrochemical aptasensor for cardiac troponin I with microwave Doppler-based heart rate variability (HRV) sensing using vector analytic demodulation (VAD) algorithms[163]. This dual-modality platform detects molecular-level myocardial injury (cTnI elevation) concurrently with autonomic dysfunction (HRV anomalies), achieving 98% accuracy in QRS complex identification under physiological noise[164]. Such integration delivers essential diagnostic context unattainable with single-mode devices, specifically by correlating nanoscale biomolecular events (e.g., cardiac troponin I fluctuations) with organ-level functional dynamics (e.g., autonomic regulation via heart rate variability) to discriminate benign biomarker variations from pathological states such as acute coronary syndromes[163,164].
While multimodal biosensing offers richer datasets, it is crucial to delineate when this complexity translates into tangible clinical benefits versus when single-mode approaches suffice. Multimodal strategies clearly outperform single-mode sensors in two specific scenarios: (1) Decoupling cross-sensitivity: In complex bio-fluids, a single sensor often responds to interfering species (e.g., ascorbic acid affecting glucose readings)[165]. A multimodal system can use a secondary modality (e.g., a selective polymer coating or an orthogonal transduction method) to calibrate and subtract these interferences in real-time[166]. (2) Correlated physiological profiling: For conditions like cardiovascular disease, measuring a single marker (e.g., ECG) provides incomplete risk assessment. Integrating biochemical markers (e.g., Troponin) with physical signals (e.g., ECG/blood pressure) creates a closed-loop diagnostic model that single-mode sensors cannot achieve[167].
However, the added complexity of multimodality is not justified when targeting highly specific, high-concentration analytes in controlled environments. For instance, in basic fitness tracking where only heart rate is required, adding biochemical sensing capabilities may introduce unnecessary power drain, data processing latency, and fabrication costs without significantly improving the user experience. Therefore, the design of “sensor-computing” platforms must be driven by the specific clinical question rather than a desire for technological novelty.
Limitations for practical applications
Increased system complexity arises from integrating heterogeneous transduction mechanisms, leading to material compatibility conflicts and signal crosstalk. For instance, conductive electrode materials (e.g., gold, carbon nanotubes) essential for electrochemical sensors can absorb or scatter light, degrading optical sensing performance[168]. Similarly, microwave antennas coupled with enzymatic electrodes may induce localized heating that denatures biomolecules[169,170]. Such interference necessitates advanced packaging strategies like microfluidic channel isolation or frequency-domain filtering[171,172], which can decrease manufacturing complexity and failure risks. Empirical studies on wearable sensors confirm that inter-modality interference is a primary contributor to reliability degradation, though specific failure rate comparisons require further clinical validation[173,174].
Enhanced power consumption and form factor constraints further limit practicality. Each modality imposes distinct power demands: optical systems require light sources, electrochemical modules need potentiostats, and impedance analyzers rely on high-frequency signal processors. Cumulatively, these increase energy consumption by 3 to 5 times compared to single-mode devices, straining battery life in wearables[175,176].
Data processing and analysis burdens scale with multimodal data heterogeneity. Platforms combining voltammetric, potentiometric, and optical outputs generate time-series, frequency-domain, and intensity datasets requiring synchronization and joint analysis. This demands computationally intensive machine learning models (e.g., convolutional neural networks (CNNs) for spatial patterns, long short-term memory networks (LSTMs) for temporal correlations)[177-179], yet unoptimized fusion can degrade accuracy. Effective analysis thus hinges on adaptive algorithms like physiologically inspired spatiotemporal fusion models to resolve signal conflicts[180].
MINIATURIZED MANUFACTURING, ALGORITHMIC PROCESSING, AND APPLICATION PRACTICES OF MULTIMODAL BIOSENSOR SYSTEMS
Fabrication strategies for multimodal biosensor systems
The advancement of miniaturized and integrated biosensors has been driven by a wide range of printing and fabrication techniques. Among these, methods such as inkjet printing, extrusion-based printing, laser-assisted bioprinting, stereolithography, volumetric printing, and microcontact printing each offer unique trade-offs in resolution, material compatibility, mechanical integrity, and fabrication complexity. Crucially, these manufacturing techniques serve as the physical enabler for Level 2: on-device intelligence. By permitting the co-fabrication of sensing elements with conductive interconnects and flexible circuit substrates, advanced printing strategies facilitate the seamless integration of biological interfaces with edge-computing hardware, ensuring robust signal transmission for downstream processing. For example, inkjet printing excels at high-resolution droplet deposition but requires low-viscosity inks, whereas extrusion techniques accommodate thicker bioinks albeit at lower resolution. Laser-assisted methods offer nozzle-free precision, though with higher cost and setup complexity, while light-based techniques like stereolithography and volumetric printing enable rapid structuring but demand specialized photopolymers. Microcontact printing provides high spatial accuracy and minimal waste for planar sensor interfaces. Table 3 summarizes these methods’ relative advantages and limitations[181]. This comparative overview guides the selection of appropriate fabrication strategies, especially when balancing electrical performance, mechanical compliance, and biocompatibility in wearable or implantable biosensing platforms.
Comparison of different bioprinting techniques
| Printing technique | Advantages | Disadvantages | Key use cases |
| Inkjet | High printing speed, Cost-effective, suitable for high-throughput production, non-contact printing method | Limited viscosity of bioinks, risk of nozzle clogging, low cell density in printed constructs, droplet size variability | Tissue engineering, high-throughput screening |
| Extrusion | Versatile bioink use, high cell density achievable, suitable for complex structures, continuous filament deposition | Poor resolution, shear stress can damage cells, slower than inkjet printing, potential for bioink leakage | 3D tissue scaffolds, vascular structures |
| Laser-based | High resolution, precision control over biomaterial deposition, minimal cell damage | High operational costs, limited material compatibility, slow printing speed | High-resolution tissue structures, cancer research |
| Volumetric | Very high printing speed, ability to print large constructs in a single step, isotropic mechanical properties | Requires specialized equipment, lower resolution compared to other methods possible light exposure issues | Large-scale tissue printing, bone scaffolds |
| Stereolithography | High resolution and surface finish, suitable for detailed structures, excellent control over polymerization | Limited to photosensitive bioinks, potential cytotoxicity, requires post-processing times | Highly detailed tissue structures, dental implants |
| Digital light projection | High resolution and accuracy, rapid printing speed, excellent control over polymerization, suitable for complex geometries | Limited to photosensitive bioinks, potential photoinitiator toxicity, high equipment costs | Complex 3D printing, precision organ printing |
| Layer-by-layer | High resolution, precision control over deposition, good for multi-materials | Slow process, requires post-processing, material limitations | Multimaterial tissue printing, biofunctional layers |
| Roll-to-roll | Scalable for large-area production, cost-effective for high-volume manufacturing, high throughput | Limited resolution, requires flexible bioinks, lower precision for microstructures | Large-area bioprinting, flexible electronics |
To further optimize for the “Level 2” on-device intelligence described in our architecture, fabrication strategies must evolve from simply patterning sensing elements to enabling the co-fabrication of heterogeneous components on a single substrate. Among the discussed techniques, inkjet and screen printing stand out as the most suitable candidates for this co-integration due to their material versatility. Inkjet printing is particularly advantageous for defining high-resolution interconnects required for interfacing with high-density flexible ICs, allowing for the precise deposition of functional inks alongside conductive silver tracks without mask changes. Conversely, screen printing offers superior throughput and thicker film deposition, making it the preferred choice for robust, low-impedance power lines needed to drive edge-computing microcontrollers.
Consequently, the choice of fabrication method directly dictates the integration strategy for computing elements. For systems requiring powerful processing (e.g., TinyML inference), flexible hybrid electronics has emerged as the dominant approach. This strategy transcends simple printing by incorporating industrial pick-and-place assembly or novel transfer printing techniques to embed rigid, thinned silicon chips onto flexible substrates. Here, printed conductive traces act as the flexible “interposer”, while advanced interconnect materials like anisotropic conductive films (ACFs) ensure reliable electrical contact between the soft sensing layers and the hard processors. In contrast, emerging fully printed electronics utilize materials such as organic electrochemical transistors (OECTs) or carbon nanotubes to offer a monolithic alternative for simpler tasks. However, their current switching speeds restrict their utility for complex algorithmic execution compared to hybrid silicon architectures. Ultimately, the ultimate objective of scalable manufacturing lies in high-throughput production lines [e.g., Roll-to-roll (R2R) processes] that can simultaneously pattern high-fidelity sensing interfaces (Level 1) and reliably integrate edge processors (Level 2), thereby bridging the gap between laboratory prototypes and commercial “sensor-computing” patches.
Printing techniques for biosensor fabrication
Printing technologies have emerged as versatile approaches for manufacturing flexible and integrated biosensor platforms, offering high spatial resolution, compatibility with diverse functional materials, and scalability for large-area production. Among these, inkjet printing, extrusion-based printing, and laser-assisted printing represent three complementary modalities, each with unique operational principles and advantages.
Inkjet printing is a key technique for creating flexible biosensor components because it enables non-contact, mask-free patterning with resolutions down to 20 to 50 µm and accommodates a variety of bioink formulations[182]. The process proceeds in three stages: the generation of droplets by thermal or piezoelectric actuation, the interaction of those droplets with the substrate surface, and the evaporation or curing of solvent to fix the patterned material[183]. Two main operating modes are used in biosensor fabrication. In continuous inkjet printers a high-pressure pump and electric field create a steady stream of droplets that may be recirculated if not deposited, which can raise contamination risks. In drop on demand systems a voltage pulse applied to a piezoelectric actuator induces a pressure wave that ejects a droplet whose volume matches the nozzle diameter. Drop on demand avoids ink recirculation, supports lower-viscosity inks, and reduces thermal stress on biomolecules[184]. This mode is particularly advantageous for integrating multiple sensitive biological or chemical components on flexible substrates, and has been widely employed in the development of next-generation wearable and photonic devices. For instance, a fully inkjet-printed multiplexed sweat sensor on flexible polyethylene terephthalate (PET) has been reported [Figure 7A-C], in which silver ink forms current collectors and sequentially printed functional inks construct enzymatic layers, mediators, pH sensing elements, and temperature sensing traces[172]. Integration with flexible circuit modules enables continuous sweat analysis with high sensitivity, low drift, and mechanical robustness.
Figure 7. Illustration of different printing-based fabrication strategies. (A-C) Diagram and photograph of a fully printed wearable sweat sensor array on PET, featuring integrated electrodes for pH, glucose, alcohol, temperature, and reference measurements. Reproduced with permission[172]. Copyright 2024, Wiley; (D) Schematic of a multi-material extrusion bioprinter, showing motorized sled for automated syringe selection and nozzle rinsing; (E) Build plate placement into bioink droplet forms a controlled filament layer; (D and E) Reproduced with permission[186]. Copyright 2021, Springer; (F) Workflow for the laser-printed paper-based analytical device: digital pattern design, laser printing and thermal baking to form fluidic barriers, surface functionalization with silver and gold nanoparticles for atrazine recognition, sample application, and smartphone-based colorimetric readout. Reproduced with permission[196]. Copyright 2023, American Chemical Society. PET: Polyethylene terephthalate; UV: ultraviolent; μPAD: microfluidic paper-based analytical device; NP: nanoparticle.
Extrusion printing is one of the most versatile approaches for fabricating 3D biosensor arrays and tissue scaffolds because it supports the deposition of bioinks with a wide range of viscosities. These inks can include low-viscosity aqueous cell suspensions as well as highly concentrated, cell-laden hydrogels, all while maintaining fine spatial resolution at the micrometer scale[185]. In a typical setup [Figure 7D and E], bioink is loaded into a syringe or cartridge and extruded through a nozzle onto a substrate in a layer-by-layer manner[186]. This process is driven by pneumatic pressure, a piston mechanism, or a screw-based system. The resolution and fidelity of the printed structures are determined by several factors, including nozzle diameter, applied extrusion force or pressure, the rheological properties of the bioink, and the movement speed of the printhead or substrate. Compared with droplet-based or light-assisted printing technologies, extrusion systems are capable of handling inks with viscosities ranging from approximately 30 millipascal-seconds to 107 millipascal-seconds, making them especially well suited for printing robust, cell-containing constructs[187].
A key advantage of extrusion printing is its compatibility with a wide variety of biomaterials and cell types, allowing for the fabrication of multimaterial and multicellular systems. However, the mechanical shear generated during the extrusion process can negatively affect cell viability. To address this, bioinks are often formulated to exhibit strong shear-thinning behavior, meaning that their viscosity decreases under shear stress, which helps reduce damage during printing. Additionally, these inks are designed to rapidly solidify or crosslink after deposition to maintain structural integrity and promote biological activity[188]. Commonly used formulations include natural and synthetic polymers such as alginate, gelatin methacrylate, and nanocellulose-based materials, which provide the necessary mechanical support and biocompatibility for various biomedical applications[189,190].
Laser-based bioprinting uses short, high-energy light pulses to deposit live-cell inks onto a target surface without any physical nozzle. A typical system comprises a pulsed laser, focusing optics, a donor ribbon (glass support, thin metal absorber, bioink layer) and a receiving substrate[191]. When the laser irradiates the metal layer, rapid vaporization creates a localized pressure bubble that ejects a picoliter droplet of bioink toward the substrate[192]. Because there is no nozzle, this method tolerates viscosities from 1 to 300 mPa·s without clogging and subjects cells to minimal mechanical stress, routinely achieving over 95% viability even at high cell densities[193,194]. Key factors controlling droplet size and placement include laser energy density, bioink film thickness and viscosity, and the spacing and surface chemistry of the receiver[195]. Figure 7F illustrates a dual-chromatic laser-printing strategy for fabricating a microfluidic paper-based analytical device[196]. One wavelength patterns hydrophobic barriers that define channel geometries, while a second wavelength deposits atrazine-sensing reagents into channel interiors. This example demonstrates how laser-based deposition can create complex, multimaterial sensor architectures in a seamless, mask-free process.
Overall, these three printing modalities offer complementary capabilities for biosensor fabrication: inkjet printing excels in high-resolution, mask-free deposition; extrusion printing enables multi-material and 3D structuring; and laser-assisted printing provides precision patterning without nozzle-related limitations.
Photopolymerization techniques for biosensor fabrication
Photopolymerization-based additive manufacturing technologies offer high-resolution, rapid prototyping capabilities that are well suited for creating complex, 3D biosensor architectures. Among these, volumetric printing, stereolithography, and digital light projection printing represent three powerful approaches that differ in curing principles, achievable geometries, and processing speeds.
Volumetric printing utilizes tomographic projection to polymerize the entire volume of photosensitive resin in one step. A series of two-dimensional light patterns, which are computed as the inverse Radon transform of the target 3D object, are projected simultaneously from multiple angles through a rotating resin bath[197]. The system typically employs a laser source, precision motion stages, and a real-time control module to coordinate light irradiation and resin rotation[198]. When the cumulative light dose exceeds the threshold, the resin cross links to form the final structure within seconds, with sub millimeter precision and a fully continuous internal architecture[199]. Since the resin remains fluid until the final exposure, there is no layer by layer “stair stepping” effect, resulting in smooth surfaces and isotropic mechanical properties. The key to volumetric printing lies in the generation of material growth functions (MGFs). These are Gaussian-shaped polymerized footprints induced by laser irradiation that accumulate spatially to form complex geometries. However, volumetric printing currently only supports single material resins and requires photosensitive formulations with narrowly tuned absorption and gelation kinetics[200].
Stereolithography is an additive manufacturing method in which a focused beam of ultraviolet or visible light induces polymerization in a photosensitive resin, creating 3D parts layer by layer. In a typical setup, a laser source is directed by galvanometer mirrors through a transparent window into a vat of liquid prepolymer. Wherever the light contacts the resin, it triggers rapid crosslinking; after one layer is cured, the build platform lowers or raises to allow fresh resin to flow beneath, and the process repeats until the final structure is complete[201]. An alternative approach uses a micromirror array to project an entire layer pattern simultaneously, curing a whole plane of material in a single exposure and thereby reducing overall build time[202]. The lateral resolution is determined by the optics of the projection system and mirror pixel size, while the vertical resolution depends on the absorption and scattering properties of the uncured resin[203]. This technology achieves sub-10-micrometer precision and allows uncured material to be washed away cleanly. Building on the high precision and versatile material properties afforded by stereolithographic fabrication, Antonelli et al. integrated photopolymerized microstructures into a Lab-on-Chip platform featuring multispectral readout[204]. The system uses a patterned visible light projector to locally polymerize vertical micropillars within a transparent resin, while a secondary red-filtered illumination path enables real-time monitoring of pillar growth. This combined curing and imaging approach supports in situ assessment of feature formation and facilitates subsequent multispectral biochemical signal analysis through machine learning.
Digital light projection printing uses a dynamic micromirror array (a digital micromirror device, DMD) to project entire cross-sectional images of the 3D model onto the resin surface. This system relies on a programmable light source, a DMD chip for pattern generation, and a printing platform to control layer stacking[205]. Each exposure cures an entire layer (tens to hundreds of micrometers thick) in fractions of a second, after which the build platform moves to refresh the resin. Lateral resolution is determined by the projected pixel size of the DMD optics, while vertical resolution is governed by the optical penetration depth of the resin and the layer step size[206]. This approach combines true layer wise control, which enables easy switching between multiple materials, with high throughput, achieving feature sizes ranging from 5 to 300 µm and build speeds of up to tens of millimeters per hour[207].
For all three methods, material formulation plays a decisive role in performance. Resin viscosity, photoinitiator concentration, and the presence of photoabsorbers or dyes must be balanced to achieve rapid curing, adequate resolution, and minimal scattering. The development of biocompatible and cell-friendly resins, particularly those curable under visible light, is expanding the applicability of these technologies to integrated biosensors and tissue engineering scaffolds[208,209]. By combining the volumetric speed of tomographic projection, the fine resolution of stereolithography, and the material versatility of digital light projection, researchers can create sophisticated, fully functional biosensor platforms within minutes while maintaining microscale precision.
Layer-by-layer assembly for biosensor integration
Layer-by-layer (LbL) assembly provides a highly versatile and modular fabrication strategy, particularly suitable for constructing flexible and functional sensor interfaces. This technique is based on the sequential adsorption of complementary materials, most commonly oppositely charged polyelectrolytes, onto a substrate. After each adsorption step, a rinsing process is performed to remove unbound species and ensure the precision of the deposited layer[210]. LbL assembly can be implemented through three main approaches[211]. In the immersion (or dipping) method, the substrate is alternately immersed into solutions containing the desired materials, with rinsing between immersions to maintain purity. The spraying method employs atomized droplets of each solution to coat the surface sequentially, allowing rapid and uniform deposition over large or non-planar substrates. The microfluidic method channels the solutions over the substrate within confined flow paths, enabling localized patterning, precise control of adsorption time, and efficient reagent use.
