Circulating tumor DNA (ctDNA) in early-stage NSCLC: a new frontier in precision oncology
Abstract
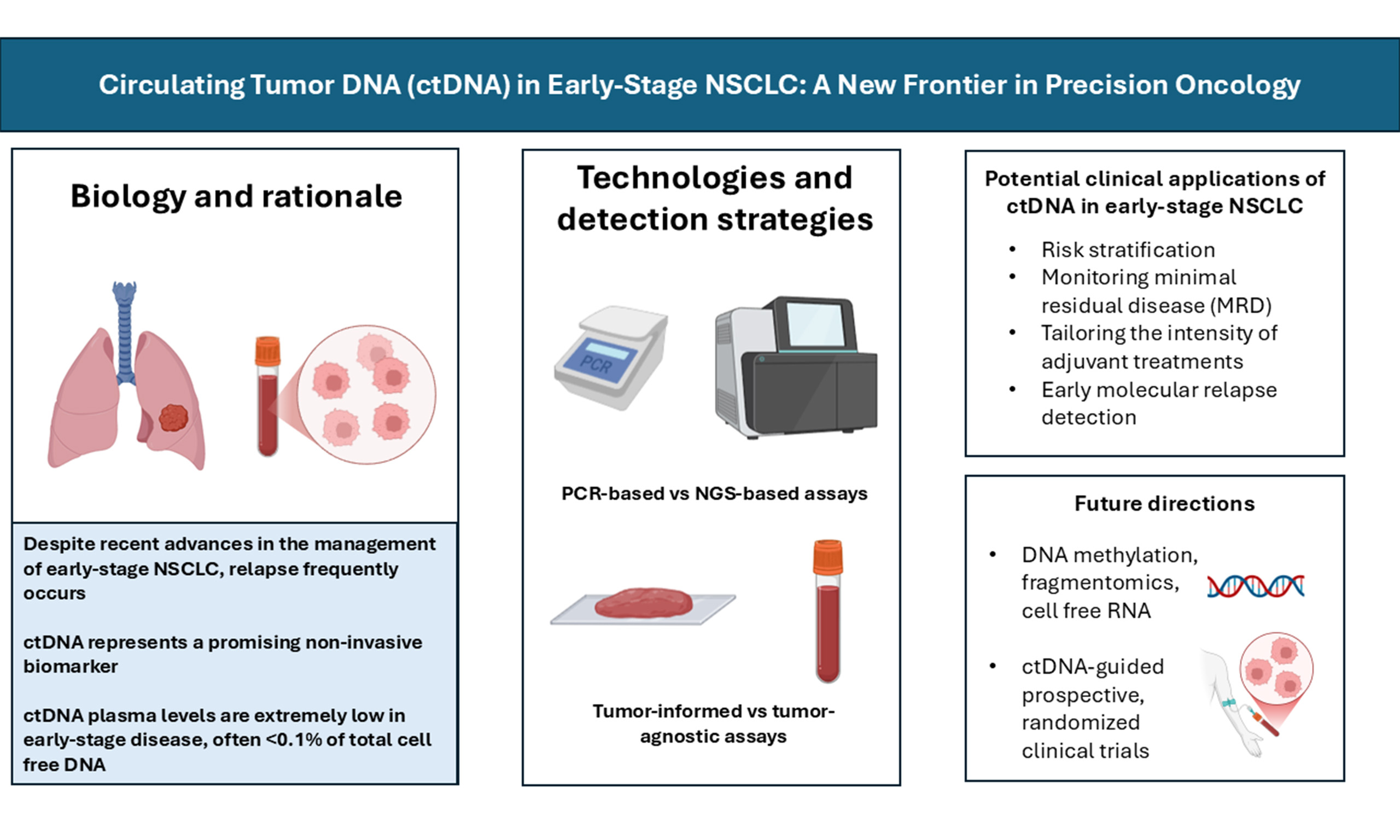
Recent advances in the management of early-stage non-small cell lung cancer (NSCLC), including perioperative immunotherapy and targeted treatments, have significantly improved outcomes. Nevertheless, a considerable proportion of patients relapse after curative-intent surgery, emphasizing the need for biomarkers that can more accurately define recurrence risk and guide individualized treatment. Circulating tumor DNA (ctDNA) has emerged as a promising, non-invasive biomarker for detecting minimal residual disease (MRD), assessing molecular response, and identifying relapse before clinical or radiologic evidence. Across retrospective and prospective studies, ctDNA detection at baseline or after treatment is consistently associated with inferior recurrence-free and overall survival, with postoperative positivity marking patients at the highest risk of relapse, often several months ahead of imaging findings. Evidence from recent pivotal trials further reinforces the prognostic significance of ctDNA dynamics in the perioperative setting. However, translation into clinical practice remains limited. ctDNA detection in early-stage disease is inherently challenging due to its low abundance and technical variability across assays. Moreover, no platform is yet validated or approved for MRD detection in this setting, and the predictive impact of ctDNA-guided treatment adaptation remains to be demonstrated. Ongoing efforts are focused on refining assay sensitivity, standardizing workflows, and conducting prospective interventional trials to establish ctDNA as a clinically actionable biomarker. This review provides an overview of current evidence, emerging technologies, and future directions toward clinical integration of ctDNA in resectable NSCLC.
Keywords
INTRODUCTION
The management of early-stage non-small cell lung cancer (NSCLC) has undergone a profound transformation in recent years, driven by the integration of either immune checkpoint inhibitors (ICIs) or targeted treatments in (neo)adjuvant setting[1-9]. Although surgical resection remains the standard curative approach for resectable stage I-III NSCLC, a substantial proportion of patients experience disease recurrence, reflecting the presence of minimal residual disease (MRD), defined as occult micrometastatic disease not detectable by conventional imaging[10-12].
Traditional clinicopathologic factors, such as tumor size and nodal status, remain insufficient to accurately predict recurrence risk or guide individualized treatment strategies[12]. This limitation emphasizes the need for sensitive and non-invasive biomarkers capable of improving risk assessment, identifying MRD, and supporting perioperative treatment decisions[13]. In the current therapeutic landscape of early-stage NSCLC, treatment selection in clinical practice is still largely based on established biomarkers, including epidermal growth factor receptor (EGFR) and anaplastic lymphoma kinase (ALK) alterations and programmed death-ligand 1 (PD-L1) expression[8,9].
Circulating tumor DNA (ctDNA) has emerged as a promising non-invasive biomarker, enabling dynamic assessment of tumor burden and molecular profile through a simple blood test. Its clinical utility is already established in advanced NSCLC, where it is used to perform molecular profiling when tissue biopsy is not feasible or insufficient[9,14,15]. In the last decade, growing interest has extended to early-stage disease, where ctDNA has been investigated as a biomarker for early detection, risk stratification, and MRD monitoring after curative-intent surgery[12]. While ctDNA-based early detection is an emerging area with potential implications for lung cancer screening[16], this review specifically focuses on its clinical utility in the perioperative management of resectable NSCLC. We integrate the most recent evidence from perioperative clinical trials, including chemoimmunotherapy and targeted therapy studies, and discuss their implications for clinical practice. Furthermore, we contextualize ctDNA within the broader liquid biopsy landscape by incorporating complementary analytes such as circulating tumor cells (CTCs). Finally, we discuss the practical challenges and potential clinical applications of ctDNA, addressing key issues related to its implementation in real-world perioperative management.
BIOLOGICAL PRINCIPLES AND TECHNOLOGIES
Biology of ctDNA in early-stage NSCLC
ctDNA is currently the most investigated biomarker among liquid biopsy analytes for its potential clinical applications in oncology. It refers to tumor-derived DNA fragments released into the bloodstream, primarily through apoptosis, necrosis, or active secretion by malignant cells[13,17,18]. In early-stage NSCLC, the amount of ctDNA is extremely low, often < 0.1% of total circulating cell-free DNA (cfDNA), compared with up to 10% or more in advanced disease[19]. This low fraction reflects the reduced tumor burden and constitutes one of the main technical hurdles for MRD detection in this setting. Data from the TRACERx study (TRAcking non-small cell lung Cancer Evolution through therapy (Rx); ClinicalTrials.gov: NCT01888601) showed median mutant allele frequencies (MAFs) of only 0.008% for stage I NSCLC, highlighting the inherent difficulty of reliable detection in small tumors[11].
Shedding of ctDNA correlates with clinical and pathological factors. Non-adenocarcinoma histology, lymphovascular invasion, and high Ki-67 expression are associated with greater ctDNA detectability, whereas small, indolent adenocarcinomas often remain ctDNA-negative[11].
Beyond tumor burden, ctDNA kinetics also contribute to the challenge of detection in early-stage disease. As ctDNA constitutes the tumor-derived fraction of circulating cfDNA, its temporal dynamics mirror the rapid turnover of cfDNA, which is cleared from the bloodstream within approximately 16 min to a few hours through renal, hepatic, and nuclease-mediated degradation[12,20]. Such kinetics confer on ctDNA the ability to capture rapid changes in tumor biology, while simultaneously requiring highly sensitive analytical technologies to ensure reliable MRD assessment.
Detection strategies and technical approaches
Analytical platforms for ctDNA detection
Detection of ctDNA relies on a range of molecular technologies that differ in analytical sensitivity, genomic breadth, and scalability. These approaches can be broadly grouped into polymerase chain reaction (PCR)-based and next-generation sequencing (NGS)-based platforms.
PCR-based methods, such as digital PCR (dPCR) and its main variants droplet digital PCR (ddPCR), and BEAMing (Beads, Emulsion, Amplification, and Magnetics), offer high analytical sensitivity and relatively fast turnaround times[21-24]. These techniques are particularly suited for detecting known mutations with high precision, often reaching limits of detection below 0.01% MAF[25-27]. However, their major limitation lies in their restricted multiplexing capacity, which makes them less suitable for MRD applications where multiple independent variants must be tracked simultaneously to minimize false negatives.
NGS technologies revolutionized ctDNA detection, with the development of several innovative approaches. Amplicon-based strategies amplify selected genomic regions to enhance sensitivity; examples include Tagged-amplicon deep sequencing (TAm-Seq), which uses a two-step amplification process to accurately detect low-frequency variants[28], and Safe-SeqS, which employs unique molecular identifiers (UMIs) to correct PCR errors, thereby lowering background noise[29]. Hybrid capture-based approaches, exemplified by CAncer Personalized Profiling by deep sequencing (CAPP-seq), provide broader genomic coverage while retaining fragment length information[30]. More recently, phased variant enrichment and detection sequencing (PhasED-seq) has been introduced as a capture-based assay that leverages clusters of mutations on the same DNA fragment to reduce both technical and biological errors, achieving detection limits in the parts-per-million[31].
