Computer vision for efficient object detection and segmentation in molecular image analysis
Abstract
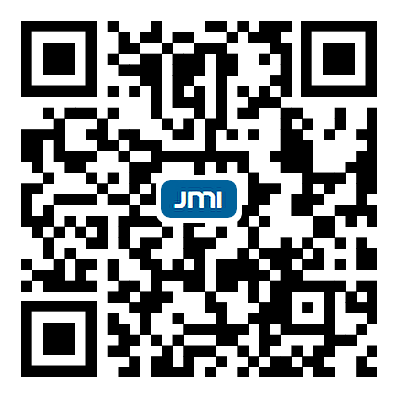
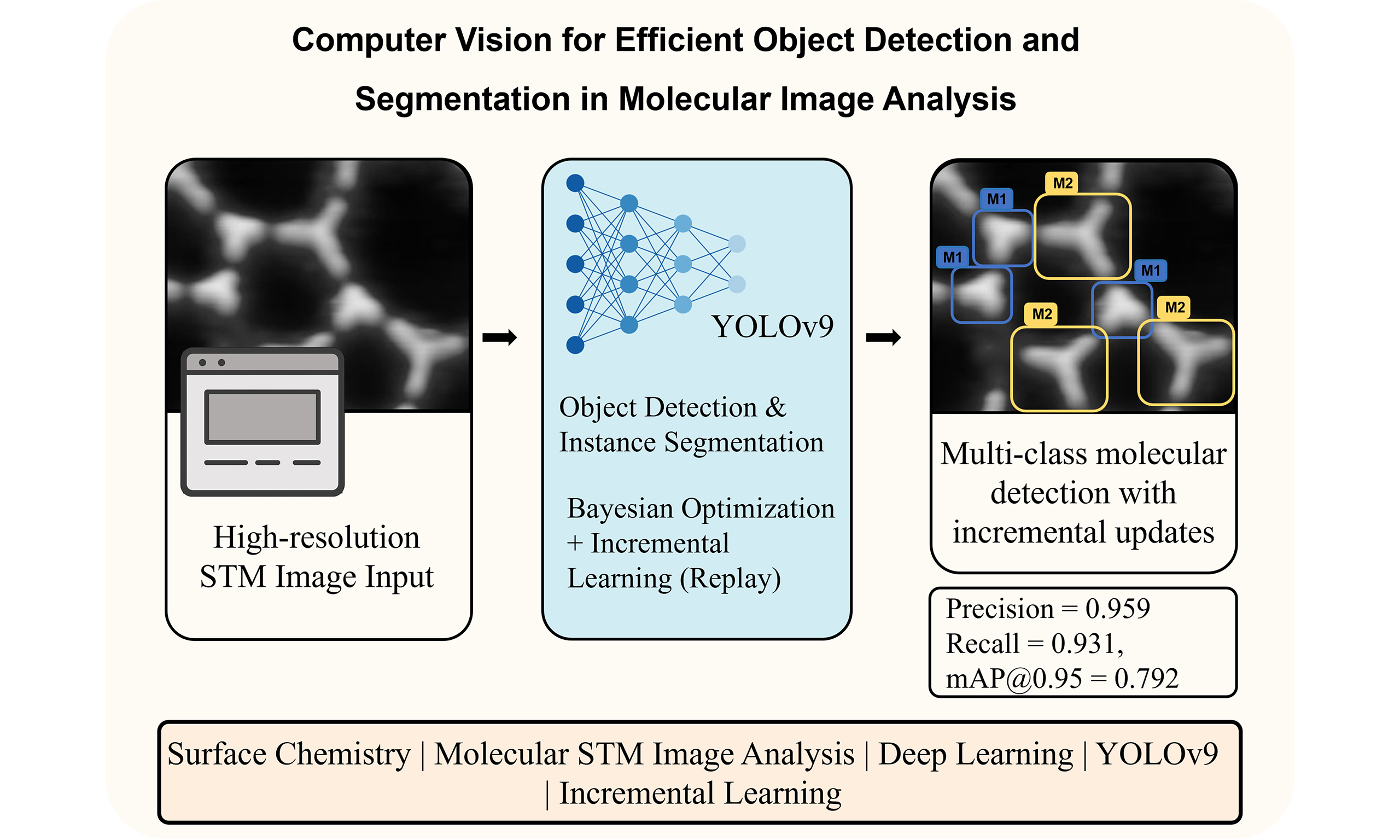
Image recognition, classification, and analysis of large sets of high-resolution molecular images are time-consuming and labor-intensive, even for human experts, due to the lack of standardized approaches. In recent years, machine learning has emerged as a powerful tool for automating image data analysis in materials science. In this work, we developed a computer vision program for efficient object detection and instance segmentation, offering a fast alternative to manual molecular image analysis. By integrating You Only Look Once version 9 (YOLOv9) with an incremental learning strategy and hyperparameter optimization, the system enables accurate detection, classification, and segmentation of molecular species across diverse scanning tunneling microscopy datasets. Our results demonstrate robust performance and minimal forgetting rates across multiple molecular categories, enabling scalable and updatable surface image analysis workflows. We anticipate that computer vision methods will see increasing applications in image data analysis within the field of on-surface chemistry.