The primary advantage of LbL assembly lies in its ability to precisely control film thickness, composition, and biofunctionalization at the nanometer scale, thereby enabling the incorporation of polymers, proteins, nanoparticles, or enzymes into tailored sensing layers. Furthermore, because it typically employs aqueous processing conditions, the method is compatible with preserving biomolecular activity, a critical consideration in biosensor integration. These attributes make LbL assembly a powerful complement to printing technologies, especially in applications requiring fine-tuned surface chemistry, hierarchical architectures, or controlled release of recognition elements in wearable or implantable biosensors[212].
Roll-to-roll manufacturing for scalable biosensor production
R2R fabrication is a continuous manufacturing technique in which flexible substrates are transferred between supply and take-up reels while passing through a sequence of processing stations. This method supports large-scale and cost-efficient production of flexible biosensors by integrating multiple fabrication steps such as patterning, functionalization, lamination, and cutting into one automated workflow[213]. The substrate, typically a polymer film, moves through a series of rollers where each station performs a specific process in-line. Patterning can be accomplished by hot embossing, photolithography, or nanoimprinting. Functionalization steps, including reagent deposition or thin-film coating, can be implemented before subsequent lamination or encapsulation. Because the substrate remains in continuous motion, R2R systems can reach production speeds of tens to hundreds of meters per hour, making the approach suitable for high-volume manufacturing of disposable biosensors.
In biosensing applications, R2R technology has been successfully applied to both electrochemical and microfluidic device fabrication[214]. As shown in Figure 8A and B, R2R gravure printing enables the scalable production of electrochemical biosensors for wearable and medical devices[215]. This process utilizes rotating cylinders to continuously deposit functional inks onto the flexible substrate. By printing multiple layers sequentially, including silver tracks, carbon working electrodes, and dielectric insulation layers, the system achieves high-throughput fabrication of uniform electrode arrays. This layer-by-layer printing strategy demonstrates the feasibility of manufacturing large-area and low-cost sensors with the precision required for medical diagnostics. Furthermore, as illustrated in Figure 8C, R2R systems can also incorporate microfluidic structures, sensing electrodes, and optical components directly onto flexible polymer substrates[216]. Hot embossing can define microchannels, die-cutting can create fluidic inlets and outlets, and automated dispensing can deposit biorecognition elements. After these steps, lamination seals the microfluidic network, and auxiliary components such as membranes or reservoirs can be added during the same production run. The integration of these processes reduces manual assembly, minimizes variability, and enables scalable manufacturing suitable for clinical diagnostics and environmental monitoring.
Figure 8. Representative fabrication strategies and device architectures for R2R printed and integrated microfluidic biosensors. (A) Schematic of R2R gravure printing for electrochemical sensors, showing the layer-by-layer deposition of functional inks to form electrode arrays on a flexible substrate; (B) Image of the R2R gravure printing system. (A and B) Reproduced with permission[215]. Copyright 2018, American Chemical Society; (C) Illustration of the R2R hot embossing process for the continuous patterning of microfluidic channels on polymer films. Reproduced with permission[216]. Copyright 2020, American Chemical Society. PET: Polyethylene terephthalate; R2R: roll-to-roll.
Signal processing and ai algorithms of multimodal biosensor systems
While section 2 focused on the physical sensing layer (Level 1), this section addresses the computational intelligence required to interpret these signals. AI in biosensing operates across two distinct tiers of our integration architecture: Level 2 (on-device intelligence): utilizing lightweight algorithms (e.g., TinyML) embedded on wearable microcontrollers to perform real-time noise reduction, feature extraction, and rapid inference at the network edge. Level 3 (system-level fusion): employing advanced deep learning models [e.g., Transformers, Large Language Models (LLMs)] to integrate multimodal data streams, enabling comprehensive health profiling and predictive analytics.
As illustrated in Figure 9, diverse AI architectures are strategically deployed across this sensor-computing hierarchy based on their computational complexity and inference requirements. Lightweight supervised learning models, such as linear regression and support vector machines (SVMs), are primarily deployed at Level 2. They enable rapid, low-power classification (e.g., risk: yes/no) and continuous parameter estimation (e.g., glucose concentration) directly on the microcontroller, minimizing data transmission costs[217,218]. Conversely, computationally intensive unsupervised learning and deep learning frameworks are typically offloaded to Level 3 (cloud/server). These models handle high-dimensional tasks such as clustering patient cohorts based on long-term glycemic trends or generating predictive health insights via LLMs[219]. Furthermore, reinforcement learning can bridge these levels by enabling adaptive sensing strategies, such as dynamically optimizing sampling rates based on real-time physiological feedback to balance power consumption and data fidelity.
Figure 9. A Signal-processing and AI methods for multimodal biosensor systems: statistics, machine learning, deep learning, reinforcement learning, and foundation models. LLM: Large Language Model; AI: artificial intelligence; SVM: support vector machine; PCA: principal component analysis; UMAP: Uniform Manifold Approximation and Projection; CNN: convolutional neural network; RNN: Recurrent Neural Network; LSTM: long short-term memory network.
Multimodal data fusion strategies: glucose monitoring as a paradigm
To illustrate the practical implementation of AI architectures, we focus on non-invasive glucose monitoring as a representative paradigm[220-223]. Glucose detection exemplifies the need for multimodal data fusion to overcome the limitations of single-analyte sensing. Effective fusion strategies are pivotal for handling heterogeneous data and are generally categorized into three levels: Data-Level (Early) Fusion for raw signal concatenation; Feature-Level (Intermediate) Fusion for merging extracted attributes; and Decision-Level (Late) Fusion for aggregating independent model predictions.
In this context, signal processing techniques are essential to extract meaningful data from raw sensor signals before fusion. For Level 2 on-device processing, techniques like wavelet packet decomposition are used to decompose signals into different frequency sub-bands[224], while empirical mode decomposition to capture nonlinear and non-stationary components[225,226]. Additionally, spectral filtering to suppress noise and physiological artifacts so that weak analyte signatures become measurable. The general process of sensing and classification involves several key stages, including data collection, acquisition and preprocessing, feature engineering, fusion and modeling, training, and finally evaluation analysis. During feature engineering (Intermediate Fusion), researchers extract spectral and temporal features such as peaks, trends, and learned embeddings with ML models including autoencoders and CNNs. Principal component analysis (PCA) is often employed to reduce redundancy and maintain interpretability. In the final stage (Level 3 Prediction), multimodal fusion integrates heterogeneous sources and ML models such as XGBoost, LSTM, and graph neural networks are used for prediction or classification, with cross-validation and bootstrapping ensuring generalizable performance.
In addition to classification, regression algorithms are used to predict continuous glucose concentrations. Representative examples include gaussian process regression for aqueous benchmarks[227], and random forest or compact CNNs for estimating fasting blood glucose from real samples[228]. These models, integrated with microstrip microwave sensors, allow for accurate predictions of glucose concentrations by capturing spectral features and repeatability behaviors. Furthermore, deep learning models such as LSTM networks are increasingly used to forecast future values based on past sensor data. Bayesian optimization fine-tunes hyperparameters like lookback windows and learning rates to optimize model performance[229].
Critical challenges and validation analysis
While AI-enabled biosensing shows immense promise, rigorous validation and critical analysis of current limitations are essential for clinical translation. Evaluation metrics for glucose prediction and classification in diabetes care using various AI methods include root mean square error (RMSE), mean absolute error (MAE), and mean absolute percentage error (MAPE) for regression tasks, with accuracy, precision, and AUC-ROC used for classification tasks[228,227,230,231]. Supplementary Table 1 [Supplementary Materials] summarizes various AI-powered glucose sensors utilizing microwave, optical, and electrochemical technologies to enhance diabetes monitoring. As shown in Supplementary Table 1, most mentioned studies formulated glucose estimation as a regression task because blood glucose is inherently a continuous variable. Results showed high accuracy in controlled settings, with some models achieving over 99% classification accuracy or strong regression performance.
However, a critical analysis reveals significant gaps remaining for effective implementation. As noted in Supplementary Table 1, most experiments were limited by small sample sizes or synthetic datasets, leading to a severe domain shift problem where models trained on controlled lab data often fail in real-world scenarios due to user variability and environmental noise (e.g., temperature and humidity)[232,233]. Consequently, robustness to motion artifacts remains a primary hurdle that necessitates hardware-aware algorithms at Level 2. Beyond signal fidelity, the “black-box” nature of deep learning models (e.g., CNNs, LSTMs) raises concerns regarding clinical trust. Although techniques like shapley additive explanations help reduce redundancy, developing fully explainable AI is required to visualize feature importance for clinicians to verify autonomous predictions. Finally, from a deployment perspective, transmitting high-resolution physiological data to the cloud exposes sensitive user information. Future systems must therefore adopt federated learning to train models locally on devices without sharing raw data, balancing personalized healthcare with rigorous privacy and security preservation.
Application demonstrations of multimodal biosensor systems
Moving beyond isolated device demonstrations, the successful deployment of sensor-computing systems relies on specific system-level design principles that dictate how modalities are combined. Our analysis of recent literature reveals three overarching strategies for selecting sensing combinations: (1) Orthogonal Verification (pairing distinct mechanisms to reduce false positives). (2) Contextual Decoupling (using physical sensors to subtract environmental noise). (3) Holistic Profiling (capturing correlated parameters for comprehensive diagnosis). The following case studies illustrate how these principles are implemented across diverse domains, ranging from clinical bioanalysis and food safety to continuous wearable health monitoring.
To illustrate the principle of Orthogonal Verification, dual-mode platforms that fuse complementary transduction mechanisms have proven highly effective in eliminating false positives. A representative example is shown in Figure 10A, which presents a chemical redox cycling-based biosensor for the self-powered photoelectrochemical (PEC) and colorimetric assay of heat shock protein (HSP)[234]. In this configuration, the detection relies on a redox cycling reaction triggered by the target analyte. This reaction simultaneously amplifies a PEC photocurrent (electrical signal) and induces a visible color shift (optical signal). The self-powered PEC mode offers high sensitivity for quantitative analysis, while the colorimetric mode provides an intuitive visual check. The consistency between these two distinct physical signals ensures that environmental noise affecting one channel (e.g., electrical drift) does not lead to a misdiagnosis, thereby achieving robust orthogonal verification.
Figure 10. Multimodal biosensing platforms employing orthogonal verification strategies. (A) Schematic illustration of a chemical redox cycling-based dual-mode biosensor for HSP, enabling self-powered PEC and colorimetric detection through a shared redox reaction. Reproduced with permission[234]. Copyright 2025, American Chemical Society; (B) Schematic of a sensitive competitive nanozyme-linked apta-sorbent assay for the dual-mode detection of OTA, utilizing aptamer-functionalized nanozymes to generate cross-validated signals in complex matrices. Reproduced with permission[235]. Copyright 2022, Royal Society of Chemistry. HSP: Heat shock protein; BSA: bovine serum albumin; ALP: alkaline phosphatase; AA: ascorbic acid; DHA: dehydroascorbic acid; TCEP: tris(2-carboxyethyl)phosphine; AAP: L-ascorbic acid 2-phosphate; PEC: photoelectrochemical; ITO: Indium Tin Oxide; NP: nanoparticle; MNP: magnetic nanoparticle; OTA: ochratoxin A; SERS: surface-enhanced Raman scattering; TMB: 3,3',5,5'-tetramethylbenzidine.
Expanding on this verification strategy in food safety, Figure 10B displays a sensitive competitive nanozyme-linked apta-sorbent assay for the dual-mode detection of ochratoxin A (OTA). Unlike direct binding assays, this platform employs a competitive mechanism where the target OTA competes with immobilized analogs for binding to aptamer-functionalized nanozymes[235]. The retained nanozymes utilize their intrinsic catalytic activity to generate dual-mode signals (e.g., colorimetric and fluorescence/optical) proportional to the target concentration. By cross-validating the signal trends from two independent readout channels, this method significantly enhances detection reliability in complex food matrices, effectively filtering out interferences that might affect a single sensing mode.
In parallel with hardware innovations, the integration of advanced computing units such as smartphones and machine learning algorithms has become a critical driver for robust orthogonal verification. For instance, Suo et al. demonstrated a multimodal platform for aflatoxin B1 (AFB1) detection that integrated electrochemistry, a commercial glucosemeter, and smartphone-based colorimetry[236]. In this system, the smartphone functioned as a core computing unit that captured colorimetric changes via its camera while simultaneously processing electrochemical data and glucose readings. This sensor-computing integration allowed for the on-site cross-validation of three distinct signal sources, which significantly reduced false positives in complex agricultural samples without relying on bulky instrumentation. Similarly, the value of data-driven enhancement in multimodal systems was highlighted by Kong et al., who developed a novel Nb2C MXene-based aptasensor for AFB1[237]. By combining electrochemical impedance spectroscopy (EIS) with PEC sensing, the platform achieved dual-mode signal acquisition. Crucially, rather than analyzing these signals in isolation, the study integrated an artificial neural network (ANN)-based regression model to fuse the non-linear outputs from both channels. This approach demonstrates how intelligent algorithms can extract correlated features from multimodal data to achieve superior detection limits and signal stability that single-mode sensing could not attain.
Transitioning to wearable health monitoring, the principle of Holistic Profiling is exemplified by the stretchable wireless bioelectronic system designed for the multiplexed management of infected chronic wounds, as illustrated in Figure 11A-D. Unlike single-analyte sensors that offer limited diagnostic insight, this platform captures a comprehensive set of physiological parameters to guide therapeutic intervention. As depicted in the layer assembly [Figure 11A], the device utilizes a soft and stretchable poly[styrene-b-(ethylene-co-butylene)-b-styrene] (SEBS) substrate to house a custom-engineered electrochemical sensor array alongside an anti-inflammatory and antimicrobial drug-loaded electroactive hydrogel. The specific sensor layout, detailed in Figure 11B, incorporates electrodes for monitoring temperature, pH, ammonium (NH4+), glucose, lactate, and UA. This multiplexed capability allows the system to correlate metabolic biomarkers with inflammatory indicators in real time. For instance, the simultaneous elevation of pH and UA can signal a distinct infection state that temperature alone might miss. Physically, the device features a compact, fingertip-sized form factor [Figure 11C and D] that maintains high flexibility and stretchability. This mechanical compliance ensures stable contact with irregular wound surfaces, such as those found on diabetic feet. By fusing these diverse data streams, the system not only diagnoses the wound status but also enables a closed-loop response, using voltage-modulated electrodes to trigger controlled drug release or electrical stimulation for accelerated healing.
Figure 11. Representative wearable biosensing platforms for epidermal analysis. (A) Schematic of a stretchable bioelectronic patch applied to a diabetic foot, showing the SEBS substrate and integrated drug-loaded hydrogel layers; (B) Layout of the multiplexed sensor array for the simultaneous detection of temperature, pH, NH4+, glucose, lactate, and UA; (C and D) Optical photographs demonstrating the flexibility and stretchability of the fingertip-sized wearable patch; (A-D) Reproduce with permission[6]. Copyright 2023, AAAS; (E-G) Fabrication, sensing mechanism, and electrode structure of the β-Bi2O3 nanoflake textile sensor; (H and I) Robustness and biocompatibility assessments, including washability and skin irritation tests; (J) In situ sweat monitoring during exercise. (E-J) Reproduced with permission[238]. Copyright 2024, Cell Press. SEBS: Poly[styrene-b-(ethylene-co-butylene)-b-styrene]; UA: uric acid; PET: polyethylene terephthalate; PAMD:
While the specialized patch described above addresses clinical wound management, integrating biosensors into daily garments requires overcoming a different set of challenges, particularly mechanical durability and washability. As shown in Figure 11E and F, Shi et al. developed a washable textile biosensor that leverages nanostructured oxides to solve this robustness bottleneck[238]. The core mechanism relies on nanostructured oxide films deposited onto textile substrates. These oxides facilitate fast ion diffusion and efficient charge transport across the textile-electrolyte interface. Biorecognition elements immobilized on this layer trigger redox reactions proportional to analyte concentration, which are amplified by the oxides’ high conductivity. Figure 11G and H demonstrate the critical advantage of this design: even after 50 machine washing cycles, the nanostructured oxide layer remains structurally intact, and the biosensor retains over 90% of its initial performance. This durability enables the device to be seamlessly integrated into wristbands for the real-time monitoring of sweat analytes during physical activity [Figure 11I and J]. From a system design perspective, this work exemplifies the strategy of signal robustness in weal-world conditions, ensuring that multimodal data streams remain reliable despite the rigorous mechanical agitation and moisture exposure inherent to daily wear.
Complementing these textile and solid-state patch approaches, flexible microfluidics offer another effective pathway for bio-fluid management in multimodal systems. For instance, Reza et al. developed a flexible biosensing patch integrated with microfluidic channels for chronic wound monitoring[6]. Unlike the open-surface contact in textile sensors, this device employs a microfluidic network to actively guide wound exudate toward a multiplexed electrochemical array. The system continuously tracks metabolites, ions, pH, and temperature, transmitting data wirelessly to a mobile unit. By correlating metabolite levels with temperature changes, this platform provides a holistic profile that can distinguish between active infection and normal inflammation. This example further highlights how diverse structural integrations, ranging from washable fibers to microfluidic channels, can be tailored to meet specific physiological monitoring needs.
While the aforementioned examples highlight the success of flexible and portable form factors in clinical analysis and surface-level physiological monitoring, the paradigm of sensor-computing integration is equally transformative in high-precision and active sensing domains. Expanding beyond soft electronics, advanced multimodal strategies have emerged in silicon-based complementary metal oxide semiconductor (CMOS) integrated circuits and bio-inspired robotic platforms. These technologies address fundamentally different constraints, shifting the focus from mechanical conformability to extreme miniaturization and in vivo accessibility, thereby enabling precision biomolecular analysis and deep-tissue diagnostics.
A representative microwave-optical dual-modality platform in 28 nm bulk CMOS co-integrates a 40 GHz quadrature VCO-based permittivity sensor with on-chip single-photon avalanche diodes (SPADs) to simultaneously acquire dielectric and optical signals[239]. Using superharmonic-coupled quadrature voltage-controlled oscillators (QVCOs) with differential varactor modulation and injection locking, the microwave path moves the measurement beyond the 1/f3 corner and achieves a thermal-noise-limited frequency sensitivity of 0.2 ppm/√Hz (≈ 0.02 aF/√Hz in Phosphate Buffered Saline at ~ 42 GHz), while decoupling dynamic range from lock range (> 115 dB) through equal exposure of coupled tanks to the biomedium. The SPAD frontend provides photon counting and fluorescence lifetime measurements with ~ 300 ps timing jitter and ~ 11.5% photon detection efficiency at 520 nm (500/s dark count at 1 V excess bias), allowing time-gated optical readout without external filters. In protein denaturation experiments, the dielectric channel resolved a 444.78 ppm frequency shift for BSA unfolding, whereas the optical channel captured the loss of bioluminescent activity in luciferase at concentrations too low for dielectric detection alone; multimodal feature fusion produced four separable clusters [native/denatured Bovine serum albumin (BSA) / luciferase (LUC)], demonstrating enhanced selectivity that neither modality achieves independently. Crucially, this integration achieves orthogonal verification at the molecular scale: the optical channel compensates for the dielectric sensor’s Limit of Detection limits in low-concentration scenarios, while the microwave sensor captures structural conformational changes invisible to optics. This mutual compensation enables a comprehensive molecular fingerprinting capability that neither modality achieves independently.
