fig5
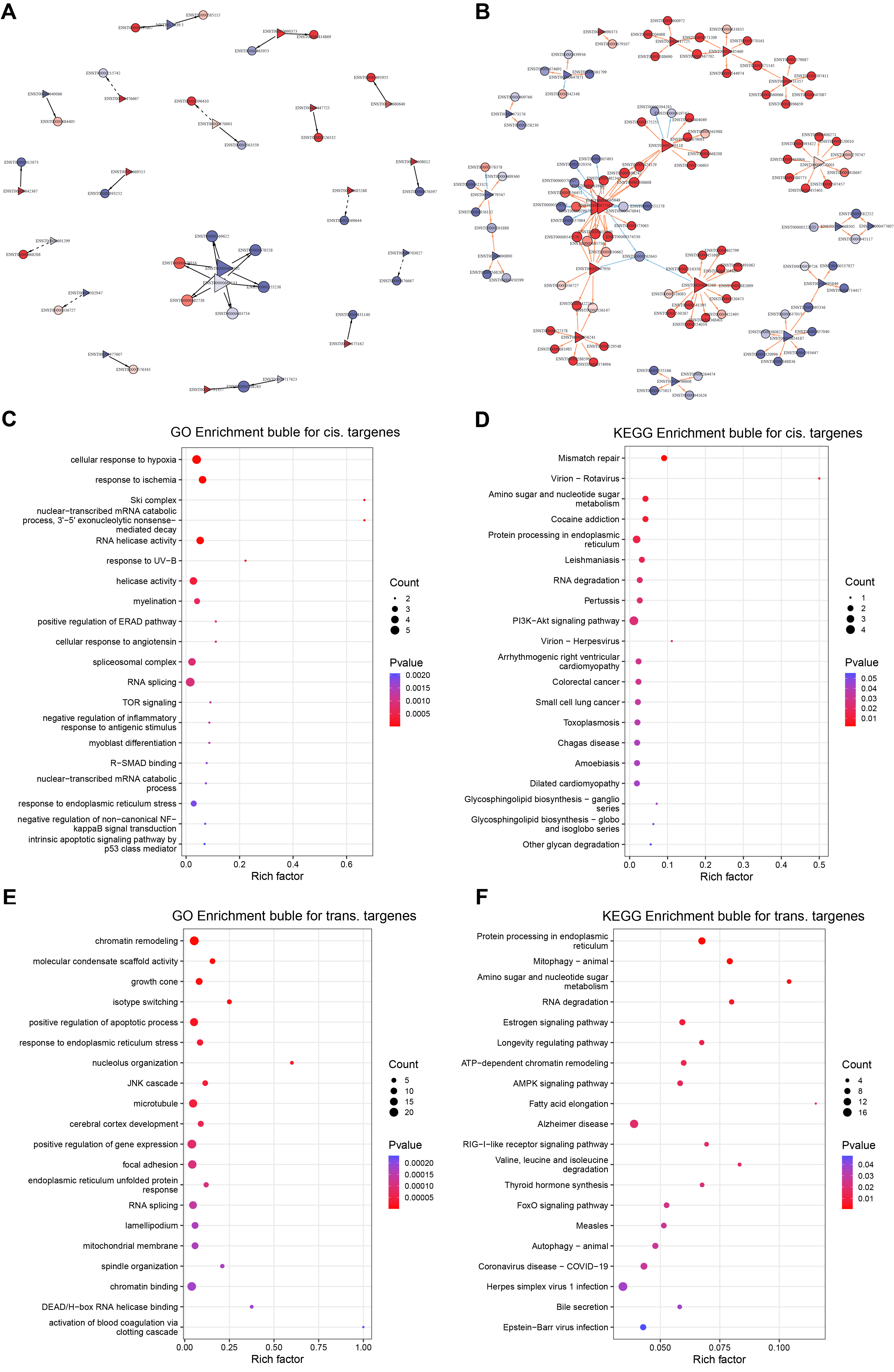
Figure 5. Predicted long non-coding RNAs (lncRNA) targets and functional enrichment profiles. (A) Cis-target regulatory network of ITGB5-responsive lncRNAs; (B) Trans-target regulatory network of ITGB5-responsive lncRNAs; (C) GO enrichment bubble plot depicting cis-target genes; (D) KEGG enrichment bubble plot depicting cis-target genes; (E) GO enrichment bubble plot depicting trans-target genes; (F) KEGG enrichment bubble plot depicting trans-target genes. DEGs: Differentially expressed genes; DETs: differentially expressed transcripts; GO: gene ontology; KEGG: kyoto encyclopedia of genes and genomes; PI3K/Akt: phosphoinositide 3-kinase/protein kinase B; UV-B: ultraviolet B; ERAD: endoplasmic reticulum-associated degradation; TOR: target of rapamycin; R-SMAD: receptor-regulated SMAD; NF: nuclear factor; JNK: c-Jun N-terminal kinase; ATP: adenosine triphosphate; AMPK: AMP-activated protein kinase; RIG: retinoic acid-inducible gene.