Although whole-exome sequencing (WES) and whole-genome sequencing (WGS) theoretically enable broad detection of tumor genomic alterations, these untargeted approaches have been mainly explored in tissue-based analyses and remain difficult to apply in liquid biopsy due to high costs, limited input material, and complex bioinformatic requirements. Feasibility is particularly challenging in early-stage disease, where ctDNA levels are extremely low. Emerging data indicate that WGS and WES may expand ctDNA detection capabilities, although their use remains largely experimental[32,33].
Assay design: tumor-informed versus tumor-agnostic approaches
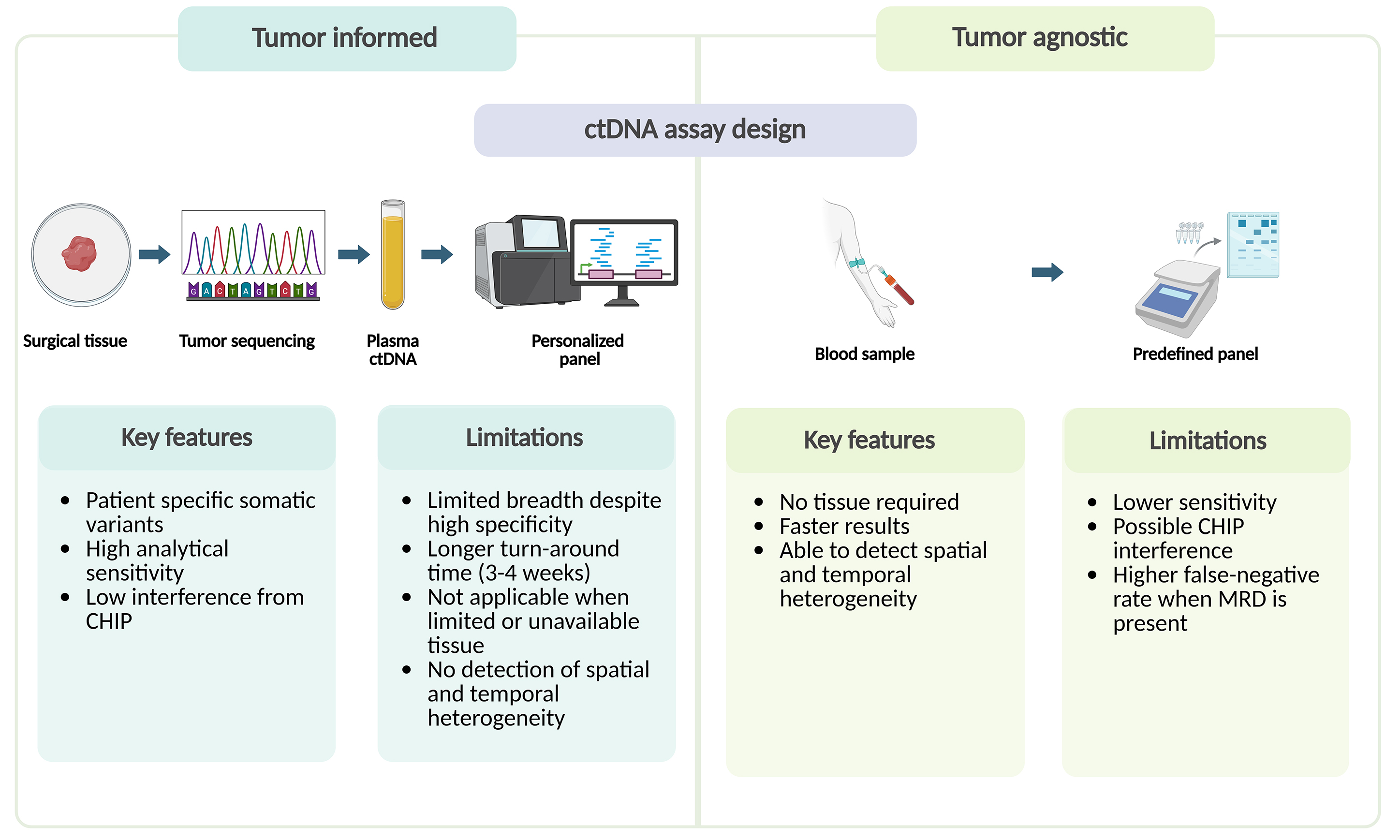
Beyond the analytical platform, ctDNA assays also differ in their design strategy, which can be either tumor-informed or tumor-agnostic[12] [Figure 1]. Tumor-informed assays involve initial sequencing of a patient’s tumor tissue to create a customized panel of somatic mutations for plasma tracking[34-38]. This strategy enhances analytical accuracy, achieving very low detection limits while reducing interference from clonal hematopoiesis of indeterminate potential (CHIP). Nonetheless, its implementation in routine clinical practice is constrained by practical issues: adequate tumor tissue must be available for sequencing, which is not always the case in early-stage disease or when only small biopsies are obtained before neoadjuvant therapy. The design of a patient-specific panel typically requires about 3-4 weeks, potentially delaying treatment decisions, and patients achieving a pathological complete response (pCR) may lack residual tumor tissue for assay generation[35]. Moreover, the number of variants incorporated in the panel directly affects assay sensitivity and the limit of detection[11,34,39]. Personalized multiplex PCR assays, such as Signatera (Natera) and RaDaR (Inivata), use tumor sequencing to identify patient-specific mutations that are subsequently tracked in plasma, reaching very high sensitivity[40,41].
Figure 1. Tumor-informed vs. tumor-agnostic ctDNA assays: key features and limitations. CHIP: Clonal hematopoiesis of indeterminate potential; ctDNA: circulating tumor DNA; MRD: minimal residual disease. Created with BioRender.com (Created in BioRender. Cammarota, A. (2026) https://BioRender.com/txdc4kj).
Tumor-agnostic (tumor-naïve) assays, by contrast, rely on predefined genomic panels that can be applied directly to plasma without prior tumor profiling[42,43]. Their main advantages are shorter turnaround time and independence from tissue availability. However, this approach is generally less sensitive and more susceptible to background noise from CHIP-related variants. From a biological standpoint, analyzing circulating DNA derived from the entire tumor burden may also enable tumor-agnostic approaches to capture both spatial and temporal genomic heterogeneity, providing a broader view of tumor evolution and disease complexity.
Other liquid biopsy analytes
Other biomarkers can be investigated through liquid biopsy in NSCLC, including CTCs and extracellular vesicles (EVs)[20]. CTCs originate from the primary tumor and/or (micro)metastatic sites by intravasation and enter the bloodstream as single cells or multicellular clusters[44]. The time CTCs remain in circulation is very short, as they are exposed to immune surveillance, anoikis, and mechanical shear forces. Within the bloodstream, multiple innate immune cell populations contribute to their rapid elimination: neutrophils exert cytotoxic effects through the release of reactive oxygen species, macrophages mediate phagocytic clearance of CTCs, and natural killer (NK) cells induce apoptosis via direct cell-cell interactions and activation of death receptor pathways, including tumor necrosis factor-related apoptosis-inducing ligand (TRAIL)-mediated signaling[45]. In parallel, hemodynamic forces such as fluid shear stress and microvascular deformation can impair membrane integrity and cytoskeletal stability, further compromising cell viability[46]. Detachment from the extracellular matrix also disrupts integrin-dependent survival signaling, including focal adhesion kinase (FAK) and phosphatidylinositol 3-kinase (PI3K)/AKT pathways, leading to activation of apoptotic cascades (anoikis), which further limits the persistence of viable CTCs in circulation[47]. On the other hand, EVs are protected by a lipid bilayer that enhances the stability of their molecular cargo, potentially allowing for greater persistence in circulation compared with cfDNA[48].
Data specifically in NSCLC are limited. Early clinical observations in patients with breast and prostate cancer undergoing surgery suggested a half-life of approximately 1-2 h, with CTC levels returning to baseline within 24 h. More recent studies using advanced animal models have demonstrated even faster clearance: around 25-30 min for single CTCs and 6-10 min for clusters[49]. Despite this limited persistence, their presence has been associated with poor clinical outcomes in several tumor types, including NSCLC, particularly when CTCs circulate as clusters exhibiting epithelial plasticity and stem-like features that enhance metastatic efficiency. CTC clusters may consist solely of tumor cells (homotypic) or include immune or stromal components (heterotypic), facilitating survival and proliferation during dissemination. Notably, interactions between CTCs and neutrophils have been shown to promote cell-cycle progression and accelerate metastatic seeding. Once disseminated, tumor cells can either proliferate into overt metastases or enter dormancy, a reversible state of long-term growth arrest influenced by intrinsic signaling pathways, such as transforming growth factor beta (TGF-β), growth arrest-specific 6 (GAS6)-AXL, and signal transducer and activator of transcription 3 (STAT3)[50]. The transition from dormancy to metastatic outgrowth remains incompletely understood and may differ between solitary and clustered disseminated tumor cells. Taken together, CTCs are key contributors to metastatic progression and relapse biology, and their enumeration and characterization may complement ctDNA-based MRD assessment by informing dissemination potential, metastatic risk, and tumor-microenvironment interactions in early-stage NSCLC. Nevertheless, despite their biological and prognostic relevance, the clinical implementation of CTCs remains limited by technical demands for enrichment and detection, and ctDNA currently represents the most widely adopted analyte for routine liquid biopsy applications.