In a distinct route to multimodal integration, bubble-inspired magnetic microrobots implement gas-responsive biosensing with co-embedded imaging and actuation[240]. These “armored” microbubbles (≈ 100-800 µm) formed by hydrophobic Fe-Ni particles at the air-liquid interface support wireless control via weak rotating or gradient magnetic fields while preserving the bubble’s intrinsic responsiveness. The air core acts as a chemical transducer: CO2 absorption in alkaline media drives size/shape changes whose rates correlate with local pH, and magnetic control can gate bubble exposure to separate navigation from sensing in complex fluids. The air-filled architecture yields strong ultrasound contrast, enabling real-time visualization and flow-stopped positioning of ~ 100 µm microrobots in mouse tail veins. From a system perspective, this represents active sensing via actuation-sensing decoupling. The magnetic modality handles the “context” (navigation and positioning), allowing the gas-responsive modality to perform targeted sensing in deep-tissue settings. This synergy solves the “access problem”, enabling precise diagnostics in anatomical regions where conventional static sensors simply cannot reach.
These diverse examples further broaden the scope and demonstrate the remarkable versatility of multimodal biosensor systems, illustrating how sensor-computing integration can be realized across fundamentally different platforms, from silicon-based monolithic integration to bio-inspired robotic architectures, while addressing distinct sensing challenges spanning molecular-scale biomolecular interactions to macroscale physiological monitoring, thereby establishing multimodal approaches as a universally applicable paradigm for next-generation intelligent biosensing systems.
CHALLENGES AND FUTURE PERSPECTIVES
Integrated multimodal biosensing platforms have advanced rapidly, offering significant potential for applications in health monitoring, food safety, and clinical diagnostics. However, translating laboratory prototypes into practical, large-scale systems faces persistent challenges. As the field progresses, and as illustrated in the strategic roadmap in Figure 12, we categorize these hurdles into three critical domains: materials and device interfaces, data and algorithmic intelligence, and system translation.
Figure 12. Strategic roadmap for addressing critical challenges in multimodal biosensing. (A) Material & Device Interfaces: schematic illustrating the physical integration of functional layers to minimize signal interference and biofouling; (B) Data Complexity & Intelligence: visualization of AI-driven strategies to process high-dimensional heterogeneous data for precise decision-making; (C) Clinical Translation Ecosystem: overview of the practical barriers to adoption, highlighting user variability, regulatory pathways, and calibration requirements. AI: Artificial intelligence.
Materials and device interfaces
A key challenge lies in the integration of different sensing modalities, which often involve incompatible materials or signal interference[241]. As depicted in Figure 12A, physical integration requires stacking distinct functional layers (e.g., optical films over conductive meshes) without compromising signal integrity. For example, conductive layers required for electrochemical detection can absorb or scatter light, disrupting optical signals. Similarly, microwave components may produce localized heating that degrades biological recognition elements. Mitigating such effects requires additional engineering strategies, including microfluidic isolation or frequency filtering, which increase system complexity and potential failure points[242]. Furthermore, in complex biological fluids, sensor performance is frequently compromised by biofouling, characterized by the accumulation of proteins and cells on sensor surfaces, which causes signal drift and limits long-term reliability[243].
To overcome these hardware limitations, future research should prioritize (1) Optically Transparent Conductors: Developing high-performance transparent electrodes (e.g., graphene meshes or silver nanowire networks) that enable electrochemical sensing without obstructing optical pathways. (2) Active Interface Engineering: Advancing zwitterionic or hydrogel-based anti-fouling coatings that are compatible with multiple transduction mechanisms, ensuring that the integration of a second modality does not compromise the lifespan of the first.
Data complexity and algorithmic intelligence
Multimodal systems generate high-dimensional datasets that are difficult to manage in real time[244,245]. This “curse of dimensionality” can lead to unobserved feature combinations, which limit the generalizability of ML models[218]. In addition, the lack of standardized data formats across sensing platforms complicates integration and analysis. Effective use of these systems depends on access to large volumes of high-quality data[246]. However, data collection remains difficult due to heterogeneity in data types, including structured signals, unstructured imaging data, and dynamic time-series[247].
The AI model offers parallel breakthroughs in data processing. As shown in Figure 12B, advanced algorithms act as a fusion engine to filter noise and align heterogeneous inputs, ranging from chaotic waveforms to unstructured point clouds, into a coherent decision stream. Emerging multimodal learning architectures such as Transformer-based models and Perceiver IO allow for unified processing of varied inputs, including physiological signals, clinical images, and text[248]. Yet, training these models requires large annotated datasets, which are often scarce in biosensing.
Addressing data challenges requires focusing on (1) Hardware-Aware Neural Architecture Search (NAS): Automatically designing lightweight AI models that fit within the power and memory constraints of on-body chips (Level 2) without sacrificing multimodal accuracy. (2) Federated Learning Frameworks: Implementing decentralized training protocols that allow models to learn from diverse user populations without transmitting raw sensitive data to the cloud, thereby solving the data scarcity problem while preserving privacy.
System translation and clinical ecosystem
Beyond technical feasibility, scalability is limited by issues of durability, cost, and regulatory barriers. Incorporating multiple sensing modules raises manufacturing expenses, which may restrict deployment in low-resource environments[249]. Furthermore, user adherence is hampered by frequent calibration requirements (e.g., finger-prick tests). Compounding this challenge is user-dependent variability, including differences in skin composition and motion artifacts, which can severely impact model generalization[250]. These interrelated barriers form a complex clinical ecosystem, as highlighted in Figure 12C, which demands coordination between user variability, regulatory safety, and cost-effective calibration. This underscores the need for adaptive AI frameworks trained on diverse, standardized datasets rather than small, homogeneous cohorts.
Navigating regulatory pathways, such as those of the U.S. Food and Drug Administration (FDA) or Conformité Européenne (CE) marking, remains inherently complex for multimodal systems. Under frameworks like Software as a Medical Device (SaMD), demonstrating that the failure of one sensing mode does not compromise patient safety is a pivotal validation challenge. Nevertheless, the integration of ecosystem-level data, which combines wearable sensors with omics data such as genomics and metabolomics, remains the ultimate goal for precision medicine, as outlined in Figure 12B[248].
To bridge the gap to clinical adoption, efforts must target (1) Self-Calibrating Architectures: Leveraging stable physical sensors (e.g., impedance or pressure) to act as internal references that automatically correct the drift of biochemical sensors, reducing user intervention. (2) Standardized Validation Protocols: Establishing industry-wide benchmarks for “Multimodal SaMD” that explicitly quantify the safety margins and cross-validation benefits (e.g., using optical data to double-check electrochemical alerts) to accelerate FDA/CE approval.
CONCLUSION
The transition from passive data collection to active health intelligence relies fundamentally on the depth of sensor-computing integration. As highlighted throughout this review, it requires not only rational design and integration of different functional components, but also requires intensive research efforts to tackle the challenges of on-sensor material engineering (Level 1), on-device algorithmic optimization (Level 2), and system-level multimodal fusion (Level 3). The future revolution of biosensing lies in the seamless coupling of the advances from these three aspects. Firstly, it is expected that enhancement on intrinsic signal quality can be achieved via novel biosensing micro/nanomaterials, thus the extracted signals can be directly processed by edge-AI algorithms for real-time decision-making. Meanwhile, high-level data fusion strategies can be utilized to unlock precise disease profiling from complex datasets. It is promising to realize intelligent and practical healthcare via hardware-aware AI and neuromorphic bio-interfaces that seamlessly bridge physical sensing with digital interpretation. Overcoming the remaining barriers on interfacial compatibility, power efficiency, and data standardization will be pivotal in translating these laboratory innovations into ubiquitous, autonomous health monitoring platforms.
DECLARATIONS
Authors’ contributions
Conceptualization: Guo, W. T.; Lin, Y.
Investigation: Guo, W. T.; Zahran, A.; El-Shal, I. H.; Guo, Y.; Eldamak A. R.; Fahmy, O. M.
Methodology: Guo, W. T.; Zahran, A.; El-Shal, I. H.; Guo, Y.
Writing - original draft: Guo, W. T.; Zahran, A.; El-Shal, I. H.; Guo, Y.
Supervision: Eldamak A. R.; Fahmy, O. M.; Elsheakh, D. N.; Yuan, Z.; Tai, H.; Lin, Y.
Project administration: Elsheakh, D. N.; Yuan, Z.; Tai, H.; Lin, Y.
Writing - review & editing: Elsheakh, D. N.; Yuan, Z.; Lin, Y.
Funding acquisition: Lin, Y.
Availability of data and materials
Not applicable.
AI and AI-assisted tools statement
Not applicable.
Financial support and sponsorship
This work was supported by the National Natural Science Foundation of China (62571225), Shenzhen Science and Technology Program (RCYX20231211090432060), the National Science Foundation for Distinguished Young Scholars (Grant No. 62225106), Natural Science Foundation of China (Grant No. U24A20229, 62471093). Open Foundation of State Key Laboratory of Electronic Thin Films and Integrated Devices (KFJJ202404).
Conflicts of interest
The authors declare there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
Supplementary Materials
REFERENCES
1. Meng, X.; Wang, J.; Diao, L.; Li, C. Construction of multi-mode photoelectrochemical immunoassays for accurate detection of cancer markers: assisted with MOF-confined plasmonic nanozyme. Anal. Chem. 2024, 96, 1336-44.
2. Lu, Y.; Ning, Y.; Li, B.; Liu, B. Dual-signal imaging mode based on fluorescence and electrochemiluminescence for ultrasensitive visualization of SARS-CoV-2 spike protein. Anal. Chem. 2024, 96, 463-70.
3. Kabay, G.; DeCastro, J.; Altay, A.; et al. Emerging biosensing technologies for the diagnostics of viral infectious diseases. Adv. Mater. 2022, 34, e2201085.
4. Lin, S.; Zheng, S.; Wu, H.; et al. Multi-analyte discriminated ECL via photonic crystal-enhanced selectivity. Anal. Chem. 2023, 95, 9116-22.
5. Dadras-Toussi, O.; Khorrami, M.; Louis Sam Titus, A. S. C.; Majd, S.; Mohan, C.; Abidian, M. R. Multiphoton lithography of organic semiconductor devices for 3D printing of flexible electronic circuits, biosensors, and bioelectronics. Adv. Mater. 2022, 34, e2200512.
6. Reza, M. S.; Sharifuzzaman, M.; Islam, Z.; et al. A flexible and multimodal biosensing patch integrated with microfluidics for chronic wound monitoring. Chem. Eng. J. 2024, 501, 157673.
7. Antiochia, R. Developments in biosensors for CoV detection and future trends. Biosens. Bioelectron. 2021, 173, 112777.
8. Fernandes, R. S.; Singh, R. K.; Goel, S.; Dey, N. Probe integrated with laser-induced graphene electrode: application towards multimodal optoelectrochemical detection of Cu(ii) ions and their remediation. J. Mater. Chem. C. 2024, 12, 15607-20.
9. Li, C.; Cheng, J.; He, Y.; et al. Polyelectrolyte elastomer-based ionotronic sensors with multi-mode sensing capabilities via multi-material 3D printing. Nat. Commun. 2023, 14, 4853.
10. Tian, J. H.; Hu, X. Y.; Hu, Z. Y.; et al. A facile way to construct sensor array library via supramolecular chemistry for discriminating complex systems. Nat. Commun. 2022, 13, 4293.
11. Ozer, T.; Henry, C. S. Review - recent advances in sensor arrays for the simultaneous electrochemical detection of multiple analytes. J. Electrochem. Soc. 2021, 168, 057507.
12. Vargas, E.; Teymourian, H.; Tehrani, F.; et al. Enzymatic/immunoassay dual-biomarker sensing chip: towards decentralized insulin/glucose detection. Angew. Chem. Int. Ed. Engl. 2019, 58, 6376-9.
13. Liu, Z.; Su, J.; Zhou, K.; Yu, B.; Lin, Y.; Li, K. H. Fully Integrated patch based on lamellar porous film assisted GaN optopairs for wireless intelligent respiratory monitoring. Nano Lett. 2023, 23, 10674-81.
14. Zhou, K.; Ding, R.; Ye, W.; et al. A fully printable and integrated system with long-term stability for versatile and multimodal perspiration tracking. Nano Lett. 2024, 24, 15092-100.
15. Shi, Y.; Zhang, Z.; Huang, Q.; Lin, Y.; Zheng, Z. Wearable sweat biosensors on textiles for health monitoring. J. Semicond. 2023, 44, 021601.
16. Gao, J.; Huang, W.; Chen, Z.; Yi, C.; Jiang, L. Simultaneous detection of glucose, uric acid and cholesterol using flexible microneedle electrode array-based biosensor and multi-channel portable electrochemical analyzer. Sens. Actuators B: Chem. 2019, 287, 102-10.
17. Ma, X.; Jiang, Z.; Lin, Y. Flexible energy storage devices for wearable bioelectronics. J. Semicond. 2021, 42, 101602.
18. Li, Z.; Lin, X.; He, D.; Ma, Y.; Lin, Y. Preface to special issue on novel semiconductor-biochemical sensors. J. Semicond. 2023, 44, 020101.
19. Ardalan, S.; Hosseinifard, M.; Vosough, M.; Golmohammadi, H. Towards smart personalized perspiration analysis: an IoT-integrated cellulose-based microfluidic wearable patch for smartphone fluorimetric multi-sensing of sweat biomarkers. Biosens. Bioelectron. 2020, 168, 112450.
20. Li, M.; Wang, L.; Liu, R.; et al. A highly integrated sensing paper for wearable electrochemical sweat analysis. Biosens. Bioelectron. 2021, 174, 112828.
21. Golparvar, A.; Kim, J.; Boukhayma, A.; Briand, D.; Carrara, S. Highly accurate multimodal monitoring of lactate and urea in sweat by soft epidermal optofluidics with single-band Raman scattering. Sens. Actuators B: Chem. 2023, 387, 133814.
22. Li, X.; Ou, R.; Yuan, Z.; Lin, Y. Wearable biosensors based on molecularly imprinted polymers. FlexMat 2025, 2, 475-92.
23. Nagel, B.; Dellweg, H.; Gierasch, L. M. Glossary for chemists of terms used in biotechnology (IUPAC Recommendations 1992). Pure Appl. Chem. 1992, 64, 143-68.
24. Cooper, J.M.; Cass, A.E.G. editors. Biosensors: a practical approach. Oxford: Oxford University Press; 2004.
25. Sarkar, S.; Hazra, S.; Patra, S.; Gogoi, M. Biosensors for cancer detection: a review. Trends Anal. Chem. 2024, 180, 117978.
26. Bahadır, E. B.; Sezgintürk, M. K. Electrochemical biosensors for hormone analyses. Biosens. Bioelectron. 2015, 68, 62-71.
27. Stoikov, D.; Ivanov, A.; Shafigullina, I.; et al. Flow-through amperometric biosensor system based on functionalized aryl derivative of phenothiazine and PAMAM-calix-dendrimers for the determination of uric acid. Biosensors 2024, 14, 120.
28. Lu, Z.; Wu, L.; Dai, X.; et al. Novel flexible bifunctional amperometric biosensor based on laser engraved porous graphene array electrodes: highly sensitive electrochemical determination of hydrogen peroxide and glucose. J. Hazard. Mater. 2021, 402, 123774.
29. Zhang, S.; Chen, Y. C.; Riezk, A.; et al. Rapid measurement of lactate in the exhaled breath condensate: biosensor optimization and in-human proof of concept. ACS Sens. 2022, 7, 3809-16.
30. Öndeş, B.; Evli, S.; Şahin, Y.; Uygun, M.; Uygun, D. A. Uricase based amperometric biosensor improved by AuNPs-TiS2 nanocomposites for uric acid determination. Microchem. J. 2022, 181, 107725.
31. Chen, L. Q.; Zhang, X. E.; Xie, W. H.; Zhou, Y. F.; Zhang, Z. P.; Cass, A. E. Genetic modification of glucose oxidase for improving performance of an amperometric glucose biosensor. Biosens. Bioelectron. 2002, 17, 851-7.
32. Han, Q.; Wang, H.; Wang, J. Multi‐Mode/Signal Biosensors: Electrochemical Integrated Sensing Techniques. Adv. Funct. Mater. 2024, 34, 2403122.
33. Bollella, P.; Gorton, L.; Antiochia, R. Direct electron transfer of dehydrogenases for development of 3rd generation biosensors and enzymatic fuel cells. Sensors (Basel). 2018, 18, 1319.
34. Wu, J.; Liu, H.; Chen, W.; Ma, B.; Ju, H. Device integration of electrochemical biosensors. Nat. Rev. Bioeng. 2023, 1, 346-60.
35. Bahrami, E.; Amini, R.; Vardak, S. Electrochemical detection of dopamine via pencil graphite electrodes modified by Cu/CuxO nanoparticles. J. Alloys Compd. 2021, 855, 157292.
36. Monteiro, T.; Coelho, A. R.; Moreira, M.; Viana, A. S.; Almeida, M. G. Interfacing the enzyme multiheme cytochrome c nitrite reductase with pencil lead electrodes: towards a disposable biosensor for cyanide surveillance in the environment. Biosens. Bioelectron. 2021, 191, 113438.
37. Zouari, M.; Campuzano, S.; Pingarrón, J. M.; Raouafi, N. Determination of miRNAs in serum of cancer patients with a label- and enzyme-free voltammetric biosensor in a single 30-min step. Mikrochim. Acta 2020, 187, 444.
38. Li, J.; Si, Y.; Park, Y. E.; et al. A serotonin voltammetric biosensor composed of carbon nanocomposites and DNA aptamer. Mikrochim. Acta 2021, 188, 146.
39. Raj, M.; Goyal, R. N. Silver nanoparticles and electrochemically reduced graphene oxide nanocomposite based biosensor for determining the effect of caffeine on Estradiol release in women of child-bearing age. Sens. Actuators B: Chem. 2019, 284, 759-67.
40. Eissa, S.; Zourob, M. Ultrasensitive peptide-based multiplexed electrochemical biosensor for the simultaneous detection of Listeria monocytogenes and Staphylococcus aureus. Mikrochim. Acta 2020, 187, 486.
41. Ding, J.; Qin, W. Recent advances in potentiometric biosensors. Trends Anal. Chem. 2020, 124, 115803.
42. Ibáñez-Redín, G.; Rosso Cagnani, G.; O Gomes, N.; et al. Wearable potentiometric biosensor for analysis of urea in sweat. Biosens. Bioelectron. 2023, 223, 114994.
43. Karimi-Maleh, H.; Orooji, Y.; Karimi, F.; et al. A critical review on the use of potentiometric based biosensors for biomarkers detection. Biosens. Bioelectron. 2021, 184, 113252.