Keywords
INTRODUCTION
In recent years, the rise of artificial intelligence (AI) technologies and advances in materials science have brought about profound transformations in traditional materials design, synthesis, and characterization[1-5]. Scanning tunneling microscopy (STM), a vital technique for investigating surface structures and adsorbates, has become widely employed in fields such as catalysis, molecular self-assembly, on-surface synthesis, and nanomaterials preparation[6-12], primarily due to its ability to provide atomic-level resolution[13-18]. However, the high-resolution images obtained from STM measurements typically exhibit considerable complexity and contain rich microscopic information. Consequently, the analysis and interpretation of such images depend heavily on the experience and subjective judgment of human experts. This reliance not only reduces research efficiency but also restricts the broader adoption and application of the cutting-edge technology.
To overcome the critical dependence on prior knowledge of human experts in high-resolution image analysis, AI techniques have increasingly been introduced[19-28]. Recent studies have demonstrated that deep learning and computer vision algorithms can effectively automate the recognition and analysis of STM images, including the automatic detection and correction of imaging artifacts[20], classification of similar molecules[22], molecular counting[19], and automated length measurement of polymer chains. These studies highlight the significant potential of integrating AI with STM technology to improve analytical accuracy and reduce manual intervention.
Nevertheless, current research remains largely limited to isolated applications tailored to specific scenarios, typically focusing only on achieving a single functionality or a particular objective. Specifically, most existing studies employ small-scale, customized datasets for training and testing, lacking systematic data preparation and a general framework for predictive modeling and result analysis. Therefore, the systematic integration of the entire AI pipeline, which includes data acquisition and preprocessing, model training, automated prediction, and multi-scale data analysis, is one of the key challenges for achieving efficient, automated, and intelligent STM-based research.
In this work, we develop a comprehensive framework that integrates multiple aspects of the machine learning workflow, including image data labeling, dataset creation, model training, inference, and data analysis. Based on the You Only Look Once version 9 (YOLOv9) model, a state-of-the-art approach that combines both object detection and instance segmentation capabilities, our program provides a one-stop solution for annotating image data, generating high-quality datasets, and training advanced models. Additionally, the program includes tools for data analysis based on prediction results, providing insights that can guide decision-making and further refine model performance. A key feature of the program is its support for incremental learning, allowing models to be continuously updated and improved as new data become available. This approach avoids the need for complete retraining from scratch and ensures that the models remain accurate and up to date.
MATERIALS AND METHODS
Data preparation and labeling
To develop a robust model for molecular image analysis, we first curated a dataset of molecular images containing various molecular assemblies. The images were pre-processed to standardize resolution and contrast, ensuring uniformity across the dataset. Using an annotation tool, molecular structures were manually labeled with bounding boxes and segmentation masks for training the object detection and instance segmentation models. Each molecular species was assigned a unique class identifier (ID), and the dataset was divided into training, validation, and test sets following an 80:10:10 split. All STM images used in this study were obtained from our group’s experiments on molecular self-assembly. To enhance dataset diversity, 2-3 original high-resolution STM images were augmented using operations such as rotation, flipping, cropping, and contrast adjustment, resulting in approximately 500 images in total. These augmentations [Supplementary Section 1] preserve the intrinsic physical and structural characteristics of the molecular features while improving the robustness of model training. This dataset was then used for object detection labeling, with molecular positions annotated using bounding boxes to form the complete training and validation sets.
Model selection and architecture
Our program is built on YOLOv9, the latest iteration of the YOLO family, which is known for efficient object detection and segmentation capabilities. Two model configurations were implemented: (1) Object detection, which identifies molecular species using bounding boxes; (2) Instance segmentation, which provides pixel-wise segmentation for precise molecular boundary identification.