POTENTIAL APPLICATIONS OF ctDNA IN EARLY-STAGE NSCLC
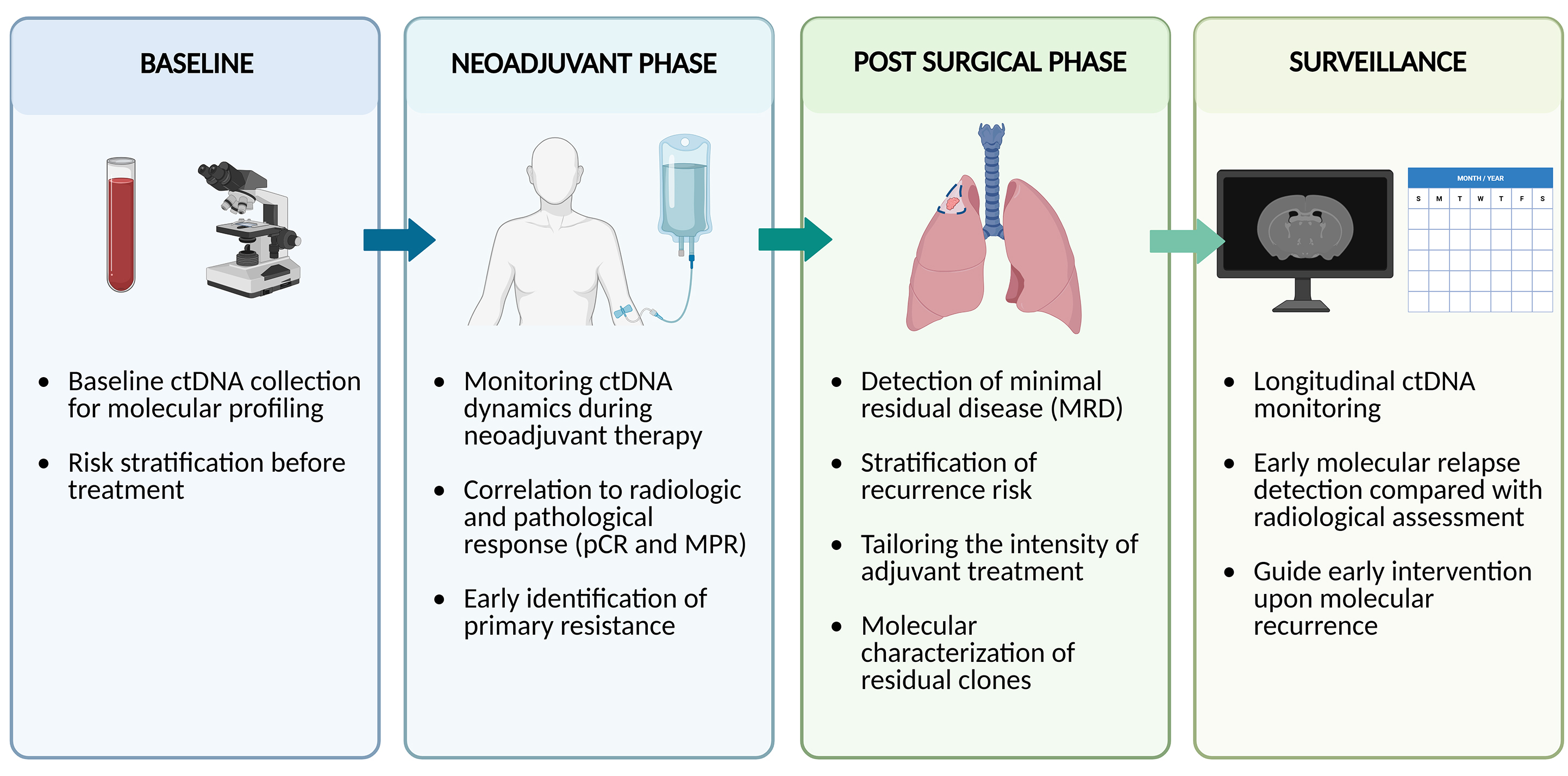
In the preoperative setting, its detection may reveal biologically aggressive disease not captured by conventional imaging or staging, potentially informing neoadjuvant treatment decisions. Postoperatively, ctDNA analysis is being explored for the identification of MRD and early relapse, thereby supporting the selection of patients who may benefit from adjuvant systemic therapy. In addition, longitudinal monitoring during or after perioperative treatment can provide early insights of therapeutic efficacy and recurrence, often preceding radiographic evidence. These applications are currently under active evaluation, with ctDNA increasingly incorporated into clinical trial designs and investigational treatment algorithms. Figure 2 summarizes the main potential clinical roles of ctDNA in early-stage NSCLC.
Figure 2. Potential clinical applications of ctDNA across perioperative management. ctDNA: Circulating tumor DNA; MPR: major pathologic response; pCR: pathologic complete response. Created with BioRender.com (Created in BioRender. Cammarota, A. (2026) https://BioRender.com/aeszxg6).
Early evidence of ctDNA in resectable NSCLC
Study characteristics and ctDNA detectability across cohorts
Over the past decade, multiple retrospective and prospective studies, with cohorts ranging from 33 to over 330 patients, have evaluated perioperative ctDNA using various platforms, most commonly NGS-based assays[11,51-68] [Table 1]. These studies generally included unselected stage I-III NSCLC patients, with the exception of the study by Jung et al., which focused on EGFR-mutant cases[61]. The study by Chaudhuri et al. also enrolled a subset of small cell lung cancer patients, and some cohorts included a minority of stage IV cases[51,53,66]. Adjuvant therapy was administered according to standard practice, while neoadjuvant chemotherapy was limited to a minority of cohorts[61,62,66]. In selected cases, definitive chemoradiotherapy or radiotherapy alone was delivered as curative-intent treatment[51,58,67].
Key retrospective and prospective studies evaluating perioperative ctDNA in resectable NSCLC
| Study | N | Presurgery ctDNA+ pts, n (%) | MRD landmark timepoint | Postsurgery ctDNA+ pts at landmark analysis, n (%) | Postsurgery ctDNA+ pts at any time, n (%) | RFS in MRD+ pts landmark/longitudinal analysis HR (95%CI) | OS in MRD+ pts landmark/longitudinal analysis HR (95%CI) | Median lead time between ctDNA detection and clinical/radiological recurrence |
| Chaudhuri et al.[51] | 40 | 37 (93.0) | 4 months | 17/32 (53.0) | 20 (54.0) | 43.4a (5.7-341.0), P < 0.001/- | 14.3 (3.2-64.1), P < 0.001/- | 5.2 months |
| Peng et al.[52] | 77 | 46 (59.7) | - | - | 30/71 (42.3) | -/2.9 (1.3-6.3), P = 0.0035 | -/3.0 (1.2-7.5), P = 0.0086 | 12.6 months |
| Li et al.[53] | 119 | 29/117 (24.8) | - | - | 37/119 (31.1) | -/3.5 (1.6-7.6), P < 0.001 | -/10.0 (1.2-85.8), P = 0.010 | 8.7 months |
| Xia et al.[54] | 330 | 69/330 (20.9) | - | - | 26/330 (7.9) | -/11.1 (6.5-19.0), P < 0.001 | - | - |
| Qiu et al.[55] | 103 | 88/91 (96.7) | ≤ 1 month and before the start of ACT | 18/85 (21.2) | 27/34 (79.0) | 4.0 (2.0-8.0), P < 0.001/8.5 (3.7-20.0), P < 0.001 | - | 88 days |
| Kuang et al.[56] | 38 | 19 (50.0) | - | - | 8/35 (22.9) | -/3.7 (CI not reported), P = 0.033 | - | - |
| Waldeck et al.[57] | 33 | 12/21 (57.0) | - | - | 4/21 (19.0) | -/0.1b,c (0.0-0.1), P = 0.013 | -/0.03a (0-0.3), P = 0.004 | 5 months |
| Gale et al.[58] | 88 | 40/78 (51.0) | ≥ 2 weeks and ≤ 4 monthsd | 10/59 (17.0) | 49/88 (56.0) | 14.8 (5.8-37.5), P < 0.001/9.8 (4.8-20.3), P < 0.001 | 5.5 (2.2-13.8) P = 0.0003/- | 212.5 days |
| Wang et al.[59] | 127 | 27/127 (21.3) | - | - | 41/127 (32.3) | -/7.6 (3.5-16.3), P < 0.0001 | - | 145 days |
| Abbosh et al.[11] | 197 | 117/178 (65.7) | ≤ 120 days | 27/108 (25.0) | 3/61 (4.9) not relapsed pts 59/70 (84.3) relapsed pts | 6.8e (3.7-12.3), P < 0.0001/- | 5.3 (2.9-9.7), P < 0.001/- | 119 days |
| Chen et al.[60] | 181 | 63/151 (41.7) | 3-7 days (timepoint B); 30 days (timepoint C) | 28/156 (17.9) (timepoint B); 19/156 (12.2) (timepoint C) | 23/110 (20.9) | 5.3f (2.7-10.5), P < 0.001 (timepoint B); 16.4 (7.8-34.3), P < 0.001 (timepoint C)/37.4 (10.8-130.2), P < 0.001 | 5.41 (1.7-16.8), P = 0.004 (timepoint B); 9.5 (3.0-29.7), P < 0.001 (timepoint C)/22.0 (2.6-188.3), P = 0.005 | 299 days |
| Jung et al.[61] | 278 | 67/278 (24.0) | 4 weeks | 16/67 (23.9) | 94/278 (33.8) | 0.5g (0.2-1.4), P = 0.11/2.7 (1.3-5.7), P = 0.03h | - | - |
| Tran et al.[62] | 85 | 48/80 (60.0) | - | - | 14/64 (21.9) | -/2.8 (1.2-6.4), P = 0.01 | -/4.0 (1.4-11.4), P = 0.0053 | - |
| Tan et al.[63] | 57 | 15/57 (26.3) | - | - | 7/57 (12.3) | -/16.1 (1.6-158.9), P < 0.0001 | - | 2.8 months |
| Ohara et al.[64] | 46 | 27/46 (58.7) | ≥ 51 days | 6/45 (13.3) | 13/46 (28.3) | 4.2 (1.6-11.0), P = 0.0018/9.8 (3.9-25.0), P < 0.0001 | 14.4 (3.8-37.0), P < 0.0001/4.9 (1.8-13.0), P = 0.00056 | 3.2 months |
| Martin et al.[65] | 108 | - | ≤ 6 months | 4/69 (5.8) | 12/108 (11.1) | 53.0 (5.5-513.0), P < 0.0001/27.0 (5.6-127.0), P < 0.0001 | - | 28 weeks |
| Oh et al.[66] | 36 | - | ≤ 6 months and before the start of ACT | 2/26 (7.7) | 9/36 (25.0) | 15.0 (1.0-253.0), P = 0.010/HR not reported, P < 0.0001 | - | 1.2-20.7 weeks |
| Schuurbiers et al.[67] | 218 | 82/165 (49.7)i | ≤ 2 weeks and ≤ 4 months | 18/141 (12.8) | 45/193 (23.3) | 12.4 (6.6-23.2), P < 0.001/ 11.4 (7.0-18.7), P < 0.001 | 6.5 (3.3-13.1), P < 0.001/ 8.1 (4.6-14.2), P < 0.001 | 206 days |
| Zhang et al.[68] | 261 | 95/261 (36.4) | 1 months | 23/261 (8.8) | 69/261 (26.4) | 0.1b,f (0.0-0.4), P < 0.001/0.1 (0.0-0.1), P < 0.001 | - | 5.2 months |
Across studies, presurgical ctDNA was detectable in approximately 20%-60% of patients, with higher rates in stage II-III compared with stage I disease, and consistently lower sensitivity in adenocarcinoma compared with squamous histology. In TRACERx, detection reached 82% (78/85) in non-adenocarcinoma vs. 42% (39/93) in adenocarcinoma and was also associated with a heavier smoking history and a higher risk of occult nodal disease[11].