44. Kulkarni, M. B.; Ayachit, N. H.; Aminabhavi, T. M. Biosensors and microfluidic biosensors: from fabrication to application. Biosensors 2022, 12, 543.
45. Ahmed, A.; Rushworth, J. V.; Hirst, N. A.; Millner, P. A. Bosensors for whole-cell bacterial detection. Clin. Microbiol. Rev. 2014, 27, 631-46.
46. Lazanas, A. C.; Prodromidis, M. I. Electrochemical impedance spectroscopy-a tutorial. ACS Meas. Sci. Au 2023, 3, 162-93.
47. Wang, S.; Zhang, J.; Gharbi, O.; Vivier, V.; Gao, M.; Orazem, M. E. Electrochemical impedance spectroscopy. Nat. Rev. Methods Primers 2021, 1, 41.
48. Lichtenberg, J. Y.; Ling, Y.; Kim, S. Non-specific adsorption reduction methods in biosensing. Sensors (Basel). 2019, 19, 2488.
49. Hassan, R. S. Wireless and battery-free biosensor based on parallel resonators for monitoring a wide range of biosignals. IEEE Trans. Microwave Theory Techn. 2022, 70, 4566-78.
50. Zapasnoy, A. S.; Belichenko, V. P.; Yakubov, V. P.; et al. Application of broadband microwave near-field sensors for glucose monitoring in biological media. Appl. Sci. 2021, 11, 1470.
51. Abdolrazzaghi, M.; Katchinskiy, N.; Elezzabi, A. Y.; Light, P. E.; Daneshmand, M. Noninvasive glucose sensing in aqueous solutions using an active split-ring resonator. IEEE Sensors J. 2021, 21, 18742-55.
52. Al-iraqi, E. Y.; Fayadh, R. A. Measurement of non-invasive blood glucose level by UWB transceiver in diabetic patient type-1. IOP Conf. Ser.: Mater. Sci. Eng. 2021, 1105, 012071.
53. Kandwal, A.; Sharma, Y. D.; Jasrotia, R.; et al. A comprehensive review on electromagnetic wave based non-invasive glucose monitoring in microwave frequencies. Heliyon 2024, 10, e37825.
54. Ma, Y.; Zhao, C.; Chen, C.; et al. Microfluidic microwave sensor based on a twisted cross-shaped structure for glucose detection. Sensors (Basel). 2025, 25, 3974.
55. Omer, A. E.; Shaker, G.; Safavi-Naeini, S.; et al. Low-cost portable microwave sensor for non-invasive monitoring of blood glucose level: novel design utilizing a four-cell CSRR hexagonal configuration. Sci. Rep. 2020, 10, 15200.
56. Dutt, S.; Thakur, R.; Kandwal, A.; et al. Emerging EM wave sensors for non-invasive glucose monitoring: review, techniques and developments. Sens. Actuators Rep. 2025, 9, 100282.
57. Zaarour, Y.; El Arroud, F.; Fernandez, T.; et al. Microwave antenna sensing for glucose monitoring in a vein model mimicking human physiology. Biosensors 2025, 15.
58. Yang, S.; Gao, S.; Zhuang, Y.; Hu, W.; Zhao, J.; Yi, Z. Non-destructive sensor for glucose solution concentration detection using electromagnetic technology. Micromachines (Basel). 2024, 15, 758.
59. Soltanian, F.; Nosrati, M.; Mobayen, S.; et al. On-body non-invasive glucose monitoring sensor based on high figure of merit (FoM) surface plasmonic microwave resonator. Sci. Rep. 2023, 13, 17527.
60. Farrugia, J.; Caruana, J.; Farhat, I.; Bonello, J.; Sammut, C. Complementary split ring resonator sensors for dielectric characterization of liquids in biosensing applications. Sens. Bio-Sens. Res. 2025, 49, 100839.
61. Gorst, A.; Zavyalova, K.; Mironchev, A.; Zapasnoy, A.; Klokov, A. Simulation and experimental study of the near field probe in the form of a folded dipole for measuring glucose concentration. Appl. Sci. 2021, 11, 5415.
62. Baghelani, M.; Abbasi, Z.; Daneshmand, M.; Light, P. E. Non-invasive continuous-time glucose monitoring system using a chipless printable sensor based on split ring microwave resonators. Sci. Rep. 2020, 10, 12980.
63. Gubeljak, P.; Xu, T.; Wlodarczyk, J.; et al. Highly sensitive glucose sensors based on gated graphene microwave waveguides. Adv. Sens. Res. 2024, 3, 2400091.
64. Lei, Y.; Zhang, D.; Wang, Q.; et al. Detection of carcinoembryonic antigen specificity using microwave biosensor with machine learning. Biosens. Bioelectron. 2025, 269, 116908.
65. Charvadeh, S. R.; Ghalibafan, J.; Abbasi, A.; Haidary, A.; Salimi, E. Utilization of the dialysate to characterize dialysis adequacy by using microwave urea biosensor based on complementary split-ring resonator (CSRR). Measurement 2025, 241, 115703.
66. Khalil, M. A.; Yong, W. H.; Islam, M. S.; et al. Cross enclosed square split ring resonator based on D.N.G. metamaterial absorber for X-band glucose sensing application. Heliyon 2024, 10, e26646.
67. Smith, D. R.; Padilla, W. J.; Vier, D. C.; Nemat-Nasser, S. C.; Schultz, S. Composite medium with simultaneously negative permeability and permittivity. Phys. Rev. Lett. 2000, 84, 4184-7.
68. Aminuzzaman, M. M.; Hossam-e-haider, M. Design and analysis of a metamaterial based biosensor to determine blood glucose concentration. IJECE. 2024, 14, 4088.
69. Gan, H.; Zhao, W.; Liu, Q.; et al. Differential microwave microfluidic sensor based on microstrip complementary split-ring resonator (MCSRR) structure. IEEE Sensors J. 2020, 20, 5876-84.
70. Ahmad, M.; Islam, M. T.; Alam, T.; et al. Coupled split square resonator based metamaterial sensor for realtime monitoring of air bubbles in IV channels. Sci. Rep. 2025, 15, 13453.
71. Nayak, R. K.; Mishra, M.; Shrivastava, K. K.; Bhoi, B.; Singh, R. Dielectric-tuned photon-photon coupling in a planar hybrid system. Chem. Phys. Impact 2025, 10, 100890.
72. Cebedio, M. C.; Rabioglio, L. A.; Gelosi, I. E.; Ribas, R. A.; Uriz, A. J.; Moreira, J. C. Analysis and design of a microwave coplanar sensor for non-invasive blood glucose measurements. IEEE Sensors J. 2020, 20, 10572-81.
73. Sameer, M.; Agarwal, P. Coplanar waveguide microwave sensor for label-free real-time glucose detection. RADIOENGINEERING 2019, 27, 491-5.
74. Kiani, S.; Rezaei, P.; Fakhr, M. Dual-frequency microwave resonant sensor to detect noninvasive glucose-level changes through the fingertip. IEEE Trans. Instrum. Meas. 2021, 70, 1-8.
75. Biurrun-Quel, C.; Teniente, J.; Del-Río, C. Reduced loss and prevention of substrate modes with a novel coplanar waveguide based on gap waveguide technology. Sensors (Basel). 2023, 23, 2909.
76. Hanna, J.; Bteich, M.; Tawk, Y.; et al. Noninvasive, wearable, and tunable electromagnetic multisensing system for continuous glucose monitoring, mimicking vasculature anatomy. Sci. Adv. 2020, 6, eaba5320.
77. Hanna, J.; Tawk, Y.; Azar, S.; et al. Wearable flexible body matched electromagnetic sensors for personalized non-invasive glucose monitoring. Sci. Rep. 2022, 12, 14885.
78. Sagar Shrikrishna, N.; Sharma, R.; Sahoo, J.; Kaushik, A.; Gandhi, S. Navigating the landscape of optical biosensors. Chem. Eng. J. 2024, 490, 151661.
79. Bratash, O.; Buhot, A.; Leroy, L.; Engel, E. Optical fiber biosensors toward in vivo detection. Biosens. Bioelectron. 2024, 251, 116088.
80. Gao, S.; Yang, X.; Wang, S.; et al. Review of optical fiber optofluidic chemical sensors and biosensors. Photonic Sens. 2024, 15, 250134.
81. Kang, J. W.; Park, Y. S.; Chang, H.; et al. Direct observation of glucose fingerprint using in vivo Raman spectroscopy. Sci. Adv. 2020, 6, eaay5206.
82. Uluç, N.; Glasl, S.; Gasparin, F.; et al. Non-invasive measurements of blood glucose levels by time-gating mid-infrared optoacoustic signals. Nat. Metab. 2024, 6, 678-86.
83. Wen, X.; Yang, X.; Ge, Z.; et al. Self-powered optical fiber biosensor integrated with enzymes for non-invasive glucose sensing. Biosens. Bioelectron. 2024, 253, 116191.
84. Guo, J.; Zhou, B.; Zong, R.; et al. Stretchable and highly sensitive optical strain sensors for human-activity monitoring and healthcare. ACS Appl. Mater. Interfaces 2019, 11, 33589-98.
85. Wu J, ; Jiang K, ; Mi H, ; et al. A rapid and sensitive fluorescence biosensor based on plasmonic PCR. Nanoscale 2021, 13, 7348-54.
86. Lin, Y.; Huang, Y.; Wang, T.; et al. Bioinspired flexible photonic sensing chips for wearable multiparameter monitoring. FlexMat 2025, 2, 365-89.
87. Xiong, Q.; Lim, C. Y.; Ren, J.; et al. Magnetic nanochain integrated microfluidic biochips. Nat. Commun. 2018, 9, 1743.
88. Feng, L.; Liu, Y.; Wang, Y.; Zhou, H.; Lu, Z.; Li, T. Ultra-compact dual-channel integrated CO2 infrared gas sensor. Microsyst. Nanoeng. 2024, 10, 151.
89. Shan, Y.; Wang, B.; Huang, H.; et al. On-site quantitative Hg2+ measurements based on selective and sensitive fluorescence biosensor and miniaturized smartphone fluorescence microscope. Biosens. Bioelectron. 2019, 132, 238-47.
90. Zhang, J.; Mai, X.; Hong, X.; Chen, Y.; Li, X. Optical fiber SPR biosensor with a solid-phase enzymatic reaction device for glucose detection. Sens. Actuators B: Chem. 2022, 366, 131984.
91. Chen, C.; Singh, R.; Huo, S.; et al. Evanescent wave-based optical biosensors for innovations, medical application and future perspectives. J. Adv. Res. 2026, 82, 295-313.
92. Zhang, C.; Huang, L.; Pu, H.; Sun, D. Magnetic surface-enhanced Raman scattering (MagSERS) biosensors for microbial food safety: fundamentals and applications. Trends Food Sci. Technol. 2021, 113, 366-81.
93. Wang, Y.; Jia, K.; Lin, J. Optical biosensors for the detection of foodborne pathogens: recent development and future prospects. Trends Anal. Chem. 2024, 177, 117785.
94. Špringer, T.; Bocková, M.; Slabý, J.; Sohrabi, F.; Čapková, M.; Homola, J. Surface plasmon resonance biosensors and their medical applications. Biosens. Bioelectron. 2025, 278, 117308.
95. Mitsubayashi, K.; Toma, K.; Iitani, K.; Arakawa, T. Gas-phase biosensors: a review. Sens. Actuators B: Chem. 2022, 367, 132053.
96. Wang, X.; Zhou, J.; Wang, H. Bioreceptors as the key components for electrochemical biosensing in medicine. Cell Rep. Phy. Sci. 2024, 5, 101801.
97. Mitsubayashi, K.; Nishio, G.; Sawai, M.; et al. A bio-sniffer stick with FALDH (formaldehyde dehydrogenase) for convenient analysis of gaseous formaldehyde. Sens. Actuators B: Chem. 2008, 130, 32-7.
98. Sellés Vidal, L.; Kelly, C. L.; Mordaka, P. M.; Heap, J. T. Review of NAD(P)H-dependent oxidoreductases: Properties, engineering and application. Biochim. Biophys. Acta. Proteins Proteom. 2018, 1866, 327-47.
99. Achmann, S.; Hämmerle, M.; Kita, J.; Moos, R. Miniaturized low temperature co-fired ceramics (LTCC) biosensor for amperometric gas sensing. Sens. Actuators B: Chem. 2008, 135, 89-95.
100. Li, R.; Liu, Y.; Zhou, J.; et al. Boosting ultrasensitive electroanalytical detection of antibiotics at triphasic interface enzymatic biosensor. Biosens. Bioelectron. 2025, 280, 117430.
101. Kudo, H.; Yamashita, T.; Ye, M.; et al. NADH-fluorometric biochemical gas sensor (bio-sniffer) for assessment of indoor air quality. In 2012 IEEE Sensors, Taipei, Taiwan, October 28-31, 2012; IEEE, 2012, pp 1-4.
102. Kudo, H.; Sawai, M.; Suzuki, Y.; et al. Fiber-optic bio-sniffer (biochemical gas sensor) for high-selective monitoring of ethanol vapor using 335 nm UV-LED. Sens. Actuators B: Chem. 2010, 147, 676-80.
103. Chaudhary, V.; Taha, B. A. L. U. C. K. Y.; et al. Nose-on-chip nanobiosensors for early detection of lung cancer breath biomarkers. ACS Sens. 2024, 9, 4469-94.
104. Ma, J.; Guan, Y.; Xing, F.; et al. Accurate and non-destructive monitoring of mold contamination in foodstuffs based on whole-cell biosensor array coupling with machine-learning prediction models. J. Hazard. Mater. 2023, 449, 131030.
105. Deng, H.; Mitsuno, H.; Kuroda, E.; Niki, S.; Kanzaki, R.; Nakamoto, T. Gas-phase odor mixture quantification based on relative comparison method using multiple olfactory receptors. Sens. Actuators B: Chem. 2024, 401, 134995.
106. Hu, J.; Hu, N.; Pan, D.; et al. Smell cancer by machine learning-assisted peptide/MXene bioelectronic array. Biosens. Bioelectron. 2024, 262, 116562.
107. Liu, S.; Hu, N.; Hu, J.; et al. DNA-mediated bioinspired MXene gas sensor array with machine learning for noninvasive cancer recognition. ACS Nano 2025, 19, 25363-84.
108. Jafari, B.; Gholizadeh, E.; Jafari, B.; et al. Highly sensitive label-free biosensor: graphene/CaF2 multilayer for gas, cancer, virus, and diabetes detection with enhanced quality factor and figure of merit. Sci. Rep. 2023, 13, 16184.
109. Gray, E. R.; Turbé, V.; Lawson, V. E.; et al. Ultra-rapid, sensitive and specific digital diagnosis of HIV with a dual-channel SAW biosensor in a pilot clinical study. NPJ Digit. Med. 2018, 1, 35.
110. Xiao, L.; Yu, S.; Cui, Q.; et al. Phthalocyanine-functionalized field-effect transistor biosensor for in situ monitoring of cell-released hydrogen sulfide. Sens. Actuators B: Chem. 2024, 406, 135402.
111. Hu, H.; Huang, H.; Li, M.; et al. A wearable cardiac ultrasound imager. Nature 2023, 613, 667-75.
112. Zeng, Y.; Yuan, R.; Fu, H.; Xu, Z.; Wei, S. Foodborne pathogen detection using surface acoustic wave biosensors: a review. RSC Adv. 2024, 14, 37087-103.
113. Xiong, H.; Zhang, X.; Sun, J.; et al. Recent advances in biosensors detecting biomarkers from exhaled breath and saliva for respiratory disease diagnosis. Biosens. Bioelectron. 2025, 267, 116820.
114. Obořilová, R.; Kučerová, E.; Botka, T.; Vaisocherová-Lísalová, H.; Skládal, P.; Farka, Z. Piezoelectric biosensor with dissipation monitoring enables the analysis of bacterial lytic agent activity. Sci. Rep. 2025, 15, 3419.
115. Shahbaz, A.; Hussain, N.; Intisar, A.; Bilal, M.; Iqbal, H. M. N. Immobilized enzymes-based biosensing cues for strengthening biocatalysis and biorecognition. Catal. Lett. 2021, 152, 2637-49.
116. Wang, L.; Sipe, D. M.; Xu, Y.; Lin, Q. A MEMS thermal biosensor for metabolic monitoring applications. J. Microelectromech. Syst. 2008, 17, 318-27.
117. Zhang, Q.; Deng, K.; Wilkens, L.; Reith, H.; Nielsch, K. Micro-thermoelectric devices. Nat. Electron. 2022, 5, 333-47.
118. Tang, X.; Qi, C.; Sun, Q. Recent progress of biosensors based on thermoelectric effects for monitoring physical activity and environment monitoring. Soft Sci. 2025, 5, 11.
119. Yuan, J.; Zhu, R.; Li, G. Self-powered electronic skin with multisensory functions based on thermoelectric conversion. Adv. Mater. Technol. 2020, 5, 2000419.
120. Kang, M.; Qu, R.; Sun, X.; et al. Self-powered temperature electronic skin based on island-bridge structure and Bi-Te micro-thermoelectric generator for distributed mini-region sensing. Adv. Mater. 2023, 35, e2309629.
121. Kus, F.; Altinkok, C.; Zayim, E.; Erdemir, S.; Tasaltin, C.; Gurol, I. Surface acoustic wave (SAW) sensor for volatile organic compounds (VOCs) detection with calix[4]arene functionalized Gold nanorods (AuNRs) and silver nanocubes (AgNCs). Sens. Actuators B: Chem. 2021, 330, 129402.
122. Xiong, J.; Sun, Z.; Yu, J.; Liu, H.; Wang, X. Thermal self-regulatory smart biosensor based on horseradish peroxidase-immobilized phase-change microcapsules for enhancing detection of hazardous substances. Chem. Eng. J. 2022, 430, 132982.
123. McMurry JF, J. R. Insulin therapy in diabetes. The importance of good glucose control. Postgrad. Med. 1986, 80, 191-4, 199-202.
124. Ciobanu, C.; Ember, K. J.; Nyiri, B. J.; et al. Potential of Raman spectroscopy for blood-based biopsy. IEEE Instrum. Meas. Mag. 2022, 25, 62-8.
125. Rossi, F.; Trakoolwilaiwan, T.; Gigli, V.; et al. Progress in nanoparticle-based electrochemical biosensors for hormone detection. Nanoscale 2024, 16, 18134-64.
126. Dong, T.; Zhu, W.; Yang, Z.; et al. Advances in heart failure monitoring: Biosensors targeting molecular markers in peripheral bio-fluids. Biosens. Bioelectron. 2024, 255, 116090.
127. Tombetti, E.; Hysa, E.; Mason, J. C.; Cimmino, M. A.; Camellino, D. Blood biomarkers for monitoring and prognosis of large vessel vasculitides. Curr. Rheumatol. Rep. 2021, 23, 17.
