fig2
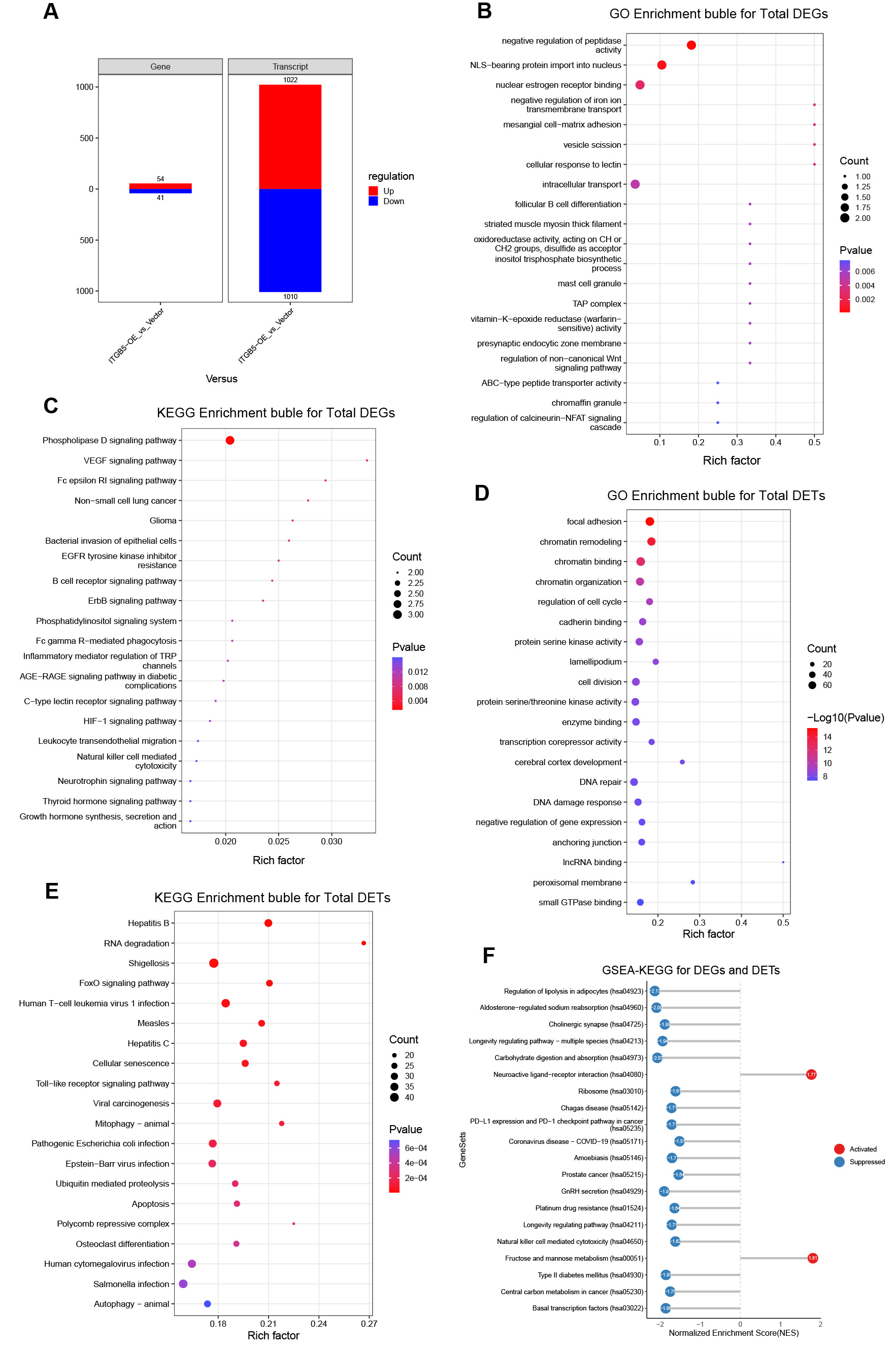
Figure 2. Functional enrichment landscapes of DEGs and DETs following ITGB5 overexpression. (A) Summary of up- and downregulated genes and transcripts; (B) Gene Ontology (GO) enrichment bubble plot for all DEGs; (C) Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment bubble plot for all DEGs; (D) GO enrichment bubble plot for all DETs; (E) KEGG enrichment bubble plot for all DETs; (F) Gene set enrichment analysis based on KEGG pathways (GSEA-KEGG) for integrated DEGs and DETs (activated pathways in red; suppressed pathways in blue). DEGs: Differentially expressed genes; DETs: differentially expressed transcripts; VEGF: vascular endothelial growth factor; TAP: transporter associated with antigen processing; NFAT: nuclear factor of activated T cells; NLS: nuclear localization signal; IRF: interferon regulatory factor; EGFR: epidermal growth factor receptor; TRP: transient receptor potential; AGE-RAGE: advanced glycation end products-receptor for advanced glycation end products; HIF-1: hypoxia-inducible factor 1; IncRNA: long non-coding RNA.