Within the TRACERx study, Abbosh et al. conducted a phylogenetic ctDNA analysis in 96 patients with resected early-stage NSCLC, using patient-specific multiplex PCR panels informed by multiregion exome sequencing of the primary tumor[34]. Preoperative ctDNA was detected in 48% of patients and was independently associated with non-adenocarcinoma histology, lymphovascular invasion, and high Ki-67 expression. In ctDNA-positive cases, clonal variants were consistently identified at high frequency, while subclonal variants were detected in about two-thirds of patients, reflecting the heterogeneity of tumor evolution. ctDNA detection preceded radiographic relapse by a median of 70 days, underscoring its ability to anticipate recurrence and capture clonal dynamics beyond imaging.
Updated TRACERx results, with a median follow-up of 4.6 years in event-free patients, further refined the prognostic role of ctDNA[11]. Among 187 evaluable patients, preoperative ctDNA detection was higher in non-adenocarcinoma (92%) than in adenocarcinoma (42%) and, in the latter, was associated with occult mediastinal nodal involvement (P = 0.001). Postoperative ctDNA was detected in 84% of patients with recurrence or incomplete resection, while false negatives were mainly observed in those lacking sampling near relapse or with isolated brain metastases, where ctDNA shedding is limited. At a postoperative landmark analysis (within 120 days after surgery), ctDNA was detectable in 25% of patients and was strongly predictive of recurrence (positive predictive value 93%, negative predictive value 68%, sensitivity 49%), with positivity increasing by stage (12% in stage I, 23% in stage II, 44% in stage III; P = 0.008). Landmark ctDNA positivity was associated with a higher risk of death (hazard ratio [HR] 5.3, 95% confidence interval [CI]: 2.9-9.7; P = 1 × 10-9) and recurrence (HR 6.8, 95%CI: 3.7-12.3; P = 6 × 10-13). During serial surveillance, 20% of landmark-negative patients later became ctDNA-positive before or at relapse, with a median lead time of approximately one year and detectable ctDNA levels as low as 0.02%.
Prognostic role of ctDNA and implications for postoperative risk stratification
The presence of detectable ctDNA before treatment consistently demonstrated prognostic value across studies, showing a significant association with poorer outcomes. Pre-treatment ctDNA positivity was associated with inferior recurrence-free survival (RFS) (HRs 2.2-4.2) and overall survival (OS) (HRs 2.0-5.5). In studies incorporating multivariable analyses, baseline ctDNA remained an independent prognostic factor for RFS and OS in two cohorts[52,54], while in another the association did not reach statistical significance[67], underscoring the heterogeneity of results across studies.
Post-treatment ctDNA assessment has been performed using either landmark analyses, which assess prognostic value at predefined postoperative timepoints to minimize immortal-time bias[69], or serial sampling enabling longitudinal monitoring. At postoperative landmark analyses, ctDNA positivity was reported in approximately 5%-25% of patients across studies. Detection at this prespecified timepoint was consistently associated with poor outcomes, with HRs for RFS ranging from 4.0 to 53.0, and for OS from 3.0 to 27.0.
Longitudinal monitoring identified ctDNA positivity in up to ~11%-56% of patients during follow-up. Across studies, longitudinal ctDNA positivity was associated with a significantly increased risk of recurrence, with HRs for RFS between 2.7 and 16.1, and for OS ranging from 2.9 to 11.4. In studies incorporating multivariable analyses, postoperative ctDNA positivity consistently emerged as an independent prognostic factor for survival outcomes[54,55,59-61,64,67,68].
Across these studies, ctDNA was investigated as an early biomarker of recurrence risk. Reported median lead times between ctDNA detection and radiological or clinical relapse ranged from less than 1 month to about 12 months, reinforcing its value as a tool for the earlier identification of molecular relapse compared with conventional imaging.
Of note, a limited number of studies observed that some patients who were ctDNA-negative after surgery subsequently relapsed with brain-only disease. This observation may reflect the limited negative predictive value of ctDNA and suggests that intracranial recurrence may be less reliably captured by plasma ctDNA assays[11,57,64,70].
Several studies have explored the prognostic implications of ctDNA status in the context of adjuvant chemotherapy, examining both dynamic changes before and after treatment as well as its potential to refine risk stratification beyond conventional clinicopathological factors. Although based on limited cohorts, findings have been consistent in showing that adjuvant chemotherapy was associated with longer RFS in ctDNA-positive patients (HRs 0.2-0.34), whereas no clear benefit was observed in ctDNA-negative patients, with some analyses even suggesting worse outcomes in this subgroup[54-56,60,68,70].
Several caveats should be considered when interpreting these findings. First, most of the studies were conducted in relatively small cohorts, which limits the statistical power and generalizability of the results. Moreover, the enrolled populations were heterogeneous, with differences in stage distribution and histologic subtype, both of which are known to impact ctDNA shedding and detectability. Additional variability derived from the definition of landmark timepoints and the frequency of blood sampling, as well as from the use of diverse assay platforms with different sensitivities and analytical thresholds. Moreover, indirect comparisons are further limited by heterogeneous preanalytical conditions, including sample handling, processing intervals, and cfDNA extraction, all of which can markedly affect cfDNA yield and ctDNA detection[71]. Taken together, these factors preclude direct comparison across studies, and the reported ranges of detection rates and hazard ratios should therefore be interpreted with caution.
Emerging role of ctDNA in practice-changing clinical trials for resectable NSCLC
As perioperative treatment strategies for resectable NSCLC evolve, ctDNA has increasingly been integrated into clinical trial designs to evaluate molecular response, MRD, and relapse risk. Exploratory biomarker analyses from several pivotal trials have provided important insights into the prognostic role of ctDNA dynamics in this setting. The main trials incorporating ctDNA analyses are summarized in Table 2.
Key practice changing clinical trials evaluating ctDNA as a biomarker in patients treated with (neo)adjuvant chemoimmunotherapy
| Study | N (overall population) | Setting | Stage | Drug | ctDNA approach strategy; assay | Survival outcomes according to baseline ctDNA status (ctDNA+ vs. ctDNA-), HR (95%CI) | Survival outcomes according to ctDNA clearance after neoadjuvant therapy (pre-surgery), HR (95%CI) | Patients with ctDNA clearance achieving pCR, n (%) |
| CheckMate 816[35] | 358 | neoadjuvant | IB to IIIA (AJCC 7th ed.) | Neoadjuvant nivolumab + CT vs. neoadjuvant CT alone | Tumor informed; ArcherDX Personalized Cancer Monitoring | - | 0.4 (0.2-1.0) for OS | 46% (11/24) (Nivolumab arm) vs. 13% (2/15) (CT alone arm) |
| NADIM[42] | 46 | perioperative | IIIA (AJCC 7th ed.) | Neoadjuvant nivolumab + CT followed by nivolumab for 1 year | Tumor agnostic; Oncomine Pan-Cancer Cell-Free assay | 4.3 (1.0-17.8), P = 0.0089 for PFS; 4.4 (0.9-21.1), P = 0.016 for OS | 3.1 (0.7-12.9), P = 0.079 for PFS; 6.9 (1.3-36.8), P = 0.0070 for OS | - |
| NADIM II[43] | 86 | perioperative | IIIA-B (AJCC 8th ed.) | Neoadjuvant nivolumab + CT followed by adjuvant nivolumab for 6 months vs. neoadjuvant CT alone | Tumor agnostic; TruSight Oncology 500 ctDNA NGS assay | 0.4 (0.2-0.9) for PFS; 0.2 (0.1-0.6) for OS | - | - |
| AEGEAN[36] | 802 | perioperative | IIA-IIIB (N2) (AJCC 8th ed.) | Neoadjuvant durvalumab + CT followed by adjuvant durvalumab for 12 cycles vs. neoadjuvant CT alone followed by adjuvant PBO | Tumor informed; Invitae Personalized Cancer Monitoring | - | Durvalumab arm: 0.3 (0.1-0.5); PBO arm: 0.5 (0.3-0.8), for EFS | - |
| CheckMate 77T[72] | 735 | perioperative | IIA-IIIB (N2) (AJCC 8th ed.) | Neoadjuvant nivolumab + CT followed by adjuvant nivolumab for 12 cycles vs. neoadjuvant CT alone followed by adjuvant PBO | Not reported | - | Nivolumab arm: ctDNA clearance + pCR vs. no ctDNA clearance + no pCR: 0.2 (0.1-0.7) for EFS | Nivolumab arm: 50% (25/50); PBO arm: 12% (3/24) |
| IMpower010 (2022;2024)[38,73] | 1,005 (600 ctDNA evaluable) | adjuvant | IB ≥ 4 cm-IIIA (AJCC 7th ed.) | Adjuvant atezolizumab vs. PBO | Tumor informed; Signatera (Natera) RUO test | - | - | - |
Role of ctDNA in neoadjuvant chemoimmunotherapy trials
The phase III CheckMate 816 trial established neoadjuvant chemoimmunotherapy as a new standard of care in resectable NSCLC, demonstrating significant improvements in pCR and event-free survival (EFS) with the addition of nivolumab to platinum-based chemotherapy[1]. An exploratory analysis within this study investigated the clinical relevance of ctDNA as a prognostic biomarker, using a tumor-informed panel for WES (ArcherDX Personalized Cancer Monitoring)[35]. Among the 358 patients enrolled, plasma samples suitable for ctDNA analysis were available for 43 (24.0%) patients in each treatment arm.
In the nivolumab plus chemotherapy arm, ctDNA clearance after neoadjuvant treatment occurred in 56% (24/43) of patients with baseline ctDNA positivity, compared to 35% (14/43) in the chemotherapy group. A higher proportion of patients achieving ctDNA clearance also had a pCR in the nivolumab plus chemotherapy group (46%) compared with the chemotherapy arm (13%). Furthermore, ctDNA dynamics provided prognostic information beyond pathologic assessment. Patients in the chemoimmunotherapy arm who cleared ctDNA experienced markedly improved long-term outcomes, with a 5-year OS rate of 75%, compared to 52.6% in those with residual ctDNA (HR 0.38, 95%CI: 0.15-1.00). Similarly, the 5-year EFS rate was higher in patients with ctDNA clearance (51.6% vs. 40.1%; HR 0.45, 95%CI: 0.18-1.11), reinforcing the prognostic value of ctDNA kinetics in the neoadjuvant setting.
These results indicate that ctDNA clearance after neoadjuvant therapy may provide additional prognostic information in resectable NSCLC, complementing established markers such as pathological assessment.