128. Maeda, Y.; Okawara, H.; Sawada, T.; et al. Implications of the onset of sweating on the sweat lactate threshold. Sensors (Basel). 2023, 23, 3378.
129. Shitanda, I.; Mitsumoto, M.; Loew, N.; et al. Continuous sweat lactate monitoring system with integrated screen-printed MgO-templated carbon-lactate oxidase biosensor and microfluidic sweat collector. Electrochim. Acta 2021, 368, 137620.
130. Ichihashi, K.; Konno, D.; Maryunina, K. Y.; et al. Selective ion exchange in supramolecular channels in the crystalline state. Angew. Chem. Int. Ed. Engl. 2019, 58, 4169-72.
131. Baker, L. B.; Wolfe, A. S. Physiological mechanisms determining eccrine sweat composition. Eur. J. Appl. Physiol. 2020, 120, 719-52.
132. Hu, M.; Zhu, K.; Wei, J.; et al. Silk fibroin-based wearable SERS biosensor for simultaneous sweat monitoring of creatinine and uric acid. Biosens. Bioelectron. 2024, 265, 116662.
133. Estiu, G.; Merz KM, J. R. The hydrolysis of urea and the proficiency of urease. J. Am. Chem. Soc. 2004, 126, 6932-44.
134. Li, H.; Xiao, Y.; Jin, L. Recent development of wearable electrochemical sensors for sweat analysis. Int. J. Electrochem. Sci. 2020, 15, 11997-2006.
135. Cho, M. T.; Goldstein, L.; Akiba, C.; et al. RAS mutational status and CEA production at initial presentation in metastatic colorectal cancer. J. Clin. Oncol. 2016, 34, 542.
136. Xu, Y.; Zhu, S.; Xia, C.; et al. Liquid biopsy-based multi-cancer early detection: an exploration road from evidence to implementation. Sci. Bull. (Beijing). 2025, 70, 2852-67.
137. Li, Z.; Wang, B.; Yin, T.; Jie, G. Self-reinforced D-MOF and autocatalytic AuPd@SnS2 dual-wavelength electrochemiluminescence biosensor for detection of ctDNA and CEA. Sens. Actuators B: Chem. 2025, 422, 136679.
138. Adhyam, M.; Gupta, A. K. A review on the clinical utility of PSA in cancer prostate. Indian J. Surg. Oncol. 2012, 3, 120-9.
139. Galle, P. R.; Foerster, F.; Kudo, M.; et al. Biology and significance of alpha-fetoprotein in hepatocellular carcinoma. Liver Int. 2019, 39, 2214-29.
140. Charkhchi, P.; Cybulski, C.; Gronwald, J.; Wong, F. O.; Narod, S. A.; Akbari, M. R. CA125 and ovarian cancer: a comprehensive review. Cancers (Basel). 2020, 12, 3730.
141. Morales, M. A.; Halpern, J. M. Guide to selecting a biorecognition element for biosensors. Bioconjugate Chem. 2018, 29, 3231-9.
142. Chen, A.; Yang, S. Replacing antibodies with aptamers in lateral flow immunoassay. Biosens. Bioelectron. 2015, 71, 230-42.
143. Liang, B.; Wang, G.; Yan, L.; et al. Functional cell surface displaying of acetylcholinesterase for spectrophotometric sensing organophosphate pesticide. Sens. Actuators B: Chem. 2019, 279, 483-9.
144. Cao, J.; Wang, M.; She, Y.; et al. Rapid colorimetric determination of the pesticides carbofuran and dichlorvos by exploiting their inhibitory effect on the aggregation of peroxidase-mimicking platinum nanoparticles. Mikrochim. Acta 2019, 186, 390.
145. Chen, Y.; Qian, C.; Liu, C.; et al. Nucleic acid amplification free biosensors for pathogen detection. Biosens. Bioelectron. 2020, 153, 112049.
146. Song, F.; Shen, Y.; Wei, Y.; et al. Botulinum toxin as an ultrasensitive reporter for bacterial and SARS-CoV-2 nucleic acid diagnostics. Biosens. Bioelectron. 2021, 176, 112953.
147. Jia, M.; Liao, X.; Fang, L.; et al. Recent advances on immunosensors for mycotoxins in foods and other commodities. Trends Anal. Chem. 2021, 136, 116193.
148. Huang, L.; Tang, D.; Yang, Z. Flexible electronic materials and devices toward portable immunoassays. FlexMat 2024, 1, 59-78.
149. Shen, L.; Chen, Y.; Pan, J.; et al. Development of a highly sensitive PbrR-based biosensor via directed evolution and its application for lead detection. J. Hazard. Mater. 2025, 488, 137489.
150. Vasanthi Sridharan, N.; Mandal, B. K. Simultaneous quantitation of lead and cadmium on an EDTA-reduced graphene oxide-modified glassy carbon electrode. ACS Omega 2022, 7, 45469-80.
151. Safwat, S. M.; Khaled, A.; Elawwad, A.; Matta, M. E. Dual-chamber microbial fuel cells as biosensors for the toxicity detection of benzene, phenol, chromium, and copper in wastewater: Applicability investigation, effect of various catholyte solutions, and life cycle assessment. Process Saf. Environ. Prot. 2023, 170, 1121-36.
152. Abu-Ali, H.; Nabok, A.; Smith, T. J. Electrochemical inhibition bacterial sensor array for detection of water pollutants: artificial neural network (ANN) approach. Anal. Bioanal. Chem. 2019, 411, 7659-68.
153. Liu, X.; Hou, Y.; Chen, S.; Liu, J. Controlling dopamine binding by the new aptamer for a FRET-based biosensor. Biosens. Bioelectron. 2021, 173, 112798.
154. Nguyen, D. C. T.; Vu Thi, Q.; Nguyen, Q. H.; et al. Engineering flexible dopamine biosensors: blended EGylated conjugated and radical polymers in organic electrochemical transistors. NPJ Flex. Electron. 2025, 9, 35.
155. Jeong, S.; Yang, D.; Beyene, A. G.; et al. High-throughput evolution of near-infrared serotonin nanosensors. Sci. Adv. 2019, 5, eaay3771.
156. Henke, A.; Kovalyova, Y.; Dunn, M.; et al. Toward serotonin fluorescent false neurotransmitters: development of fluorescent dual serotonin and vesicular monoamine transporter substrates for visualizing serotonin neurons. ACS Chem. Neurosci. 2018, 9, 925-34.
157. Al-Rawhani, M. A.; Hu, C.; Giagkoulovits, C.; et al. Multimodal integrated sensor platform for rapid biomarker detection. IEEE Trans. Biomed. Eng. 2020, 67, 614-23.
158. Zhao, Z.; Cheng, J.; Ren, Y.; et al. Multimodal sensing platform based on Fe3O4/DEX/PDA@Au(Raman reporters)@Au nanocomposites for sweat biomarkers monitoring. Biosens. Bioelectron. 2025, 286, 117629.
159. Yan, R.; Sang, G.; Yin, B.; et al. Temperature self-calibrated pH sensor based on GO/PVA-coated MZI cascading FBG. Opt. Express 2021, 29, 13530-41.
160. Yakovleva, M.; Bhand, S.; Danielsson, B. The enzyme thermistor - a realistic biosensor concept. A critical review. Anal. Chim. Acta 2013, 766, 1-12.
161. De J. Souza, E. L.; Santos, T. L. Transient compensation for thermistor-based sensors in constant temperature configuration. Sensor. Actuat. A-Phys. 2020, 305, 111920.
162. Wang, A.; Zou, Y.; Liu, S.; et al. Comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo using highly responsive biosensors. Nat. Protoc. 2024, 19, 1311-47.
163. Qin, X.; Li, D.; Qin, X.; et al. Electrochemical detection of the cardiac biomarker cardiac troponin I. VIEW 2024, 5, 20240025.
164. Dong, S.; Li, Y.; Gu, C.; Mao, J. Robust cardiac timing detection technique with vectors analytic demodulation in doppler cardiogram sensing. IEEE Trans. Microwave Theory Techn. 2024, 72, 4866-77.
165. Chen, D.; Wang, X.; Zhang, K.; et al. Glucose photoelectrochemical enzyme sensor based on competitive reaction of ascorbic acid. Biosens. Bioelectron. 2020, 166, 112466.
166. Zhang, Y.; Discekici, E. H.; Burns, R. L.; Somervell, M. H.; Hawker, C. J.; Bates, C. M. Single-step, spin-on process for high fidelity and selective deposition. ACS Appl. Polym. Mater. 2019, 2, 481-6.
167. Zhang, N.; Zhang, Y.; Chen, J.; et al. Multi-signal acquisition system for continuous blood pressure monitoring. Sensors (Basel). 2025, 25, 5910.
168. Huang, Y.; Sun, X.; Zhang, Z.; et al. Metal-organic framework-based dual-mode biosensors: Mechanisms and applications. Coord. Chem. Rev. 2025, 545, 217006.
169. Dai, L.; Zhao, X.; Guo, J.; et al. Microfluidics-based microwave sensor. Sensor. Actuat. A-Phys. 2020, 309, 111910.
170. Li, T.; Zhang, J.; Bu, P.; Wu, H.; Guo, J.; Guo, J. Multi-modal nanoprobe-enabled biosensing platforms: a critical review. Nanoscale 2024, 16, 3784-816.
171. Wang, E.; Chen, X.; Li, Y.; Fu, Z.; Huang, J. Lower limb motion intent recognition based on sensor fusion and fuzzy multitask learning. IEEE Trans. Fuzzy Syst. 2024, 32, 2903-14.
172. Ma, S.; Wan, Z.; Wang, C.; et al. Ultra-sensitive and stable multiplexed biosensors array in fully printed and integrated platforms for reliable perspiration analysis. Adv. Mater. 2024, 36, e2311106.
173. Huber, D.; Falco-jonasson, L.; Talary, M.; et al. Multi-sensor data fusion for non-invasive continuous glucose monitoring. In 2007 10th International Conference on Information Fusion, Quebec City, QC, Canada, July 9-12, 2007; IEEE, 2007, pp 1-10.
174. Agarwal, S.; Varghese, R. T.; Savian, R.; Low Wang, C. C.; Galindo, R. J. Moving beyond glycated hemoglobin to glucose patterns: newer indications for continuous glucose monitor use. Endocr. Pract. 2025, 31, 1046-53.
175. Xu, J.; Chen, Y.; Chen, H.; Cui, Q.; Tao, X. Use of two-mode circuitry and optimal energy efficient power control under target delay-outage constraints. In 2017 IEEE 28th Annual International Symposium on Personal, Indoor, and Mobile Radio Communications (PIMRC), Montreal, QC, Canada, October 8-13, 2017; IEEE, 2017, pp 1-7.
176. Wu, S.; Kan, H.; Gao, J.; et al. Convolutional neural networks-motivated high-performance multi-functional electronic skin for intelligent human-computer interaction. Nano Energy 2024, 122, 109313.
177. Zhang, D.; Xiang, Y.; Zou, Q.; Zhu, K.; Hu, N. Electromechanical integrated recording of single cardiomyocyte in situ by multimodal microelectrode biosensing system. Biosens. Bioelectron. 2022, 212, 114387.
178. Li, Z.; Li, Q.; Zhu, Z.; Hu, Z.; Wu, X. Multi-scale spatio-temporal fusion with adaptive brain topology learning for fMRI based neural decoding. IEEE J. Biomed. Health Inform. 2023, PP.
179. Guo, W.; Lei, Y.; Zhao, X.; et al. Printed-scalable microstructure BaTiO3/ecoflex nanocomposite for high-performance triboelectric nanogenerators and self-powered human-machine interaction. Nano Energy 2024, 131, 110324.
180. Yin, Y.; Zhao, Z.; Na, J.; Xue, C.; Qin, K. Physiologically inspired spatiotemporal adaptive multimodal fusion model for blood glucose prediction. Biomed. Signal Process. Control. 2025, 109, 107998.
181. Byrne, R.; Carrico, A.; Lettieri, M.; Rajan, A. K.; Forster, R. J.; Cumba, L. R. Bioinks and biofabrication techniques for biosensors development: a review. Mater. Today. Bio 2024, 28, 101185.
182. Wilson WC, J. R.; Boland, T. Cell and organ printing 1: protein and cell printers. Anat. Rec. A. Discov. Mol. Cell. Evol. Biol. 2003, 272, 491-6.
183. Boland, T.; Mironov, V.; Gutowska, A.; Roth, E. A.; Markwald, R. R. Cell and organ printing 2: fusion of cell aggregates in three-dimensional gels. Anat. Rec. A. Discov. Mol. Cell. Evol. Biol. 2003, 272, 497-502.
184. Li, J.; Rossignol, F.; Macdonald, J. Inkjet printing for biosensor fabrication: combining chemistry and technology for advanced manufacturing. Lab Chip 2015, 15, 2538-58.
185. Zhang, T.; Zhao, W.; Xiahou, Z.; Wang, X.; Zhang, K.; Yin, J. Bioink design for extrusion-based bioprinting. Appl. Mater. Today 2021, 25, 101227.
186. Grigoryan, B.; Sazer, D. W.; Avila, A.; et al. Development, characterization, and applications of multi-material stereolithography bioprinting. Sci. Rep. 2021, 11, 3171.
187. Gurkan, U. A.; El Assal, R.; Yildiz, S. E.; et al. Engineering anisotropic biomimetic fibrocartilage microenvironment by bioprinting mesenchymal stem cells in nanoliter gel droplets. Mol. Pharm. 2014, 11, 2151-9.
188. Chimene, D.; Lennox, K. K.; Kaunas, R. R.; Gaharwar, A. K. Advanced bioinks for 3D printing: a materials science perspective. Ann. Biomed. Eng. 2016, 44, 2090-102.
189. Malda, J.; Visser, J.; Melchels, F. P.; et al. 25th anniversary article: engineering hydrogels for biofabrication. Adv. Mater. 2013, 25, 5011-28.
190. Kolesky, D. B.; Truby, R. L.; Gladman, A. S.; Busbee, T. A.; Homan, K. A.; Lewis, J. A. 3D bioprinting of vascularized, heterogeneous cell-laden tissue constructs. Adv. Mater. 2014, 26, 3124-30.
191. Rutz, A. L.; Lewis, P. L.; Shah, R. N. Toward next-generation bioinks: tuning material properties pre- and post-printing to optimize cell viability. MRS Bull. 2017, 42, 563-70.
192. Guvendiren, M.; Molde, J.; Soares, R. M.; Kohn, J. Designing biomaterials for 3D printing. ACS Biomater. Sci. Eng. 2016, 2, 1679-93.
193. Xiong, R.; Zhang, Z.; Chai, W.; Chrisey, D. B.; Huang, Y. Study of gelatin as an effective energy absorbing layer for laser bioprinting. Biofabrication 2017, 9, 024103.
194. Zhai, X.; Ma, Y.; Hou, C.; et al. 3D-printed high strength bioactive supramolecular polymer/clay nanocomposite hydrogel scaffold for bone regeneration. ACS Biomater. Sci. Eng. 2017, 3, 1109-18.
195. Liu, W.; Heinrich, M. A.; Zhou, Y.; et al. Extrusion bioprinting of shear-thinning gelatin methacryloyl bioinks. Adv. Healthc. Mater. 2017, 6, e1601451.
196. Moulahoum, H. Dual chromatic laser-printed microfluidic paper-based analytical device (μPAD) for the detection of atrazine in water. ACS Omega 2023, 8, 41194-203.
197. Rodríguez-pombo, L.; Xu, X.; Seijo-rabina, A.; et al. Volumetric 3D printing for rapid production of medicines. Addit. Manuf. 2022, 52, 102673.
198. Piao, C.; Du, X.; Xu, Y.; To, S.; Zhu, L.; Zhu, Z. Time-dependent volumetric printing of precision lenses through dynamic laser writing. Int. J. Extrem. Manuf. 2025, 7, 045007.
199. Bhattacharya, I.; Toombs, J.; Taylor, H. High fidelity volumetric additive manufacturing. Addit. Manuf. 2021, 47, 102299.
200. Rizzo, R.; Ruetsche, D.; Liu, H.; Zenobi-Wong, M. Optimized photoclick (bio)resins for fast volumetric bioprinting. Adv. Mater. 2021, 33, e2102900.
201. Zhou, J.; Li, Q.; Tian, Z.; Yao, Q.; Zhang, M. Recent advances in 3D bioprinted cartilage-mimicking constructs for applications in tissue engineering. Mater. Today. Bio 2023, 23, 100870.
202. Grigoryan, B.; Paulsen, S. J.; Corbett, D. C.; et al. Multivascular networks and functional intravascular topologies within biocompatible hydrogels. Science 2019, 364, 458-64.
203. Raman, R.; Bhaduri, B.; Mir, M.; et al. High-resolution projection microstereolithography for patterning of neovasculature. Adv. Healthc. Mater. 2016, 5, 610-9.
204. Antonelli, G.; Mencattini, A.; Massimiani, M.; et al. Development of integrated optical biosensors based on low-cost stereolithography fabrication and multispectral signature for Lab-On-Chip applications. Sens. Actuators B: Chem. 2024, 401, 135009.
205. Zhao, Z.; Tian, X.; Song, X. Engineering materials with light: recent progress in digital light processing based 3D printing. J. Mater. Chem. C. 2020, 8, 13896-917.
206. He, Y., Gao, Q. and Jin, Y. Digital light projection-based 3D bioprinting. In Cell Assembly with 3D Bioprinting; Wiley, 2021; pp 89-111.
207. Ge, L.; Dong, L.; Wang, D.; Ge, Q.; Gu, G. A digital light processing 3D printer for fast and high-precision fabrication of soft pneumatic actuators. Sensor. Actuat. A-Phys. 2018, 273, 285-92.
208. Tripathi, S.; Mandal, S. S.; Bauri, S.; Maiti, P. 3D bioprinting and its innovative approach for biomedical applications. MedComm (2020). 2023, 4, e194.
209. Budharaju, H.; Sundaramurthi, D.; Sethuraman, S. Embedded 3D bioprinting - an emerging strategy to fabricate biomimetic & large vascularized tissue constructs. Bioact. Mater. 2024, 32, 356-84.
210. Wang, H.; Zhou, C.; Liang, J.; Yu, H.; Peng, F. An enhanced glucose biosensor modified by Pt/sulfonated-MWCNTs with layer by layer technique. Int. J. Electrochem. Sci. 2008, 3, 1180-5.
211. Hasan, E.; Lewis, C. J.; Hernandez, J. G.; Gentile, P.; Ferreira, A. M. Layer-by-layer assembly: advancing skin repair, one layer at a time. RSC Adv. 2025, 15, 13908-23.
212. Mariani, S.; Robbiano, V.; Strambini, L. M.; et al. Layer-by-layer biofunctionalization of nanostructured porous silicon for high-sensitivity and high-selectivity label-free affinity biosensing. Nat. Commun. 2018, 9, 5256.
213. Liu, H.; Zhong, H.; Yuan, Q.; et al. Roll-to-roll manufacturing of breathable superhydrophobic membranes. Small Methods 2024, 8, e2400038.
