fig4
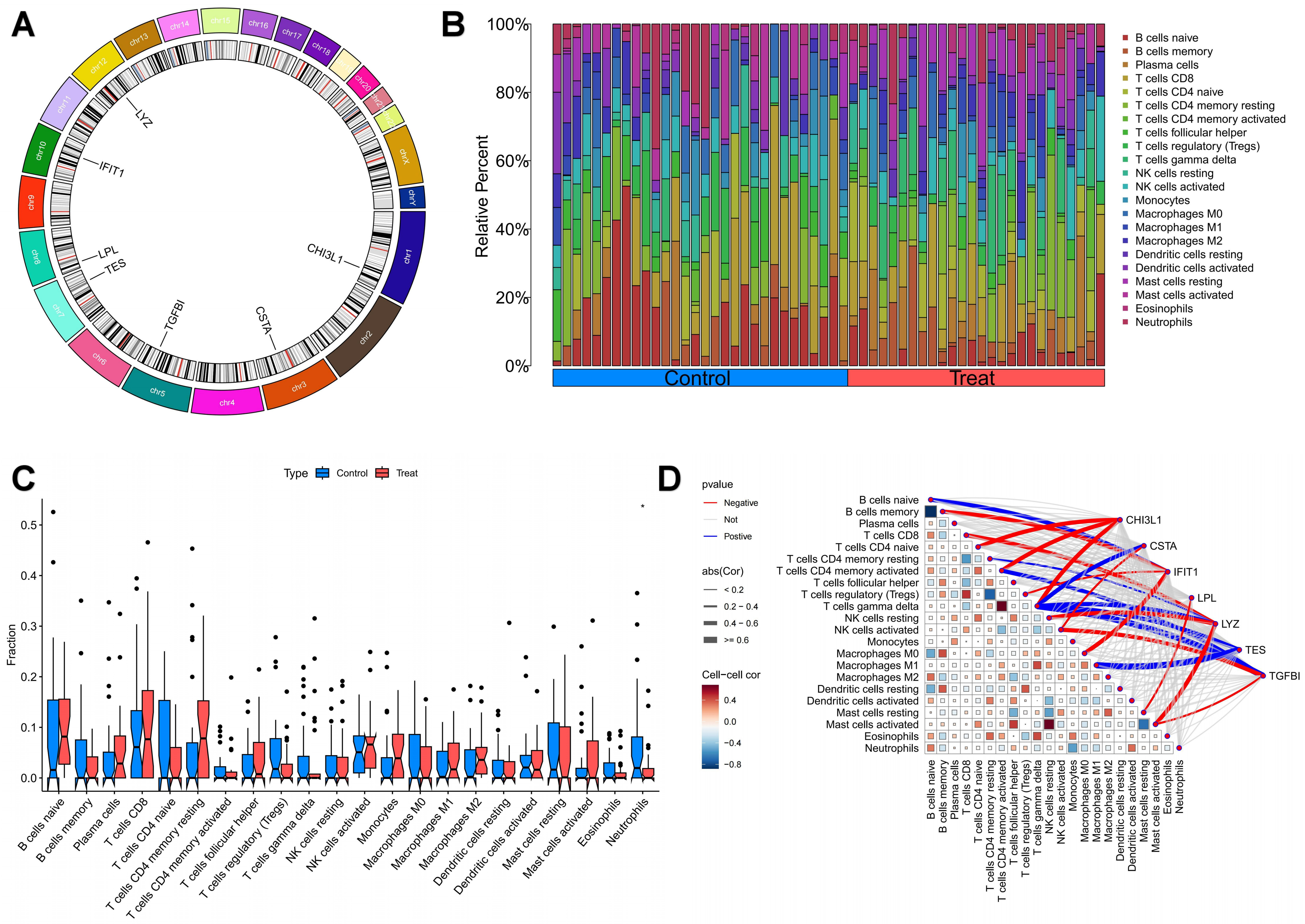
Figure 4. Visualization of immune infiltration assessment for key genes. (A) Circos plot of intersecting genes; (B) Stacked histogram of immune cell proportions between DKD groups and control groups; (C) Box plot comparing 22 immune cells between DKD groups and control groups; (D) Heatmap of correlations between 22 types of immune cells and key genes. Differences in immune cell infiltration between groups (C) were analyzed using the Wilcoxon rank-sum test. Correlations between key genes and immune cells in DKD samples (D) were assessed using Spearman’s rank correlation analysis. DKD: Diabetic kidney disease; LYZ: lysozyme; IFIT1: interferon induced protein with tetratricopeptide repeats 1; LPL: lipoprotein lipase; TES: testin LIM domain protein; TGFBI: transforming growth factor beta induced; CSTA: cystatin A; CHI3L1: chitinase 3-like 1; NK: natural killer.