Role of ctDNA in perioperative chemoimmunotherapy trials
The NADIM trial (ClinicalTrials.gov: NCT03081689) was a phase II, single-arm study that established perioperative chemoimmunotherapy as a highly promising approach in resectable stage IIIA NSCLC[74]. Patients (N = 46) received three cycles of nivolumab plus chemotherapy followed by surgery and one year of adjuvant nivolumab. At 5-year follow-up, the trial continued to report remarkably favorable outcomes, with long-term survival significantly outperforming historical controls[42]. A prespecified exploratory objective was to assess ctDNA as a potential biomarker of treatment efficacy and prognosis. At baseline, ctDNA was detectable in 30 of 43 (69.8%) patients with available samples, and levels were significantly correlated with tumor size (P = 0.021). Quantitative assessment of ctDNA burden—measured as the summatory mutant allele fraction (sumMAF)—demonstrated strong prognostic value. Patients with low baseline ctDNA levels (< 1% sumMAF) experienced superior 5-year progression-free survival (PFS) and OS, at 83.8% (95%CI: 65.3-92.9) and 86.2% (95%CI: 67.1-94.6), respectively, compared to 48.6% (95%CI: 19.2-73.0) and 56.3% (95%CI: 24.4-79.1) in those with sumMAF ≥ 1% (HR 4.26; 95%CI: 1.02-17.75, P = 0.0089 and HR 4.38; 95%CI: 0.91-21.10, P = 0.016 for PFS and OS, respectively). Of note, four of the five patients with disease progression despite low baseline ctDNA burden harbored unfavorable genomic alterations, including one EGFR and three kelch-like ECH-associated protein 1 (KEAP1) mutations, highlighting the potential additive prognostic relevance of mutational profile[75]. Clearance of ctDNA following neoadjuvant therapy—defined as undetectable or < 0.1% sumMAF—was also significantly associated with improved long-term outcomes. Among the 40 patients with evaluable post-treatment samples, 27 (67.5%) achieved ctDNA negativity prior to surgery. Patients who cleared ctDNA showed better 5-year survival: PFS was 85.2% and OS reached 92.3%, vs. 60.6% and 59.2%, respectively, in patients with residual detectable ctDNA (HR 3.06, 95%CI: 0.73-12.90,
Additional evidence comes from the NADIM II trial, which enrolled patients with resectable stage IIIA and IIIB NSCLC, thereby extending the evaluation of perioperative chemoimmunotherapy to a more advanced disease stage[43]. In this randomized phase II study, patients received perioperative nivolumab plus platinum-based chemotherapy or chemotherapy alone. Baseline ctDNA was detectable in 57 of 86 patients and correlated with tumor burden. Patients with low baseline ctDNA levels (MAF < 5%) experienced significantly better outcomes than those with high ctDNA levels (HR 0.38, 95%CI: 0.15-0.95 for PFS; HR 0.22, 95%CI: 0.07-0.64 for OS). Post-treatment plasma samples collected before surgery were available in a subset of patients. Following neoadjuvant therapy, ctDNA was undetectable in 67% (18/27) of patients in the chemoimmunotherapy arm, compared to 44% (4/9) in the chemotherapy arm.
Further insights on the prognostic utility of ctDNA in the perioperative setting comes from the phase III AEGEAN trial, which evaluated neoadjuvant platinum-based chemotherapy with or without durvalumab followed by surgery and one year of adjuvant durvalumab or placebo in patients with resectable stage II-IIIB NSCLC[3]. A comprehensive exploratory analysis focused on ctDNA dynamics at multiple timepoints: prior to each neoadjuvant therapy cycle, at surgery, and during adjuvant therapy (cycles 1, 3/4, and 10/11). A MRD landmark assessment was conducted at adjuvant cycle 1 day 1, corresponding to a median of 6.9 weeks after resection. WES of pretreatment tumor biopsies was used to identify somatic variants for the development of patient-specific, tumor-informed assays applied to plasma samples. At baseline, ctDNA was detectable in the majority of patients (89.6%, N = 283, 142 patients in the durvalumab arm and 141 in placebo arm), and its clearance during neoadjuvant treatment was observed more frequently in patients achieving a pathological response[76]. Among patients who were ctDNA-positive at baseline, all those who achieved pCR and over 93% of those with major pathologic response (MPR) had achieved ctDNA clearance by pre-surgical assessment. Conversely, lack of early clearance—particularly by cycle 2 day 1—had a high negative predictive value
In the phase III CheckMate 77T trial, patients with resectable stage IIA-IIIB (N2) NSCLC were randomized to receive neoadjuvant nivolumab plus platinum-based chemotherapy followed by surgery and adjuvant nivolumab or placebo[4]. Among biomarker-evaluable patients (N = 190 across both arms), baseline ctDNA was detectable in 76 (33%) patients in the nivolumab arm and 64 (28%) patients in the placebo arm[72]. Pre-surgical ctDNA clearance occurred more frequently with nivolumab (66% vs. 38%) and was associated with longer EFS across treatment groups, with the greatest benefit observed in the nivolumab arm (HR 0.41, 95%CI: 0.20-0.86). Importantly, the association between ctDNA clearance and improved EFS was evident regardless of pCR status. The EFS benefit with nivolumab was consistent across key genomic subgroups, including patients harboring KRAS, serine/threonine kinase 11 (STK11), or KEAP1 mutations[72].
Role of ctDNA in adjuvant immunotherapy trials
In the phase III IMpower010 trial, patients with completely resected stage IB-IIIA NSCLC received 1-4 cycles of adjuvant platinum-based chemotherapy before randomization to atezolizumab or best supportive care[5]. A prespecified exploratory analysis evaluated ctDNA at two key timepoints: post-surgery (before adjuvant chemotherapy) and post-chemotherapy (cycle 1 of adjuvant treatment in post-surgery ctDNA-positive patients), using a tumor-informed Signatera (Natera) assay[38]. Among 600 ctDNA-evaluable patients with stage II-IIIA disease, 112 of 534 (21%) were ctDNA-positive after surgery. Detection of ctDNA after surgery or completion of adjuvant chemotherapy was associated with inferior DFS. The addition of adjuvant atezolizumab after chemotherapy improved DFS and OS in PD-L1-positive population, irrespective of ctDNA status, and also showed benefit in longitudinal ctDNA monitoring. Among patients who were ctDNA-negative after adjuvant chemotherapy, the median time to conversion to ctDNA positivity was longer with atezolizumab (HR 0.60, 95%CI: 0.31-1.17), and DFS was also prolonged in this subgroup (31.3 vs. 13.3 months; HR 0.70, 95%CI: 0.37-1.34)[73].
In the phase III BR.31 trial, adjuvant durvalumab did not improve survival outcomes in resected EGFR/ALK-negative NSCLC[77]. A retrospective tumor-informed MRD analysis, performed in 1,131 patients (80% of the trial population), showed that postoperative MRD positivity was identified in approximately 10% of cases (116/1131) and was strongly associated with inferior OS in the placebo arm. A survival benefit with durvalumab was observed in MRD-positive patients within both PD-L1 ≥ 25% (P = 0.011) and ≥ 1%
Role of ctDNA in targeted therapy approaches for early-stage NSCLC
Based on the results of the phase III ADAURA trial, three years of adjuvant osimertinib are now recommended for patients with completely resected stage IB-IIIA EGFR-mutant NSCLC[6]. A retrospective, post hoc, tumor-informed MRD analysis was performed in a subset of 220 patients[37]. At randomization, MRD was detectable in 8% of patients (5/112, 4% with osimertinib; 13/108, 12% with placebo), all with stage II-IIIA disease. Baseline MRD positivity appeared to be associated with poor prognosis, with 17 of 18 patients experiencing a DFS event, although the subgroup size was small. Four of five MRD-positive patients receiving osimertinib achieved ctDNA clearance, compared with none in the placebo arm. During follow-up, MRD events occurred in 31% (68/220) of patients. Among those MRD-negative at randomization, MRD-free status was maintained in 91% of patients in the osimertinib arm versus 58% in the placebo arm. In the osimertinib group, most MRD or DFS events (68%) occurred after treatment discontinuation, often within 12 months. Across both arms, MRD positivity preceded clinical recurrence by a median of 4.7 months. At 36 months, the DFS and MRD event-free rate was 86% vs. 36% for osimertinib and placebo, respectively (HR 0.23, 95%CI: 0.15-0.36).
Recently, preliminary findings from the ADMIN study evaluating ctDNA dynamics in patients with resected NSCLC harboring uncommon EGFR mutations treated with adjuvant osimertinib were presented[78]. Treatment was given with or without prior adjuvant chemotherapy. ctDNA dynamics were assessed using a tumor-informed assay. Among 51 enrolled patients, 43 were evaluable for MRD analysis. ctDNA was detected in 23.3% (10/43), and half of these patients (5/10) achieved clearance after 12 weeks of osimertinib. At baseline, 18.6% (8/43) were plasma EGFR mutation-positive, with clearance observed in 62.5% (5/8).
In the phase III NeoADAURA trial, neoadjuvant osimertinib-containing regimens significantly improved MPR over chemotherapy alone in resectable, EGFR-mutated stage II-IIIB NSCLC[79]. An exploratory analysis from NeoADAURA evaluated the associations between baseline and pre-surgical MRD status with EFS and MPR. MRD testing was performed using the Personalis NeXT Personal® assay, an ultra-sensitive tumor-informed approach, in 189 patients. Compared with single EGFR mutation testing by Cobas® (30% detection rate), tumor-informed MRD analysis improved baseline ctDNA detection (71%). Patients without detectable MRD generally had less extensive disease at baseline, in terms of stage (II vs. III), tumor size, and nodal involvement. Notably, MRD negativity was associated with longer EFS (HR 0.24, 95%CI: 0.07-0.80). Treatment with osimertinib-containing regimens resulted in higher rates of MRD clearance compared with the control arm. Moreover, both pre-surgical MRD clearance and undetectable MRD status were associated with MPR (P = 0.0378 and P = 0.0546, respectively)[80]. The prognostic relevance of ctDNA was further supported by a phase II multi-institutional trial (NCT03433469) investigating neoadjuvant osimertinib in resectable stage I-IIIA EGFR-mutated NSCLC[81]. Among 23 evaluable patients, ctDNA testing was performed with the Guardant Reveal assay. Baseline ctDNA detection prior to treatment (C1D1) was independently associated with inferior EFS, with a median of 8.5 months in ctDNA-positive patients compared with 45.6 months in ctDNA-negative patients (HR 3.78, 95%CI: 1.09-13.13; P = 0.036). Similarly, ctDNA detection after one cycle of osimertinib (C1D28) was associated with shorter EFS in univariate analysis (18.7 vs. 45.6 months; HR 5.90, 95%CI: 1.09-32.01; P = 0.038), although this did not remain significant in multivariable analysis[82].