214. Fung, C.; Lloyd, J.; Samavat, S.; Deganello, D.; Teng, K. Facile fabrication of electrochemical ZnO nanowire glucose biosensor using roll to roll printing technique. Sens. Actuators B: Chem. 2017, 247, 807-13.
215. Bariya, M.; Shahpar, Z.; Park, H.; et al. Roll-to-roll gravure printed electrochemical sensors for wearable and medical devices. ACS Nano 2018, 12, 6978-87.
216. Liedert, C.; Rannaste, L.; Kokkonen, A.; et al. Roll-to-roll manufacturing of integrated immunodetection sensors. ACS Sens. 2020, 5, 2010-7.
217. Lamaakal, I.; Essahraui, S.; Maleh, Y.; et al. A comprehensive survey on tiny machine learning for human behavior analysis. IEEE Internet Things J. 2025, 12, 32419-43.
218. Jin, X.; Liu, C.; Xu, T.; Su, L.; Zhang, X. Artificial intelligence biosensors: challenges and prospects. Biosens. Bioelectron. 2020, 165, 112412.
219. Mansour, M.; Saeed Darweesh, M.; Soltan, A. Wearable devices for glucose monitoring: A review of state-of-the-art technologies and emerging trends. Alex. Eng. J. 2024, 89, 224-43.
220. Joshi, A. M.; Jain, P.; Mohanty, S. P.; Agrawal, N. iGLU 2.0: A new wearable for accurate non-invasive continuous serum glucose measurement in IoMT framework. IEEE Trans. Consumer Electron. 2020, 66, 327-35.
221. Di Filippo, D.; Sunstrum, F. N.; Khan, J. U.; Welsh, A. W. Non-invasive glucose sensing technologies and products: a comprehensive review for researchers and clinicians. Sensors (Basel). 2023, 23.
222. Siegel, P. H.; Tang, A.; Virbila, G.; Kim, Y.; Chang, M. C. F.; Pikov, V. Compact non-invasive millimeter-wave glucose sensor. In 2015 40th International Conference on Infrared, Millimeter, and Terahertz waves (IRMMW-THz), Hong Kong, China, August 23-28, 2015; IEEE, 2015, pp 1-3.
223. Gharbi, M. E.; Fernández-garcía, R.; Gil, I. Textile antenna-sensor for in vitro diagnostics of diabetes. Electronics 2021, 10, 1570.
224. Shi, J.; Fernández-García, R.; Gil, I. Sensor technologies for non-invasive blood glucose monitoring. Sensors (Basel). 2025, 25, 3591.
225. Tanaka, T.; Mandic, D. P. Complex empirical mode decomposition. IEEE Signal Process. Lett. 2007, 14, 101-4.
226. Shensa, M. The discrete wavelet transform: wedding the a trous and Mallat algorithms. IEEE Trans. Signal Process. 1992, 40, 2464-82.
227. Bamatraf, S. M.; Ramahi, O. M.; Aldhaeebi, M. A. Non-invasive aqueous glucose monitoring using microwave sensor with machine learning. In 2021 IEEE International Symposium on Antennas and Propagation and USNC-URSI Radio Science Meeting (APS/URSI), Singapore, Singapore, December 4-10, 2021; IEEE, 2021, pp 1875-6.
