fig2
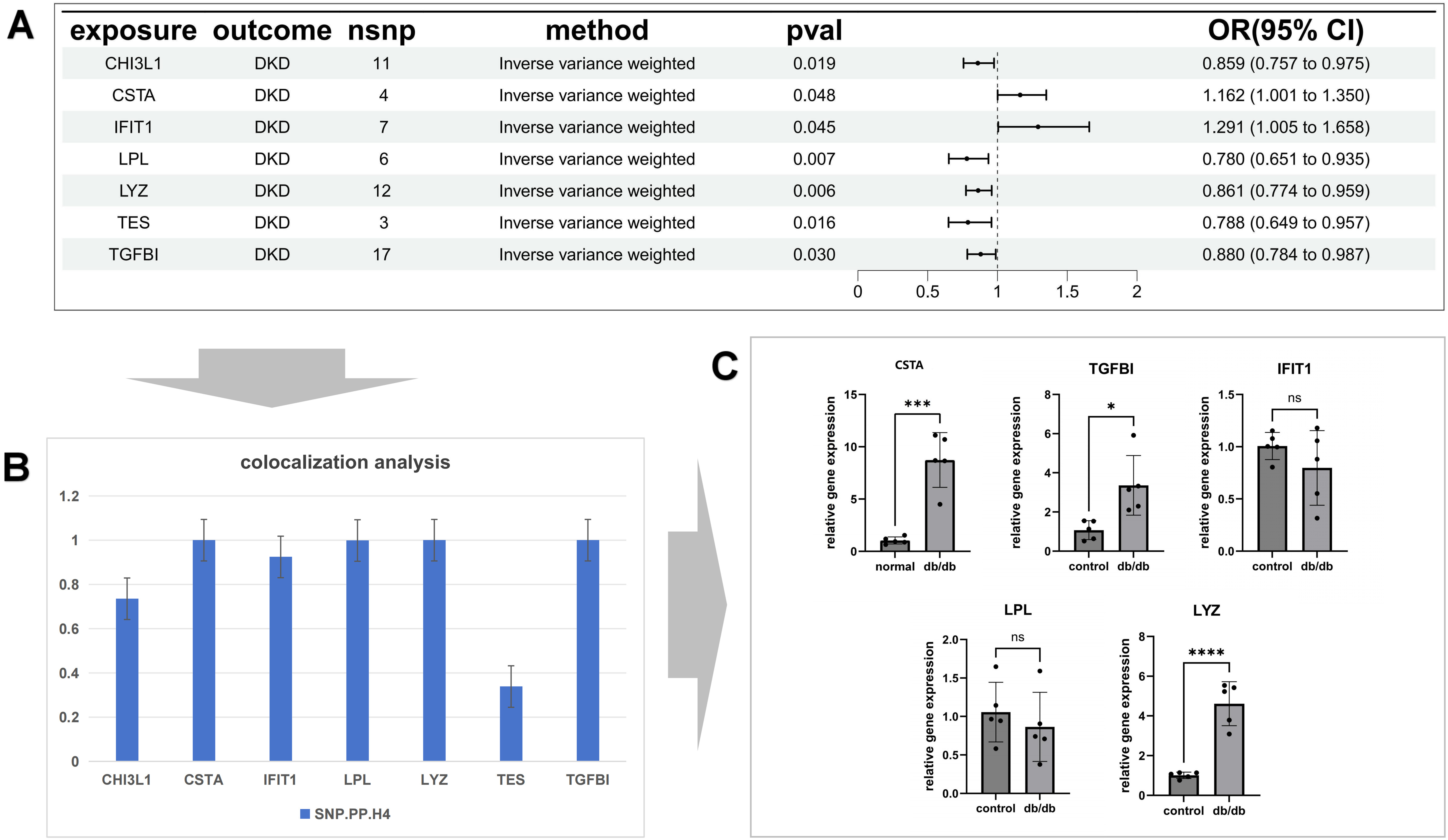
Figure 2. Identification of potential causal proteins for DKD via MR and experimental validation. (A) Forest plot of MR analysis showing the causal effects of seven plasma proteins on DKD risk using the IVW method. The OR and 95%CI are presented; (B) Colocalization analysis evaluating the probability of shared causal variants between protein expression and DKD. The y-axis represents the PP.H4; (C) Validation of relative gene expression levels for candidate targets (CSTA, TGFBI, IFIT1, LPL, and LYZ) in kidney tissues of db/db mice compared with controls (normal). Data are presented as mean ± SD. Statistical significance was determined using unpaired Student’s t-tests (*P < 0.05, ***P < 0.001, ****P < 0.0001; ns, not significant). DKD: Diabetic kidney disease; MR: Mendelian randomization; IVW: inverse-variance weighted; OR: odds ratio; CI: confidence interval; CSTA: cystatin A; TGFBI: transforming growth factor beta induced; IFIT1: interferon induced protein with tetratricopeptide repeats 1; LPL: lipoprotein lipase; LYZ: lysozyme; SD: standard deviation; CHI3L1: chitinase 3-like 1; TES: testin LIM domain protein; SNP: single nucleotide polymorphism.