In a single-center pilot study (NORA trial) of 25 patients with stage IA-IIIA EGFR-mutant NSCLC, longitudinal tumor-informed ctDNA monitoring was integrated into a treatment strategy of neoadjuvant osimertinib (two 28-day cycles), surgery, and three years of adjuvant osimertinib[83]. Personalized ctDNA assays were developed for 20 patients, with baseline ctDNA positivity in six (30%). All achieved clearance after the first neoadjuvant cycle, irrespective of stage. Postoperatively, 95% (18/19) of recurrence-free patients remained ctDNA-negative, showing strong concordance between MRD status and disease control. One patient became ctDNA-positive while on adjuvant osimertinib without radiological recurrence, and another relapsed 2.6 months after treatment discontinuation despite ctDNA negativity. In terms of efficacy, neoadjuvant osimertinib achieved an objective response rate of 44% and a major pathological response rate of 24%, with no complete pathological responses reported.
POTENTIAL ROLE OF CTDNA-GUIDED RISK-ADAPTED ADJUVANT STRATEGIES
Adjuvant platinum-based chemotherapy remains the established postoperative standard for resected stage II-IIIA NSCLC, providing a modest but consistent survival benefit in unselected patients[84]. The advent of adjuvant targeted and immune therapies has further expanded this landscape, offering biomarker-driven strategies for molecularly defined subgroups[6,7,85]. In addition, perioperative approaches combining neoadjuvant chemoimmunotherapy followed by one year of adjuvant anti-PD-(L)1 therapy have reshaped the treatment paradigm for resectable NSCLC, achieving unprecedented rates of pCR[2-4]. In this evolving scenario, the ability to tailor adjuvant therapy intensity represents an unmet clinical need. While current regimens are administered to all eligible patients, a substantial proportion may already be cured by surgery alone and are therefore exposed to unnecessary toxicity. Conversely, a subset with MRD may require treatment intensification. ctDNA-based MRD monitoring offers a promising tool to personalize perioperative management, identifying patients who could benefit from therapy escalation or, conversely, those suitable for de-escalation or omission of adjuvant therapy. Multiple ongoing randomized studies are prospectively evaluating ctDNA-guided treatment strategies, aiming to determine whether MRD can serve as a reliable predictive biomarker to individualize therapeutic decisions in resected NSCLC. The phase III MERMAID-1 (NCT04385368) and MERMAID-2 (NCT04642469) trials represented the first randomized efforts to test ctDNA-guided adjuvant strategies in resected NSCLC. Both used tumor-informed MRD assays to guide risk-adapted adjuvant treatment but were discontinued early, as the approval of ICIs for early-stage disease made the control arms outdated within the evolving standard of care. Several MRD-guided phase II-III studies are currently ongoing to prospectively evaluate ctDNA as a tool to guide perioperative treatment decisions in resected NSCLC. These trials, summarized in Table 3, explore adjuvant immunotherapy and targeted therapies for oncogene-driven tumors, all sharing the goal of refining postoperative management through ctDNA-informed risk stratification. Collectively, they are expected to clarify whether MRD can serve as a predictive biomarker to individualize therapy intensity in early-stage disease.
Key ongoing clinical trials of MRD-guided adjuvant management in early-stage NSCLC
| Study | Phase | Population | MRD-guided adjuvant treatment | MRD test timing |
| ECTOP-1022 (NCT06323148) | Randomized phase 3 | EGFR-mutant stage II-IIIA (N1-N2) NSCLC | MRD+: CT MRD-: treatment based on clinico-pathologic assessment 1:1 randomization to osimertinib vs observation | Not reported |
| UMBRELLA (NCT06332274) | Phase 3 | Stage II-III NSCLC and other solid tumors | MRD+ after completion of surgery and perioperative treatments: tislelizumab | Not reported |
| Seagull (NCT05286957) | Randomized phase 2 | Stage IIA-IIIB NSCLC | MRD+: CT + tislelizumab MRD-: CT | Not reported |
| ADAPT-Lung/ CTONG2508 (NCT07120698) | Phase 2 | Stage II-IIIB NSCLC with no pCR after neoadjuvant CT-IO | MRD+: sintilimab MRD-: observation | 3-7 days, 28 days, then q3 months post-surgery |
| ADAPT-E (NCT04585477) | Phase 2 | Stage I-III NSCLC, MRD+ post-definitive therapy (surgery or RT) and adjuvant CT | Durvalumab | Not reported |
| CONTINUE (NCT06426511) | Phase 2 | Stage IB-IIIA NSCLC after adjuvant therapy | MRD+: toripalimab MRD-: observation | Not reported |
| NCT06780839 | Phase 2 | Stage IA3-IIIB (T3N2) ALK-positive NSCLC | MRD+: ensartinib; if MRD+ persists: ensartinib + CT MRD-: ensartinib; if MRD turns +: ensartinib + CT | 14-50 days post-surgery; q12 weeks for 2 years, then q24 weeks in year 3 |
FUTURE PERSPECTIVES AND CHALLENGES
Despite substantial progress in assay development, several limitations continue to hinder the clinical implementation of ctDNA in early-stage NSCLC. Addressing these limitations will be critical to enable the transition of ctDNA from a promising biomarker to a tool that can be effectively used in clinical practice. A key challenge is sensitivity, as ctDNA input is inherently low in early-stage tumors and in low-shedding histologies, which constrains reliable detection even with advanced NGS technologies. Several strategies have been proposed to overcome current sensitivity barriers and further lower the limit of detection, including longitudinal sampling to track more plasma samples, advanced error-suppression methods, enrichment of minor alleles, and increasing the amount of cfDNA[31,86]. In addition, there is marked heterogeneity in the assays used across studies, with a lack of standardization and no platforms currently approved by regulatory authorities for MRD assessment in this setting. Sequencing approaches also remain limited by high costs and longer turnaround times, restricting their applicability for real-time clinical decision-making.
Both tumor-informed and tumor-agnostic strategies present advantages and drawbacks. Tumor-informed assays, which track patient-specific mutations identified from tumor tissue, offer high sensitivity and specificity but require sufficient tissue and involve longer timelines for panel design. Tumor-agnostic approaches, in contrast, do not rely on prior tumor sequencing and can provide faster results, yet they are generally less sensitive and may be confounded by clonal hematopoiesis or non-tumor-derived signals. Beyond analytical variability, inconsistencies in preanalytical workflows, including blood collection, transport, plasma isolation, storage, and DNA extraction, constitute an additional source of heterogeneity[71]. The development and implementation of harmonized operating procedures will be crucial to ensure reproducibility and reliability of results.
To overcome current limitations, future developments are focusing on ultrasensitive detection methods and the integration of multi-omic approaches. Beyond genomic profiling, complementary strategies include methylation-based analyses (epigenomics), fragmentation profiling of cfDNA (fragmentomics), and the study of circulating cell-free RNA (cfRNA), which may provide insights into tumor biology and microenvironmental interactions.
DNA methylation patterns provide information on tissue-of-origin and can enhance detection by capturing tumor-derived fragments that may not harbor point mutations[87]. Fragmentomic signatures, including fragment length patterns, end motifs, and nucleosome positioning, also distinguish tumor-derived cfDNA from background leukocyte-derived fragments, reflecting differences in chromatin organization and nuclease activity[88,89]. By capturing these structural features, fragmentomic analyses represent a promising approach to improve detection sensitivity, particularly in early-stage NSCLC where ctDNA levels are often extremely low.
RNA-based liquid biopsy strategies have been explored in NSCLC, leveraging circulating tumor RNA (ctRNA) and exosomal RNA. Although still in early development, integration of RNA expression signatures may provide complementary information on tumor activity and immune microenvironment[90,91]. Proteomic approaches, particularly in combination with cfDNA, have also shown promise in early detection and may further refine risk stratification for MRD-positive patients[92].
Another unresolved issue is the optimal timing for ctDNA assessment in the postoperative setting. Available studies have used heterogeneous timepoints, and results suggest that sampling performed too early after surgery (e.g., within a few days) may be less reliable, likely due to postoperative inflammation and tissue healing[93]. Standardized timepoints for landmark and longitudinal monitoring are needed to improve interpretability and clinical application.
The clinical management of patients who convert from ctDNA-negative to ctDNA-positive status during follow-up also remains uncertain. No guidelines currently recommend ctDNA to guide follow-up in NSCLC. Standard imaging schedules, such as CT chest, remain the backbone of surveillance[8,9]. Incorporating MRD into these algorithms raises several questions: should MRD-positive patients undergo intensified imaging, PET/CT, or brain MRI? Should MRD-negative patients undergo de-escalated imaging to reduce radiation exposure and cost? At present, these scenarios are hypothetical. Without prospective data, altering follow-up based on MRD status risks both overutilization of imaging and missed opportunities for early intervention. Moreover, whether earlier detection of recurrence via ctDNA translates into improved survival remains unproven.
In this context, CTCs may offer complementary information to ctDNA in early-stage NSCLC. Higher perioperative CTC counts in pulmonary veins or peripheral blood have been consistently associated with an increased risk of recurrence and mortality[94]. In patients who are ctDNA-negative after surgery but later relapse, recurrence may reflect disseminated tumor cells that persist in dormant niches and reactivate at a later stage[50]. In breast cancer, persistent or newly detectable CTCs during post-adjuvant surveillance have shown prognostic significance across multiple large studies[49]. Although similar data in NSCLC are limited, these findings support the potential utility of CTCs monitoring for risk stratification during follow-up. Assessing CTCs in combination with ctDNA may improve diagnostic sensitivity compared with single-modality testing, supporting future evaluation of multimodal liquid biopsy approaches for early detection and postoperative surveillance. While accumulating evidence indicates that ctDNA-positive patients derive survival benefit from adjuvant chemotherapy, whereas ctDNA-negative patients do not, the available data are insufficient to justify treatment escalation or de-escalation outside clinical trials.
Randomized studies in which therapeutic decisions are guided by ctDNA status will be crucial to establish the predictive value of ctDNA and to determine its role in tailoring perioperative systemic therapies.
Overall, perioperative ctDNA has consistently shown prognostic value when detectable both before and after treatment. However, its predictive significance remains unproven. This distinction is particularly relevant in the current therapeutic landscape, where perioperative chemoimmunotherapy and targeted therapies are practice-changing strategies, and treatment selection is still exclusively based on EGFR, ALK, and PD-L1 status.
CONCLUSION
ctDNA has emerged as a promising biomarker in early-stage NSCLC, offering dynamic and non-invasive assessment of MRD and recurrence risk. Across retrospective and prospective studies, ctDNA detection before and after curative-intent therapy has consistently correlated with poorer outcomes, with postoperative positivity identifying patients at highest risk of relapse, often months prior to radiological progression. These findings highlight the strong prognostic value of ctDNA in the perioperative setting. However, several challenges still preclude routine clinical use. Limitations related to detection sensitivity, methodological variability, and the timing of sampling remain significant barriers. Importantly, although ctDNA positivity is consistently prognostic, its predictive role in guiding therapeutic decisions remains unproven.