228. Sattari, M. A.; Hayati, M. Detection of fasting blood sugar using a microwave sensor and convolutional neural network. Sci. Rep. 2025, 15, 22937.
229. Kazemi, N.; Abdolrazzaghi, M.; Light, P. E.; Musilek, P. In-human testing of a non-invasive continuous low-energy microwave glucose sensor with advanced machine learning capabilities. Biosens. Bioelectron. 2023, 241, 115668.
230. Belfarsi, E. A.; Flores, H.; Valero, M. Reliable noninvasive glucose sensing via CNN-based spectroscopy, arXiv 2025;arXiv:2506.13819. Available online: https://doi.org/10.48550/arXiv.2506.13819. [accessed 15 April 2026].
231. Naresh, M.; Nagaraju, V. S.; Kollem, S.; Kumar, J.; Peddakrishna, S. Non-invasive glucose prediction and classification using NIR technology with machine learning. Heliyon 2024, 10, e28720.
232. Dai, B.; You, S.; Wang, K.; et al. Deep learning-enabled filter-free fluorescence microscope. Sci. Adv. 2025, 11, eadq2494.
233. Glehr, G.; Riquelme, P.; Kronenberg, K.; et al. Restricting datasets to classifiable samples augments discovery of immune disease biomarkers. Nat. Commun. 2024, 15, 5417.
234. Hu, X. B.; Wang, Y. L.; Lv, J. L.; et al. A chemical redox cycling-based dual-mode biosensor for self-powered photoelectrochemical and colorimetric assay of heat shock protein. ACS Sens. 2025, 10, 196-203.
235. Li, M.; Wang, H.; Yu, X.; et al. A sensitive and simple competitive nanozyme-linked apta-sorbent assay for the dual-mode detection of ochratoxin A. Analyst 2022, 147, 2215-22.
236. Suo, Z.; Lu, T.; Liu, J.; et al. Electrochemistry-glucosemeter-smartphone integrated multi-mode biosensor for accurate detection of aflatoxin B1. Anal. Chim. Acta 2025, 1333, 343398.
237. Kong, Y.; Li, Z.; Zhang, L.; et al. A novel Nb2C MXene based aptasensor for rapid and sensitive multi-mode detection of AFB1. Biosens. Bioelectron. 2023, 242, 115725.
238. Shi, Y.; Zhou, K.; Ma, X.; et al. Washable textile biosensors enabled by nanostructured oxides with fast ion diffusion. Device 2024, 2, 100503.
239. Zhang, L.; Niknejad, A. M. Design and analysis of a microwave-optical dual modality biomolecular sensing platform. IEEE J. Solid-State Circuits 2020, 55, 639-49.