Exploratory analyses from practice-changing trials—including CheckMate 816, NADIM, AEGEAN, CheckMate 77T, IMpower010, and ADAURA—demonstrate that ctDNA clearance during neoadjuvant or perioperative therapy is associated with improved long-term survival, complementing pathological response as a marker of treatment efficacy. Nonetheless, these data remain hypothesis-generating, as none of the trials were designed to tailor therapy based on ctDNA status.
Future directions include the standardization of assays and sampling schedules, as well as the integration of multi-omic approaches such as methylation profiling, fragmentomics, and RNA-based analyses to improve sensitivity. Prospective randomized studies, including ctDNA-guided clinical trials, will ultimately be essential to establish whether ctDNA can serve not only as a prognostic but also as a predictive biomarker. If validated, ctDNA-guided strategies could personalize perioperative therapy, enabling treatment escalation for high-risk patients while sparing low-risk patients from unnecessary toxicity.
DECLARATIONS
Acknowledgments
We thank Antonella Cammarota for assistance with the creation of Figure 1, Figure 2, and the graphical abstract using BioRender.com.
Authors’ contributions
Conceptualization, data curation: Masini S, Finocchiaro G
Methodology, writing - review and editing: Masini S, Finocchiaro G, Toschi L
Resources: Masini S, Fila M, Rudini M, Finocchiaro G
Writing - original draft preparation: Masini S, Fila M, Finocchiaro G
Supervision: Santoro A, Toschi L, Finocchiaro G
All authors have read and agreed to the published version of the manuscript.
Availability of data and materials
Not applicable.
AI and AI-assisted tools statement
During the preparation of this manuscript, the AI tool ChatGPT (OpenAI, based on the GPT-5.3 model) was used solely for language editing. The tool did not influence the study design, data collection, analysis, interpretation, or the scientific content of the work. All authors take full responsibility for the accuracy, integrity, and final content of the manuscript.
Financial support and sponsorship
None.
Conflicts of interest
All authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Forde PM, Spicer J, Lu S, et al. Neoadjuvant nivolumab plus chemotherapy in resectable lung cancer. N Engl J Med. 2022;386:1973-85.
2. Wakelee H, Liberman M, Kato T, et al. Perioperative pembrolizumab for early-stage non-small-cell lung cancer. N Engl J Med. 2023;389:491-503.
3. Heymach JV, Harpole D, Mitsudomi T, et al. Perioperative durvalumab for resectable non-small-cell lung cancer. N Engl J Med. 2023;389:1672-84.
4. Cascone T, Awad MM, Spicer JD, et al. Perioperative nivolumab in resectable lung cancer. N Engl J Med. 2024;390:1756-69.
5. Felip E, Altorki N, Zhou C, et al. Adjuvant atezolizumab after adjuvant chemotherapy in resected stage IB-IIIA non-small-cell lung cancer (IMpower010): a randomised, multicentre, open-label, phase 3 trial. Lancet. 2021;398:1344-57.
6. Wu YL, Tsuboi M, He J, et al. Osimertinib in resected EGFR-mutated non-small-cell lung cancer. N Engl J Med. 2020;383:1711-23.
7. Wu YL, Dziadziuszko R, Ahn JS, et al. Alectinib in resected ALK-positive non-small-cell lung cancer. N Engl J Med. 2024;390:1265-76.
8. Zer A, Ahn MJ, Barlesi F, et al. Early and locally advanced non-small-cell lung cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann Oncol. 2025;36:1245-62.
9. National Comprehensive Cancer Network. NCCN clinical practice guidelines in oncology (NCCN Guidelines®). Non-Small Cell Lung Cancer. Version 5 2026. Available from: https://www.nccn.org/professionals/physician_gls/pdf/nscl.pdf [Last accessed on 23 Apr 2026].
10. Goldstraw P, Chansky K, Crowley J, et al. The IASLC lung cancer staging project: proposals for revision of the TNM stage groupings in the forthcoming (Eighth) edition of the TNM classification for lung cancer. J Thorac Oncol. 2016;11:39-51.
11. Abbosh C, Frankell AM, Harrison T, et al. Tracking early lung cancer metastatic dissemination in TRACERx using ctDNA. Nature. 2023;616:553-62.
12. Normanno N, Morabito A, Rachiglio AM, et al. Circulating tumour DNA in early stage and locally advanced NSCLC: ready for clinical implementation? Nat Rev Clin Oncol. 2025;22:215-31.
13. Corcoran RB, Chabner BA. Application of cell-free DNA analysis to cancer treatment. N Engl J Med. 2018;379:1754-65.
14. Hendriks LE, Kerr KM, Menis J, et al. Oncogene-addicted metastatic non-small-cell lung cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann Oncol. 2023;34:339-57.
15. Rolfo C, Mack P, Scagliotti GV, et al. Liquid biopsy for advanced NSCLC: a consensus statement from the international association for the study of lung cancer. J Thorac Oncol. 2021;16:1647-62.
16. Marinello A, Tagliamento M, Pagliaro A, et al. Circulating tumor DNA to guide diagnosis and treatment of localized and locally advanced non-small cell lung cancer. Cancer Treat Rev. 2024;129:102791.
17. Stejskal P, Goodarzi H, Srovnal J, Hajdúch M, van 't Veer LJ, Magbanua MJM. Circulating tumor nucleic acids: biology, release mechanisms, and clinical relevance. Mol Cancer. 2023;22:15.
18. Diaz LA Jr, Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014;32:579-86.
19. Newman AM, Bratman SV, To J, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20:548-54.
20. Bartolomucci A, Nobrega M, Ferrier T, et al. Circulating tumor DNA to monitor treatment response in solid tumors and advance precision oncology. NPJ Precis Oncol. 2025;9:84.
21. Herbreteau G, Vallée A, Knol AC, et al. Quantitative monitoring of circulating tumor DNA predicts response of cutaneous metastatic melanoma to anti-PD1 immunotherapy. Oncotarget. 2018;9:25265-76.
22. Takayama Y, Suzuki K, Muto Y, et al. Monitoring circulating tumor DNA revealed dynamic changes in KRAS status in patients with metastatic colorectal cancer. Oncotarget. 2018;9:24398-413.
23. Diehl F, Schmidt K, Choti MA, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985-90.
24. García-Foncillas J, Alba E, Aranda E, et al. Incorporating BEAMing technology as a liquid biopsy into clinical practice for the management of colorectal cancer patients: an expert taskforce review. Ann Oncol. 2017;28:2943-9.
25. Corless BC, Chang GA, Cooper S, et al. Development of novel mutation-specific droplet digital PCR assays detecting TERT promoter mutations in tumor and plasma samples. J Mol Diagn. 2019;21:274-85.
26. Dong L, Wang S, Fu B, Wang J. Evaluation of droplet digital PCR and next generation sequencing for characterizing DNA reference material for KRAS mutation detection. Sci Rep. 2018;8:9650.
27. Vessies DCL, Greuter MJE, van Rooijen KL, et al. Performance of four platforms for KRAS mutation detection in plasma cell-free DNA: ddPCR, Idylla, COBAS z480 and BEAMing. Sci Rep. 2020;10:8122.
28. Forshew T, Murtaza M, Parkinson C, et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med. 2012;4:136ra68.
29. Kinde I, Wu J, Papadopoulos N, Kinzler KW, Vogelstein B. Detection and quantification of rare mutations with massively parallel sequencing. Proc Natl Acad Sci USA. 2011;108:9530-5.
30. Newman AM, Lovejoy AF, Klass DM, et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat Biotechnol. 2016;34:547-55.
31. Kurtz DM, Soo J, Co Ting Keh L, et al. Enhanced detection of minimal residual disease by targeted sequencing of phased variants in circulating tumor DNA. Nat Biotechnol. 2021;39:1537-47.
32. Zviran A, Schulman RC, Shah M, et al. Genome-wide cell-free DNA mutational integration enables ultra-sensitive cancer monitoring. Nat Med. 2020;26:1114-24.
33. Martín-Arana J, Gimeno-Valiente F, Henriksen TV, et al. Whole-exome tumor-agnostic ctDNA analysis enhances minimal residual disease detection and reveals relapse mechanisms in localized colon cancer. Nat Cancer. 2025;6:1000-16.
34. Abbosh C, Birkbak NJ, Wilson GA, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature. 2017;545:446-51.
35. Forde PM, Spicer JD, Provencio M, et al. Overall survival with neoadjuvant nivolumab plus chemotherapy in lung cancer. N Engl J Med. 2025;393:741-52.
36. Reck M, Gale D, Zhu Z, et al. Association of post-surgical MRD status with neoadjuvant ctDNA dynamics, genomic mutations, and clinical outcomes in patients with resectable NSCLC (R-NSCLC) from the phase 3 AEGEAN trial. J Clin Oncol. 2025;43:8009.
37. Herbst RS, John T, Grohé C, et al. Molecular residual disease analysis of adjuvant osimertinib in resected EGFR-mutated stage IB-IIIA non-small-cell lung cancer. Nat Med. 2025;31:1958-68.
38. Wakelee H, Reck M, Felip E, et al. 1211P IMpower010: ctDNA status and 5y DFS follow up in patients (pts) with resected NSCLC who received adjuvant chemotherapy (chemo) followed by atezolizumab (atezo) or best supportive care (BSC). Ann Oncol. 2024;35:S779-80.
39. Black JRM, Bartha G, Abbott CW, et al. Ultrasensitive ctDNA detection for preoperative disease stratification in early-stage lung adenocarcinoma. Nat Med. 2025;31:70-6.
40. Coombes RC, Page K, Salari R, et al. Personalized detection of circulating tumor DNA antedates breast cancer metastatic recurrence. Clin Cancer Res. 2019;25:4255-63.
41. Heider K, Gale D, Ruiz-valdepenas A, et al. Abstract 735: sensitive detection of ctDNA in early stage non-small cell lung cancer patients with a personalized sequencing assay. Cancer Res. 2020;80:735.
42. Provencio M, Serna-Blasco R, Nadal E, et al. Overall survival and biomarker analysis of neoadjuvant nivolumab plus chemotherapy in operable stage IIIA non-small-cell lung cancer (NADIM phase II trial). J Clin Oncol. 2022;40:2924-33.
43. Provencio M, Nadal E, González-Larriba JL, et al. Perioperative nivolumab and chemotherapy in stage III non-small-cell lung cancer. N Engl J Med. 2023;389:504-13.