240. Xu, Z.; Sun, H.; Chen, Y.; Yu, H. H.; Deng, C. X.; Xu, Q. Bubble-inspired multifunctional magnetic microrobots for integrated multidimensional targeted biosensing. Nano Lett. 2024, 24, 13945-54.
241. Gu, M.; Guo, C. F.; Song, Y. Multimodal bioelectronics: a pathway to digital health management. Matter 2025, 8, 102048.
242. Buttkewitz, M. A.; Heuer, C.; Bahnemann, J. Sensor integration into microfluidic systems: trends and challenges. Curr. Opin. Biotechnol. 2023, 83, 102978.
243. Ding, Q.; Wang, K.; Shi, F.; et al. Direct charge-transfer chemosensing materials for painless multimodal disease diagnosis. Acc. Mater. Res. 2025, 6, 1435-50.
244. Liu, T.; Wu, Q.; Liu, H.; et al. A crosslinked eutectogel for ultrasensitive pressure and temperature monitoring from nostril airflow. Nat. Commun. 2025, 16, 3334.
245. Wang, J.; Huang, D.; Zheng, D.; Shen, F.; Zhang, Y. Selectively quantify toxic pollutants in water by machine learning empowered electrochemical biosensors. Environ. Sci. Technol. 2025, 59, 616-27.
246. Blankenberg, D.; Coraor, N.; Von Kuster, G.; Taylor, J.; Nekrutenko, A.; Galaxy Team. Integrating diverse databases into an unified analysis framework: a galaxy approach. Database (Oxford). 2011, 2011, bar011.
247. Amará, J.; Ströele, V.; Braga, R.; Bauer, M. Sensor data integration using ontologies for event detection, In: L. Barolli (Ed.), Advanced information networking and applications. Cham: Springer International Publishing, 2023. pp. 171-83.
248. Acosta, J. N.; Falcone, G. J.; Rajpurkar, P.; Topol, E. J. Multimodal biomedical AI. Nat. Med. 2022, 28, 1773-84.
249. Elbadawi, M.; Ong, J. J.; Pollard, T. D.; Gaisford, S.; Basit, A. W. Additive manufacturable materials for electrochemical biosensor electrodes. Adv. Funct. Mater. 2020, 31, 2006407.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Copyright
Data & Comments
Data
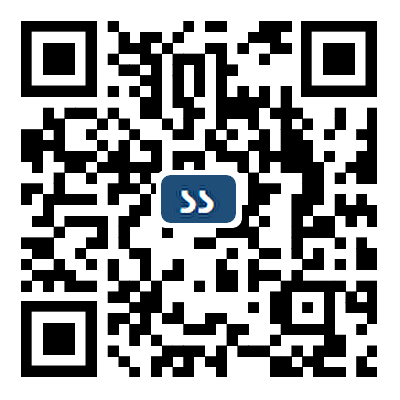







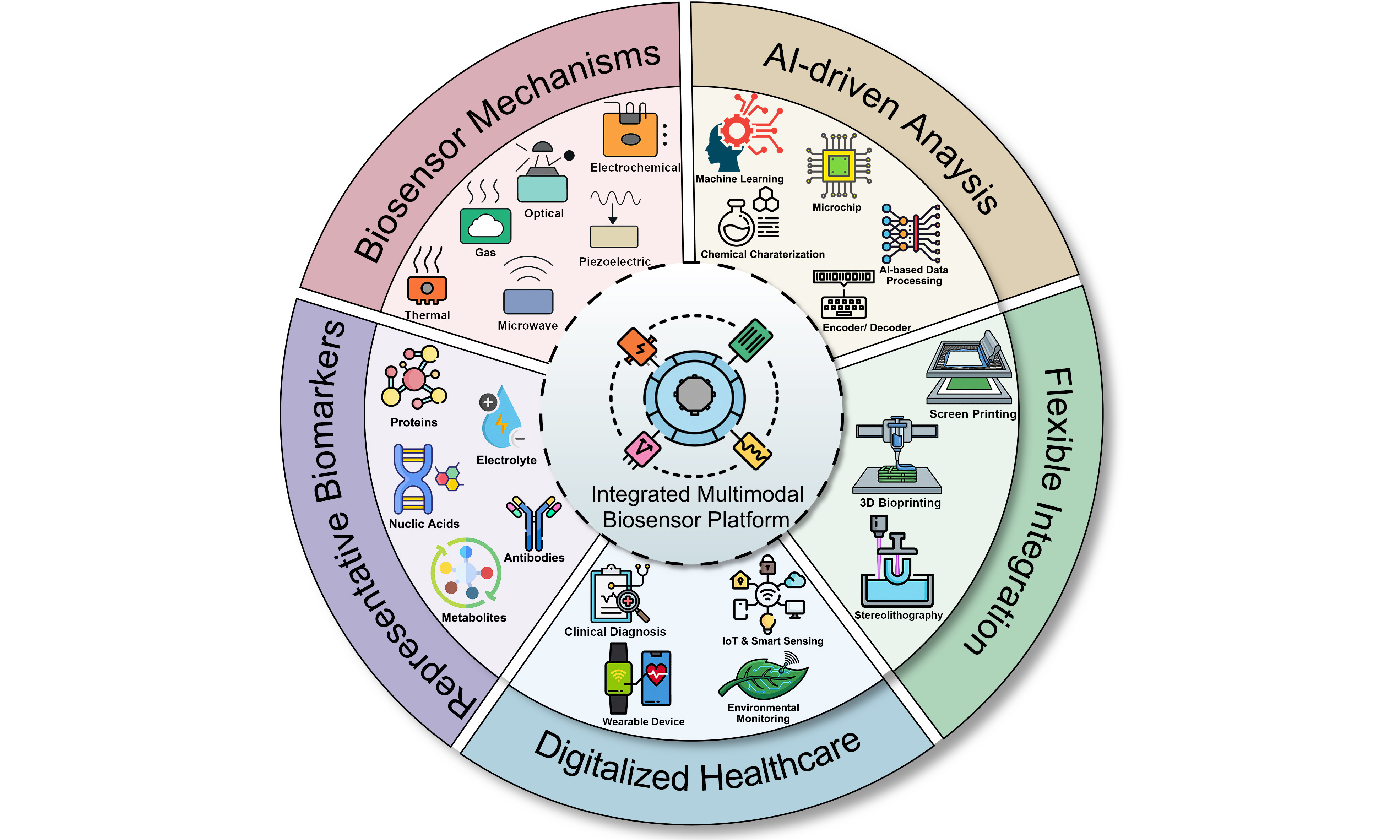
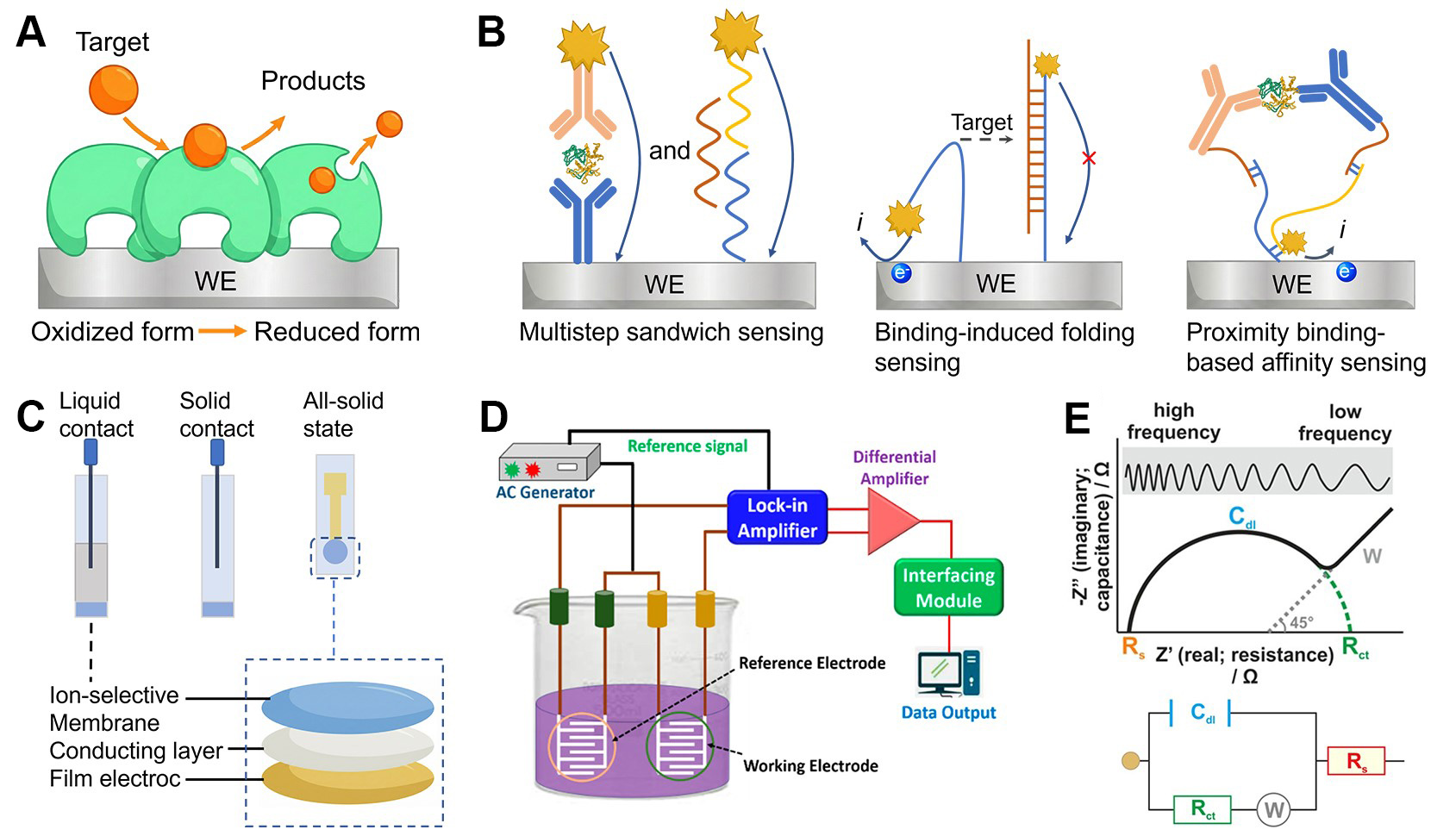
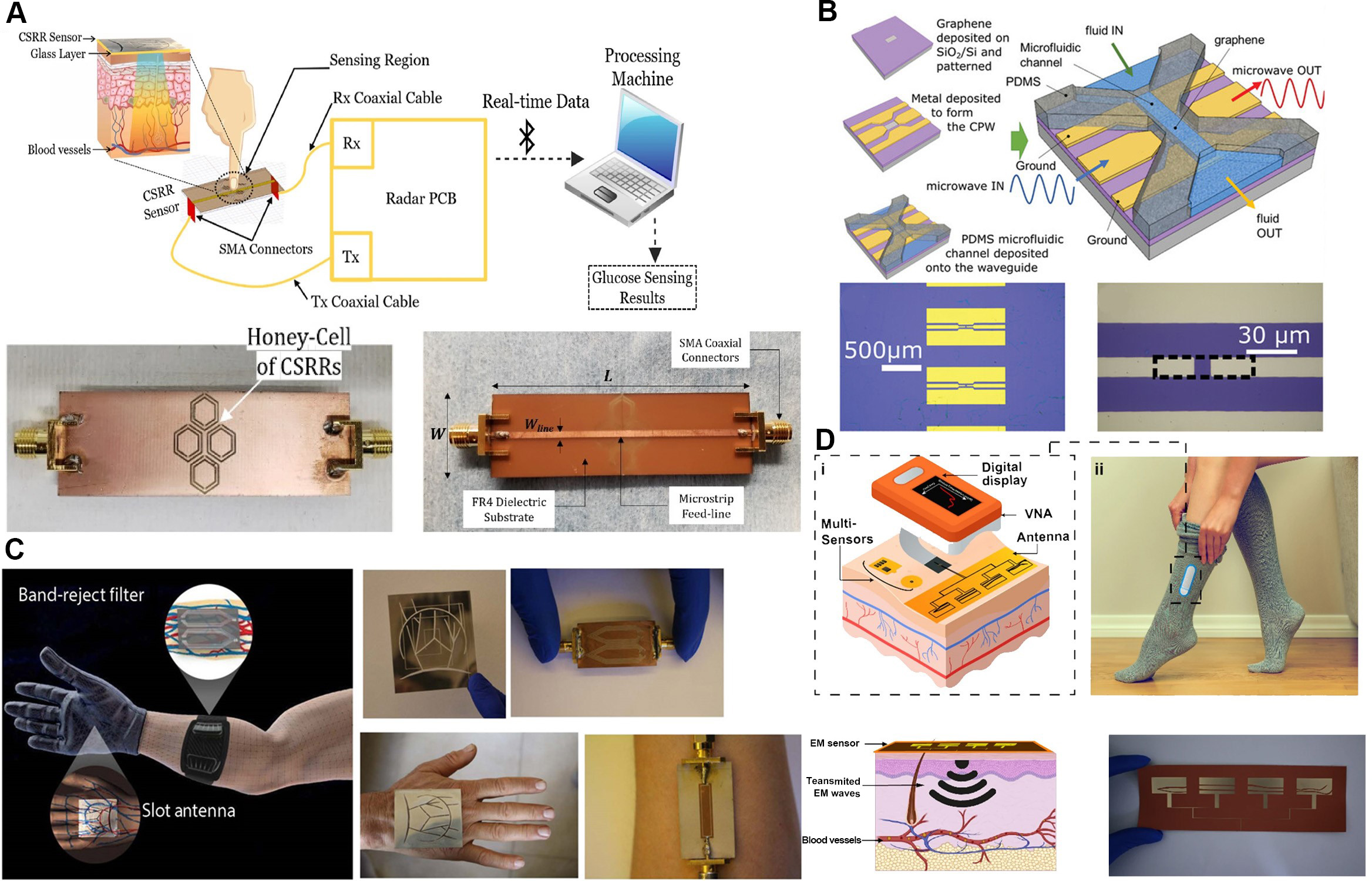
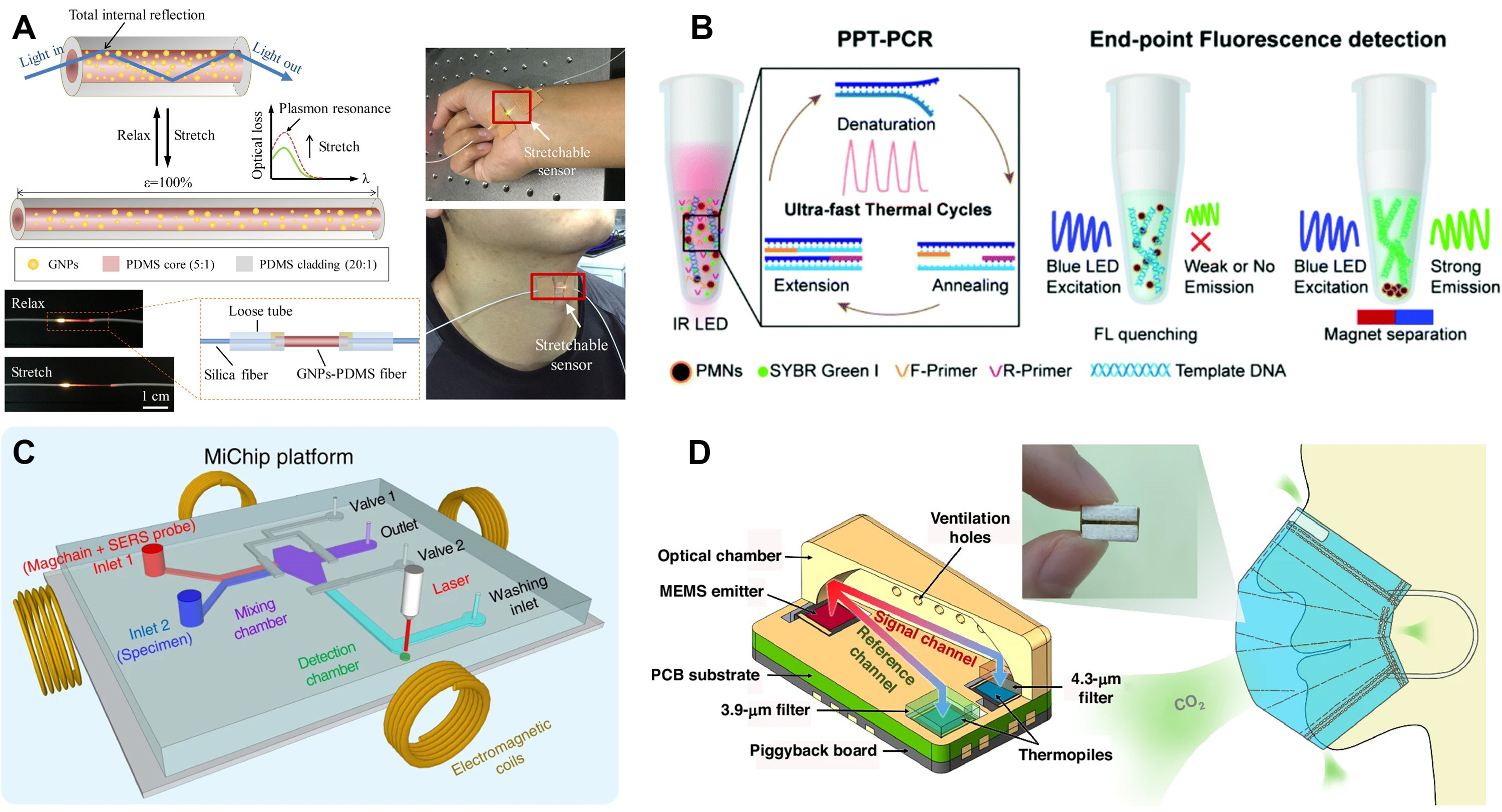
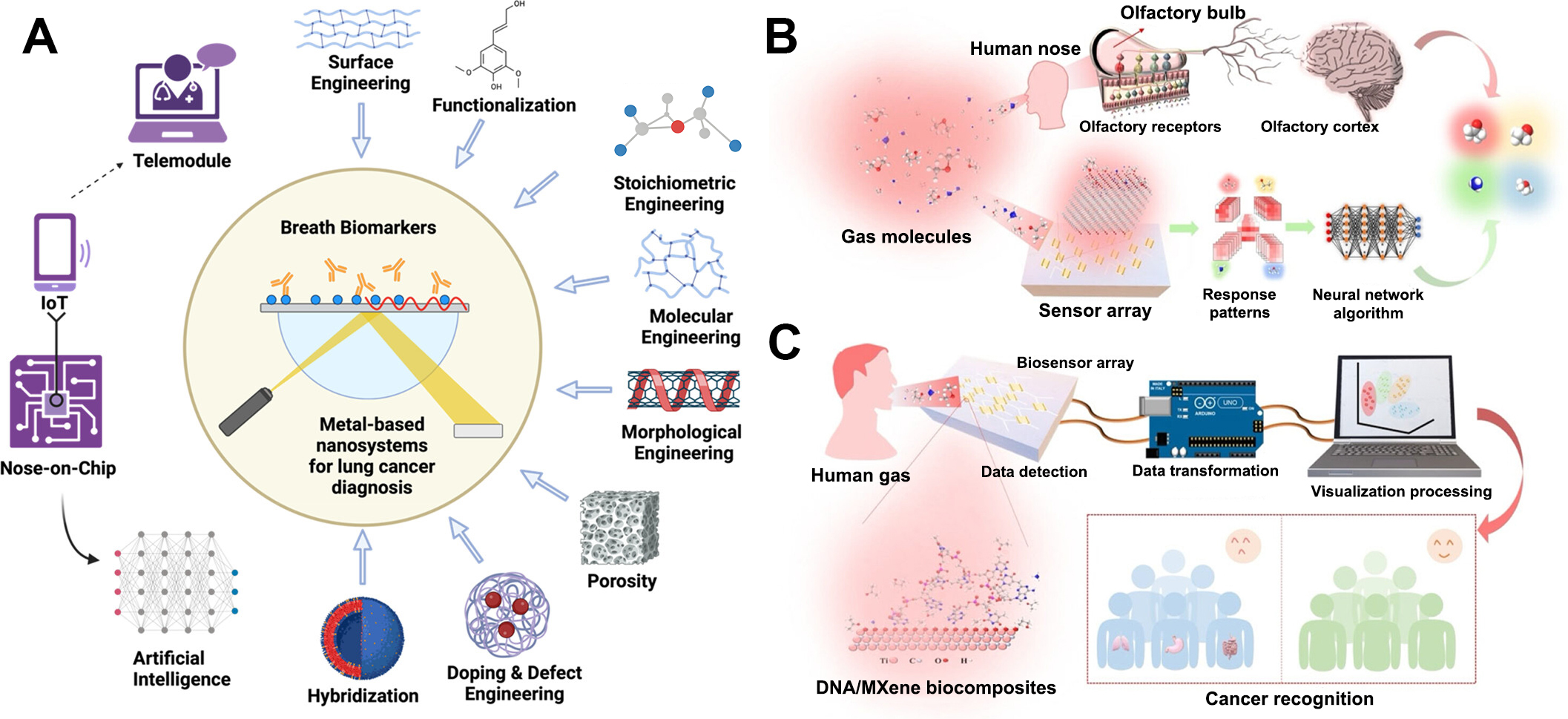
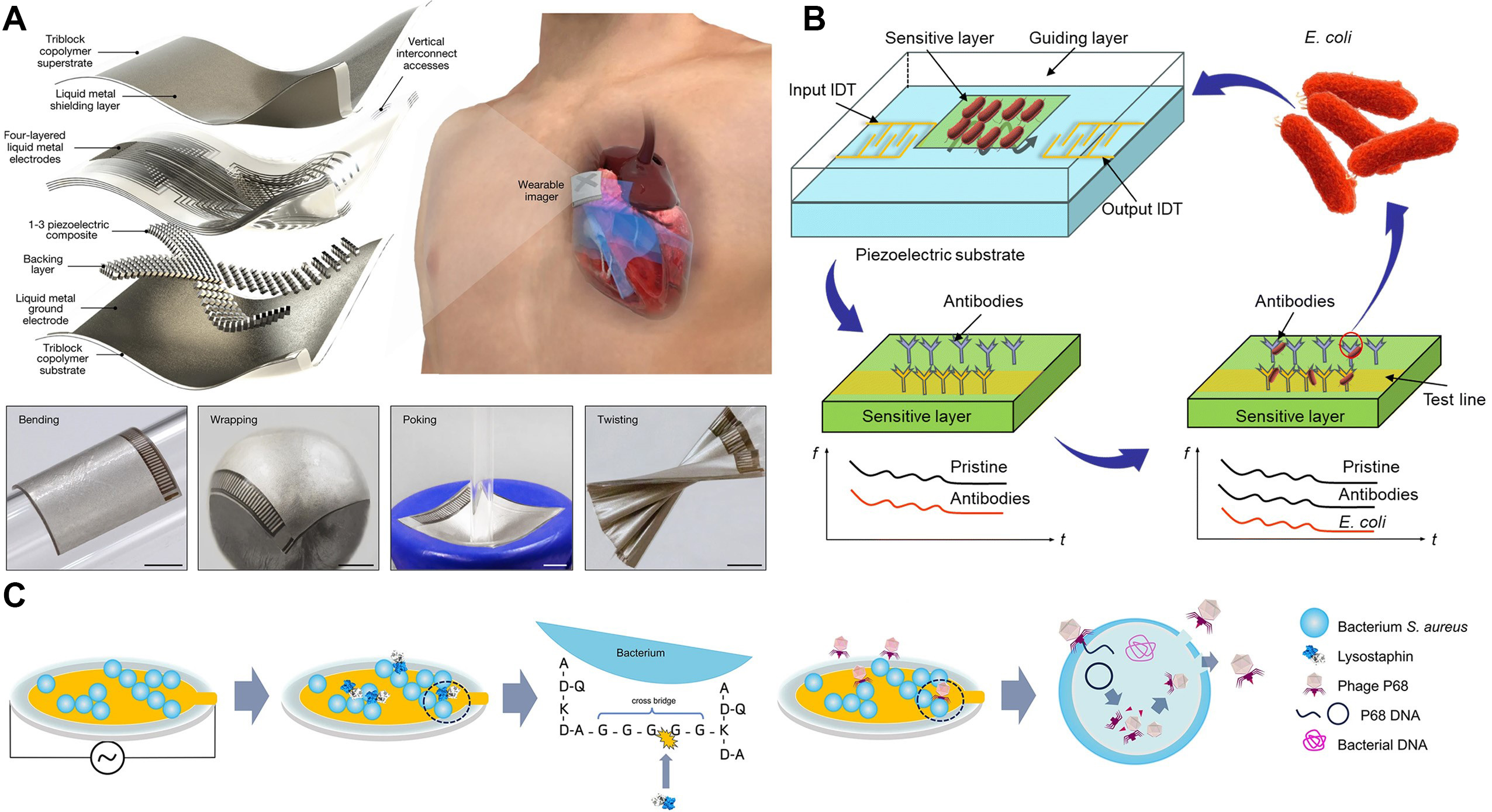
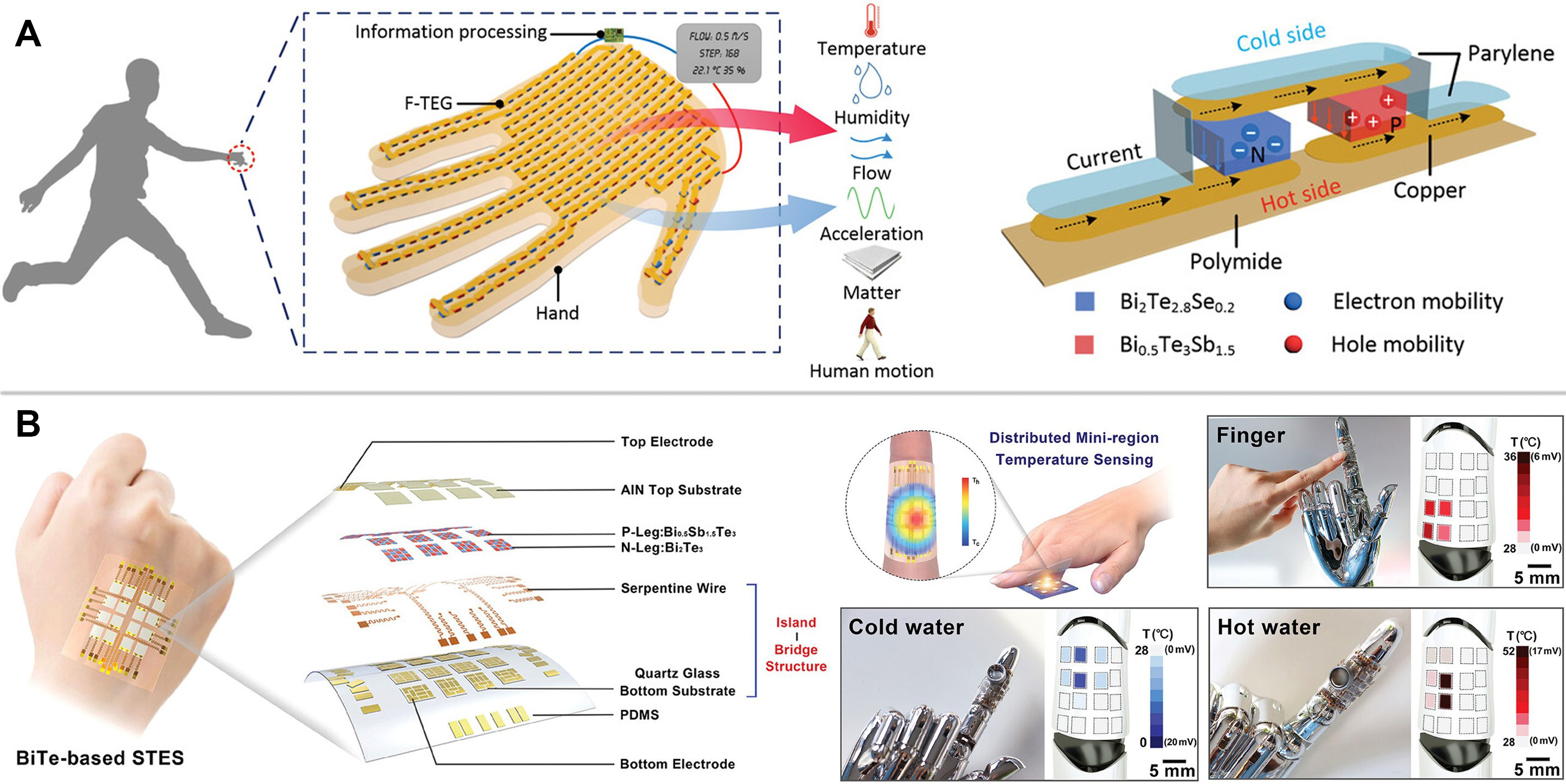
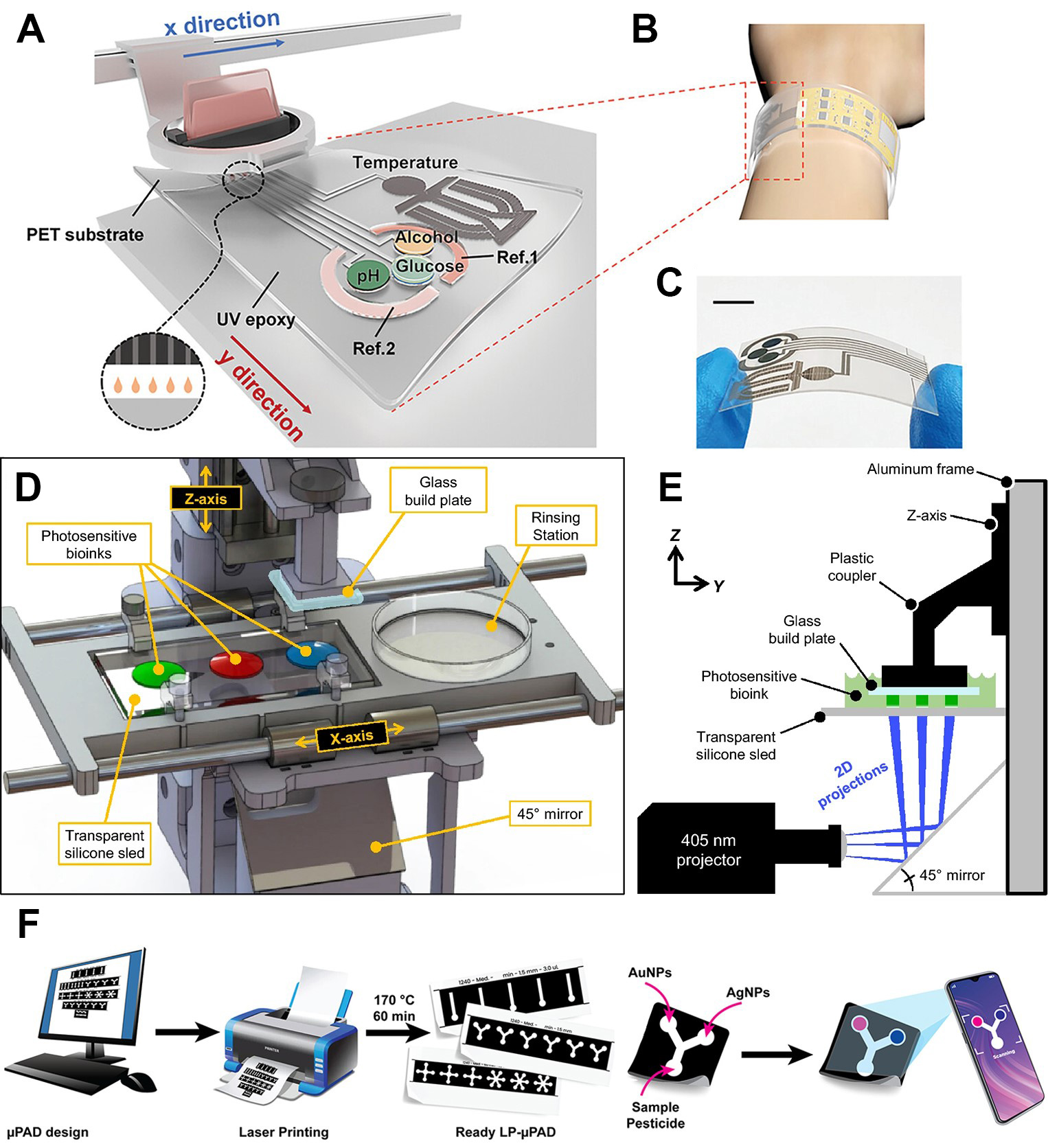
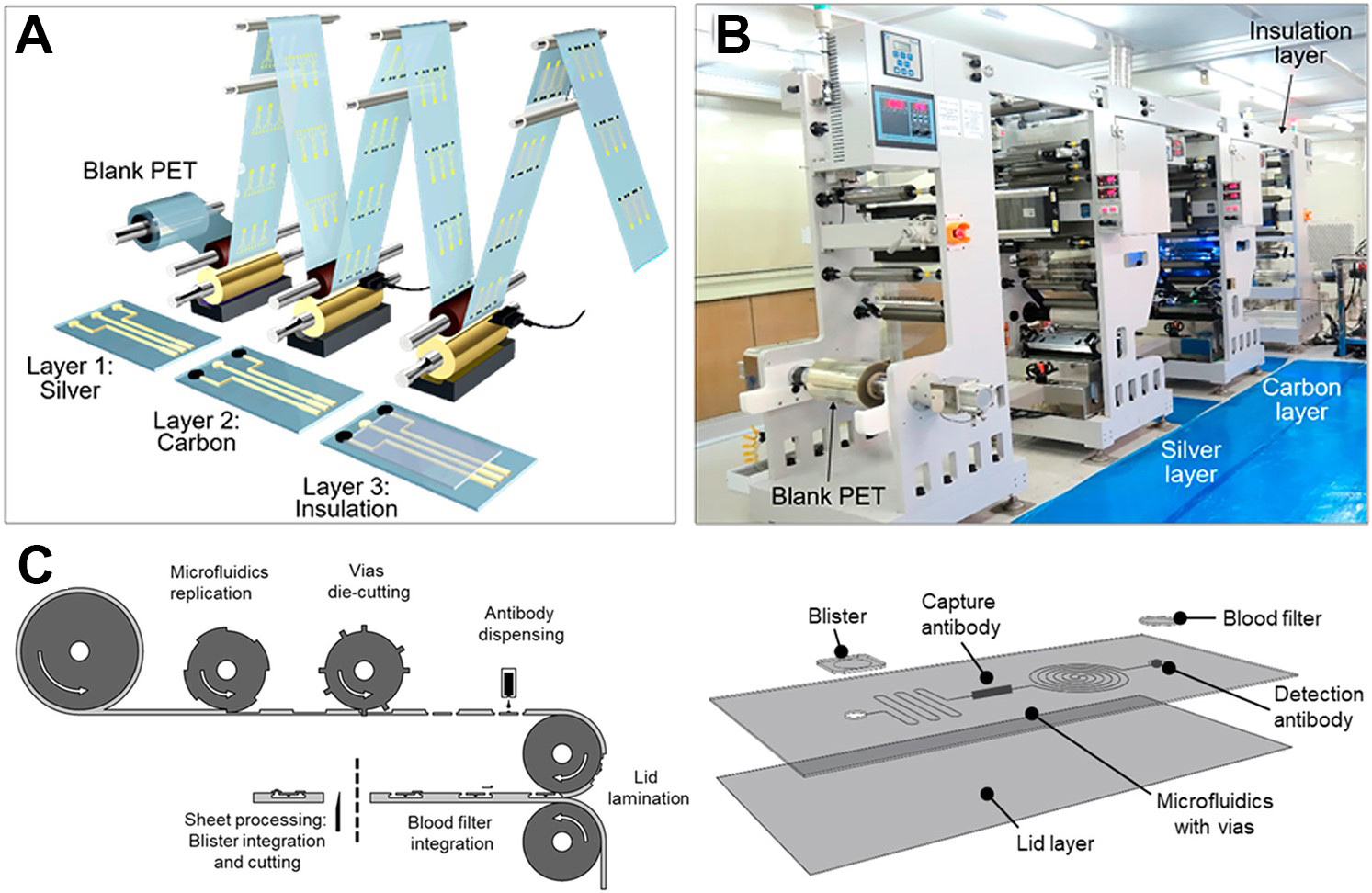
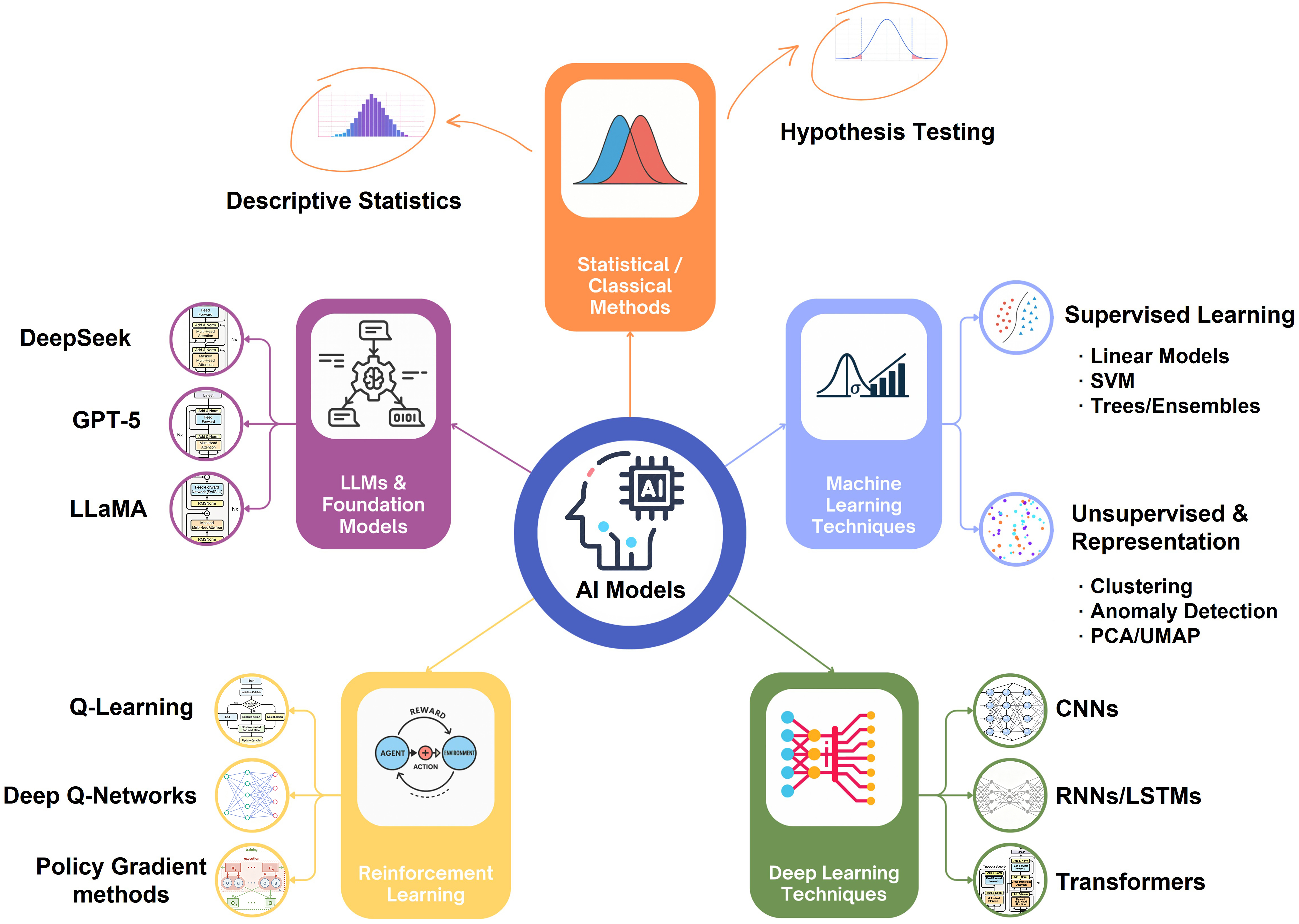
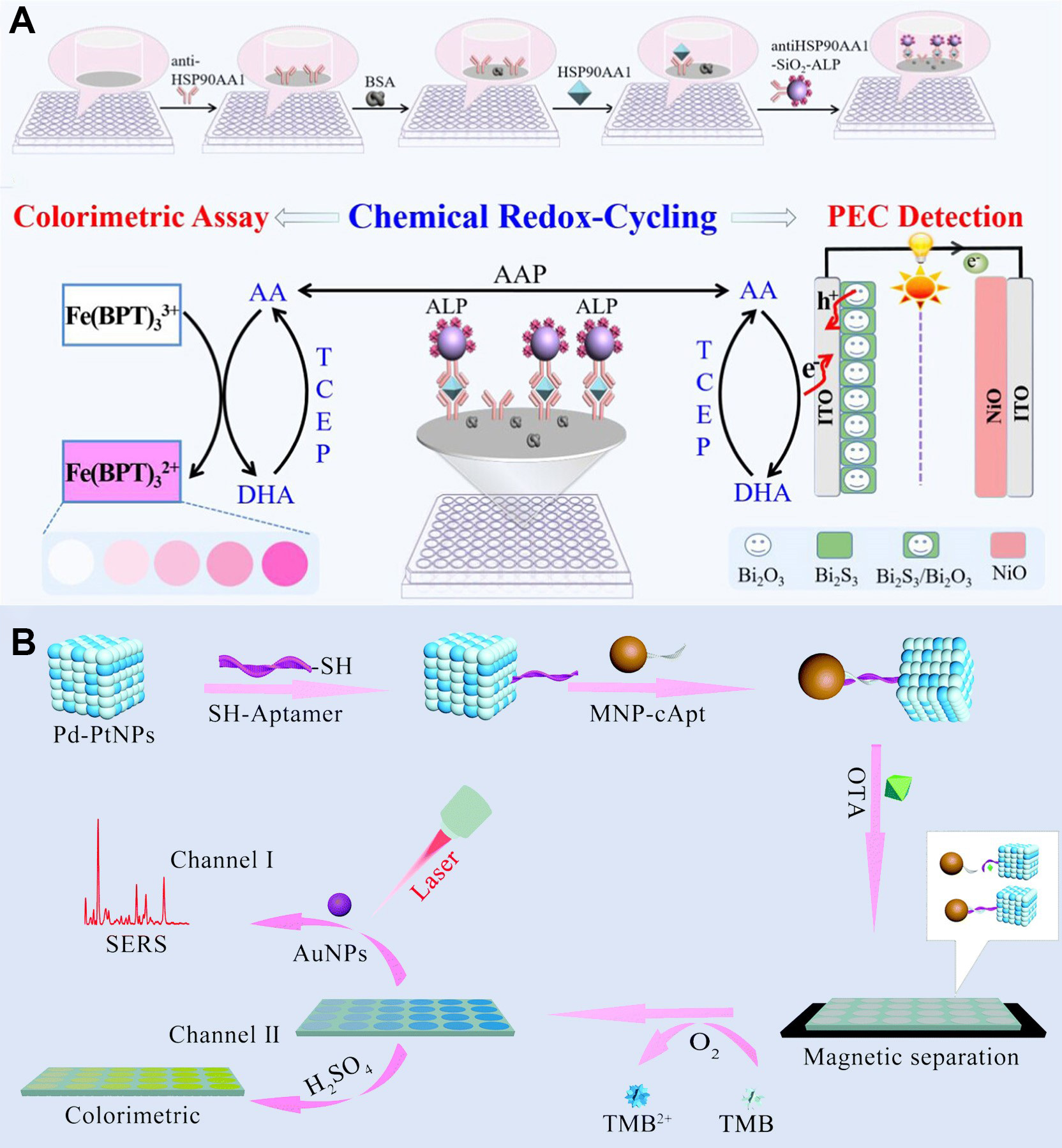
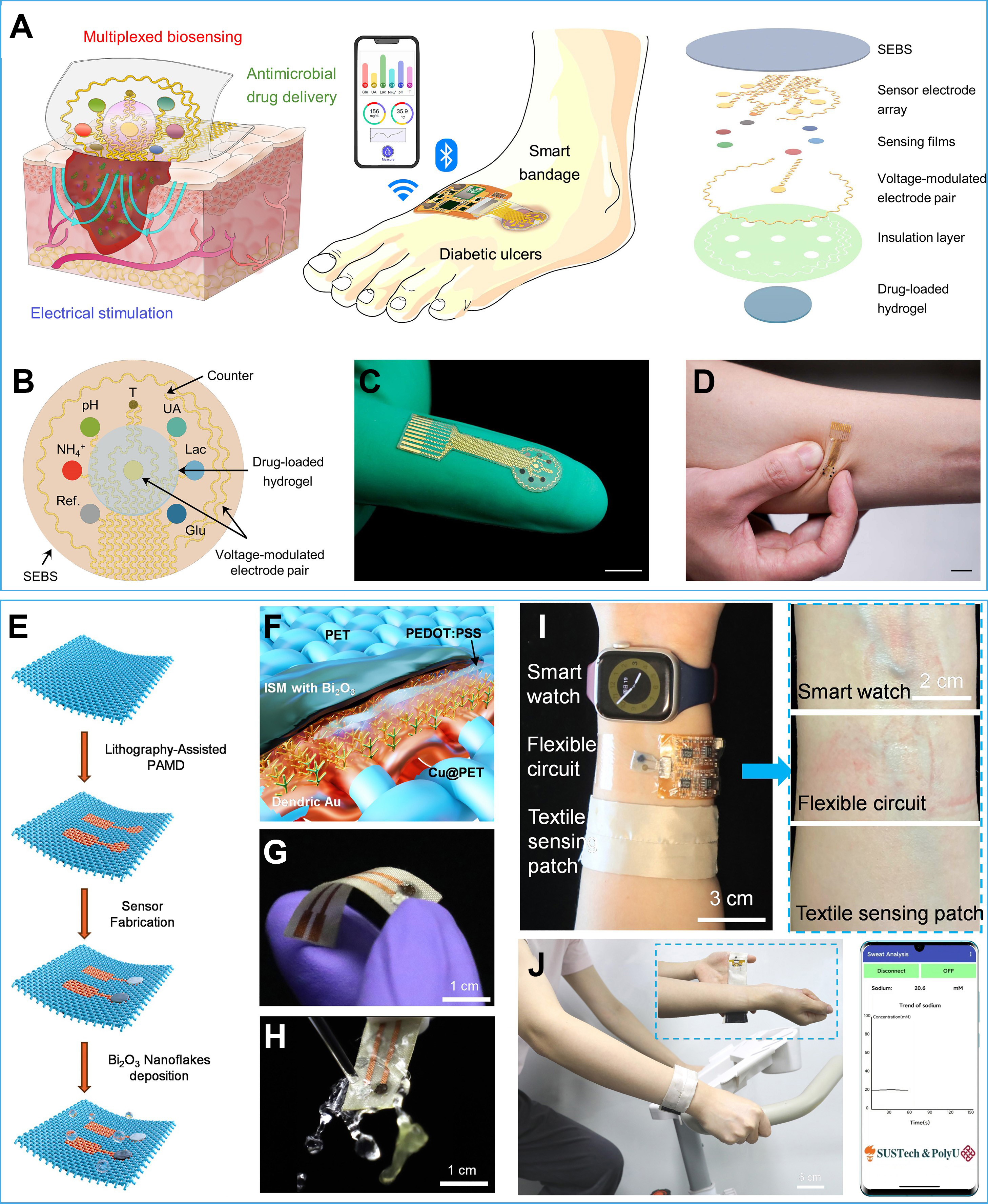
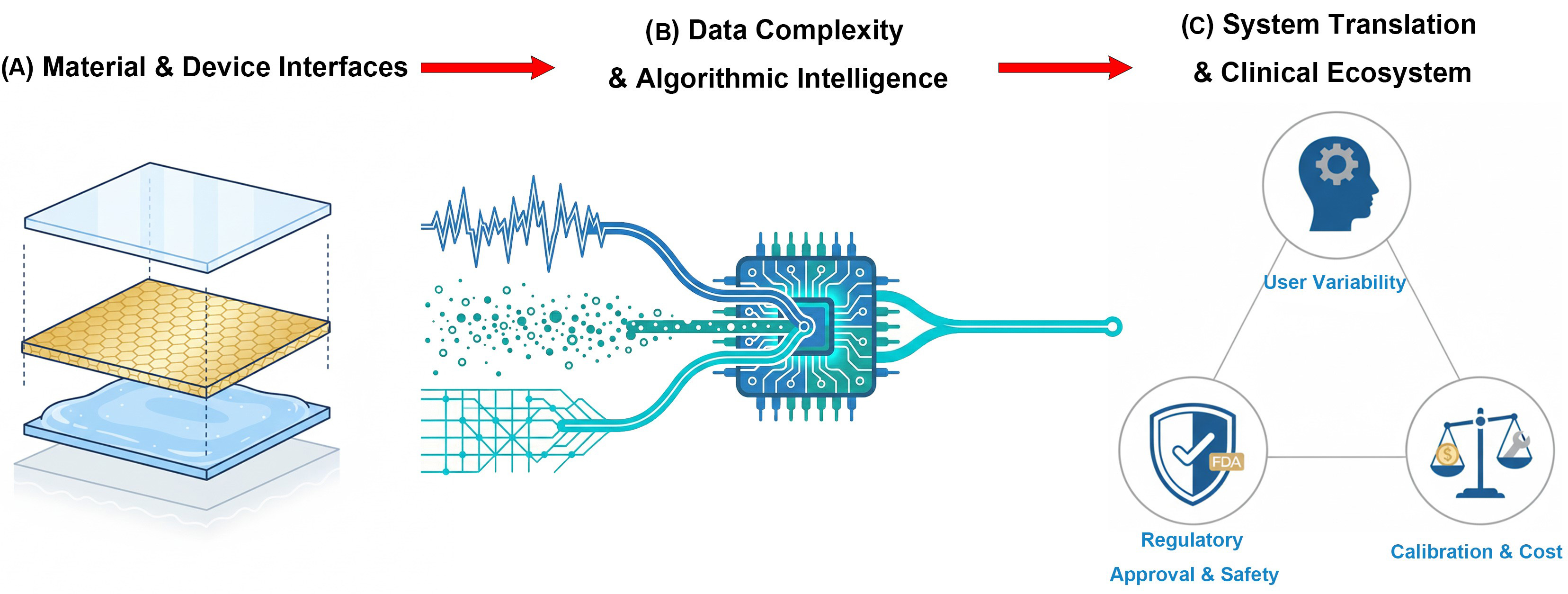












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].