44. Andrikou K, Rossi T, Verlicchi A, et al. Circulating tumour cells: detection and application in advanced non-small cell lung cancer. Int J Mol Sci. 2023;24:16085.
45. Ward MP, E Kane L, A Norris L, et al. Platelets, immune cells and the coagulation cascade; friend or foe of the circulating tumour cell? Mol Cancer. 2021;20:59.
46. Moose DL, Krog BL, Kim TH, et al. Cancer cells resist mechanical destruction in circulation via RhoA/actomyosin-dependent mechano-adaptation. Cell Rep. 2020;30:3864-74.e6.
48. Tsering T, Nadeau A, Wu T, Dickinson K, Burnier JV. Extracellular vesicle-associated DNA: ten years since its discovery in human blood. Cell Death Dis. 2024;15:668.
49. Alix-Panabières C, Pantel K. From discovery to diagnosis: a perspective for circulating tumor cells in personalized oncology. Cancer Discov. 2025;15:1985-2001.
50. Nasr MM, Lynch CC. How circulating tumor cluster biology contributes to the metastatic cascade: from invasion to dissemination and dormancy. Cancer Metastasis Rev. 2023;42:1133-46.
51. Chaudhuri AA, Chabon JJ, Lovejoy AF, et al. Early detection of molecular residual disease in localized lung cancer by circulating tumor DNA profiling. Cancer Discov. 2017;7:1394-403.
52. Peng M, Huang Q, Yin W, et al. Circulating tumor DNA as a prognostic biomarker in localized non-small cell lung cancer. Front Oncol. 2020;10:561598.
53. Li N, Wang BX, Li J, et al. Perioperative circulating tumor DNA as a potential prognostic marker for operable stage I to IIIA non-small cell lung cancer. Cancer. 2022;128:708-18.
54. Xia L, Mei J, Kang R, et al. Perioperative ctDNA-based molecular residual disease detection for non-small cell lung cancer: a prospective multicenter cohort study (LUNGCA-1). Clin Cancer Res. 2022;28:3308-17.
55. Qiu B, Guo W, Zhang F, et al. Dynamic recurrence risk and adjuvant chemotherapy benefit prediction by ctDNA in resected NSCLC. Nat Commun. 2021;12:6770.
56. Kuang PP, Li N, Liu Z, et al. Circulating Tumor DNA Analyses as a potential marker of recurrence and effectiveness of adjuvant chemotherapy for resected non-small-cell lung cancer. Front Oncol. 2020;10:595650.
57. Waldeck S, Mitschke J, Wiesemann S, et al. Early assessment of circulating tumor DNA after curative-intent resection predicts tumor recurrence in early-stage and locally advanced non-small-cell lung cancer. Mol Oncol. 2022;16:527-37.
58. Gale D, Heider K, Ruiz-Valdepenas A, et al. Residual ctDNA after treatment predicts early relapse in patients with early-stage non-small cell lung cancer. Ann Oncol. 2022;33:500-10.
59. Wang S, Li M, Zhang J, et al. Circulating tumor DNA integrating tissue clonality detects minimal residual disease in resectable non-small-cell lung cancer. J Hematol Oncol. 2022;15:137.
60. Chen K, Yang F, Shen H, et al. Individualized tumor-informed circulating tumor DNA analysis for postoperative monitoring of non-small cell lung cancer. Cancer Cell. 2023;41:1749-62.e6.
61. Jung HA, Ku BM, Kim YJ, et al. Longitudinal monitoring of circulating tumor DNA from plasma in patients with curative resected stages I to IIIA EGFR-mutant non-small cell lung cancer. J Thorac Oncol. 2023;18:1199-208.
62. Tran HT, Heeke S, Sujit S, et al. Circulating tumor DNA and radiological tumor volume identify patients at risk for relapse with resected, early-stage non-small-cell lung cancer. Ann Oncol. 2024;35:183-9.
63. Tan AC, Lai GGY, Saw SPL, et al. Detection of circulating tumor DNA with ultradeep sequencing of plasma cell-free DNA for monitoring minimal residual disease and early detection of recurrence in early-stage lung cancer. Cancer. 2024;130:1758-65.
64. Ohara S, Suda K, Sudhaman S, et al. Clinical significance of perioperative minimal residual disease detected by circulating tumor DNA in patients with lung cancer with a long follow-up data: an exploratory study. JTO Clin Res Rep. 2025;6:100762.
65. Martin TK, Dinerman A, Sudhaman S, et al. Early real-world experience monitoring circulating tumor DNA in resected early-stage non-small cell lung cancer. J Thorac Cardiovasc Surg. 2024;168:1349-59.e2.
66. Oh Y, Yoon SM, Lee J, et al. Personalized, tumor-informed, circulating tumor DNA assay for detecting minimal residual disease in non-small cell lung cancer patients receiving curative treatments. Thorac Cancer. 2024;15:1095-102.
67. Schuurbiers MMF, Smith CG, Hartemink KJ, et al. Recurrence prediction using circulating tumor DNA in patients with early-stage non-small cell lung cancer after treatment with curative intent: a retrospective validation study. PLoS Med. 2025;22:e1004574.
68. Zhang JT, Liu SY, Gao X, et al. Follow-up analysis enhances understanding of molecular residual disease in localized non-small cell lung cancer. Clin Cancer Res. 2025;31:1305-14.
69. Giobbie-Hurder A, Gelber RD, Regan MM. Challenges of guarantee-time bias. J Clin Oncol. 2013;31:2963-9.
70. Zhang JT, Liu SY, Gao W, et al. Longitudinal undetectable molecular residual disease defines potentially cured population in localized non-small cell lung cancer. Cancer Discov. 2022;12:1690-701.
71. der Leest P, Schuuring E. Critical factors in the analytical work flow of circulating tumor DNA-based molecular profiling. Clin Chem. 2024;70:220-33.
72. Cascone T, Awad MM, Spicer J, et al. Perioperative nivolumab (NIVO) vs placebo (PBO) in patients (pts) with resectable NSCLC: updated survival and biomarker analyses from CheckMate 77T. J Clin Oncol. 2025;43:LBA8010.
73. Felip E, Srivastava M, Reck M, et al. 1O IMpower010: ctDNA status in patients (pts) with resected NSCLC who received adjuvant chemotherapy (chemo) followed by atezolizumab (atezo) or best supportive care (BSC). Immuno Oncol Technol. 2022;16:100106.
74. Provencio M, Nadal E, Insa A, et al. Neoadjuvant chemotherapy and nivolumab in resectable non-small-cell lung cancer (NADIM): an open-label, multicentre, single-arm, phase 2 trial. Lancet Oncol. 2020;21:1413-22.
75. Provencio M, Nadal E, Insa A, et al. Perioperative chemotherapy and nivolumab in non-small-cell lung cancer (NADIM): 5-year clinical outcomes from a multicentre, single-arm, phase 2 trial. Lancet Oncol. 2024;25:1453-64.
76. Reck M, Gale D, Zhu Z, et al. LBA49 associations of ctDNA clearance (CL) during neoadjuvant Tx with pathological response and event-free survival (EFS) in pts with resectable NSCLC (R-NSCLC): expanded analyses from AEGEAN. Ann Oncol. 2024;35:S1239.
77. Goss G, Darling G, Westeel V, et al. LBA68 CCTG BR.31: Adjuvant durvalumab (D) in resected non-small-cell lung cancer (NSCLC): Final overall survival (OS) and minimal residual disease (MRD) analyses. Ann Oncol. 2025;36:S1609.
78. Liu C, Leng X, Yue D, et al. PT1.07.03 adjuvant osimertinib in IB-IIIB NSCLC with uncommon EGFR mutations (admin): mutation profile and ctDNA dynamics. J Thorac Oncol. 2025;20:S601.
79. He J, Tsuboi M, Weder W, et al. Neoadjuvant osimertinib for resectable EGFR-mutated non-small cell lung cancer. J Clin Oncol. 2025;43:2875-87.
80. Blakely C, Robichaux J, Lee S, et al. OA02.02 molecular residual disease (MRD) analysis from NeoADAURA: neoadjuvant osimertinib ± chemotherapy in resectable EGFRm NSCLC. J Thorac Oncol. 2025;20:S12-3.
81. Blakely CM, Urisman A, Gubens MA, et al. Neoadjuvant osimertinib for the treatment of stage I-IIIA epidermal growth factor receptor-mutated non-small cell lung cancer: a phase II multicenter study. J Clin Oncol. 2024;42:3105-14.
82. Brown T, Gubens M, Mulvey C, et al. P1.07.40 detection of ctDNA prior to neoadjuvant osimertinib treatment correlates with decreased EFS in EGFR-mutated NSCLC. J Thorac Oncol. 2025;20:S153-4.
83. Lee JB, Choi SJ, Shim HS, et al. Neoadjuvant and adjuvant osimertinib in stage IA to IIIA, EGFR-mutant NSCLC (NORA). J Thorac Oncol. 2025;20:641-50.
84. Pignon JP, Tribodet H, Scagliotti GV, et al. Lung adjuvant cisplatin evaluation: a pooled analysis by the LACE collaborative group. J Clin Oncol. 2008;26:3552-9.
85. Felip E, Altorki N, Zhou C, et al. Overall survival with adjuvant atezolizumab after chemotherapy in resected stage II-IIIA non-small-cell lung cancer (IMpower010): a randomised, multicentre, open-label, phase III trial. Ann Oncol. 2023;34:907-19.
86. Moding EJ, Nabet BY, Alizadeh AA, Diehn M. Detecting liquid remnants of solid tumors: circulating tumor DNA minimal residual disease. Cancer Discov. 2021;11:2968-86.
87. Liu MC, Oxnard GR, Klein EA, Swanton C, Seiden MV, CCGA Consortium. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann Oncol. 2020;31:745-59.
88. Cristiano S, Leal A, Phallen J, et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature. 2019;570:385-9.
89. Tsui WHA, Jiang P, Lo YMD. Cell-free DNA fragmentomics in cancer. Cancer Cell. 2025;43:1792-814.
90. Larson MH, Pan W, Kim HJ, et al. A comprehensive characterization of the cell-free transcriptome reveals tissue- and subtype-specific biomarkers for cancer detection. Nat Commun. 2021;12:2357.
91. Nesselbush MC, Luca BA, Jeon YJ, et al. An ultrasensitive method for detection of cell-free RNA. Nature. 2025;641:759-68.
92. Cohen JD, Li L, Wang Y, et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science. 2018;359:926-30.
93. Henriksen TV, Reinert T, Christensen E, et al. The effect of surgical trauma on circulating free DNA levels in cancer patients-implications for studies of circulating tumor DNA. Mol Oncol. 2020;14:1670-9.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Copyright
Data & Comments
Data
















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].