Model training and evaluation
The YOLOv9 model was trained using a stochastic gradient descent (SGD) optimizer with momentum, and weight decay was applied for regularization. Training was performed for 100 epochs with an adaptive learning rate schedule. The following performance metrics were used for evaluation: F1 Score, which balances precision and recall for classification accuracy;
The implementation was developed in Python, using PyTorch as the deep learning framework. Training was conducted on an NVIDIA 3070Ti GPU with 8 GB VRAM, with a training time of approximately 20 min per full training cycle.
RESULTS AND DISCUSSION
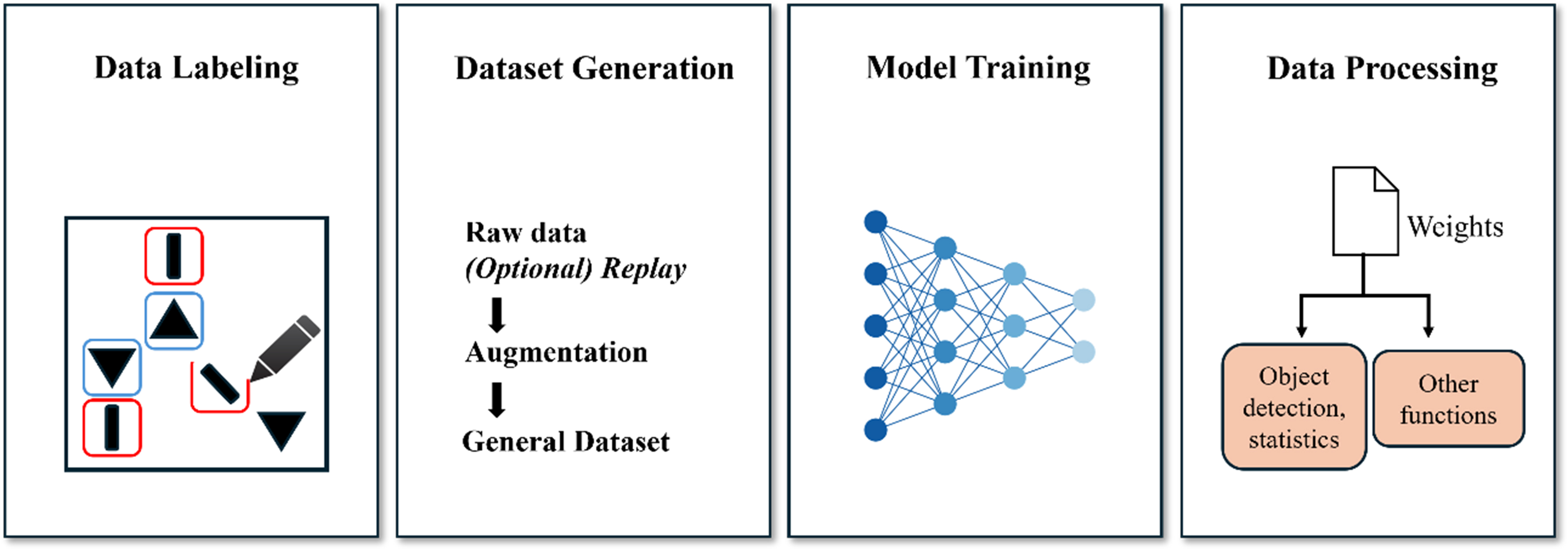
Our framework consists of four main components: data labeling, dataset generation, model training/prediction, and data processing. In the training section, users can choose between two models: object detection or instance segmentation, both of which are based on the YOLOv9 model [Supplementary Section 2]. The overall framework architecture is illustrated in Figure 1.
The process begins with the “Data Labeling” stage, where the data are annotated to provide ground truth for model training. Following this, the system checks whether the labeled data correspond to a new category. If a new category is detected, the system performs the “Load Class ID from Index Table” step to assign a unique ID, which is then used in the “Dataset Generation” step. The generated dataset serves as the foundation for model training, ensuring that all relevant categories are included. Additionally, when generating the dataset, users can enable the incremental learning feature to create a more generalized dataset. In the incremental learning phase, the model is updated with newly added data without complete retraining from scratch, significantly enhancing its adaptability, especially as new types of species and patterns are discovered in the experimental image data.
After dataset creation, the “Train & Detect” step utilizes the YOLOv9 model to perform both object detection and instance segmentation, identifying key features in the image data. YOLOv9, the latest version of the YOLO family of models[29], combines advanced detection algorithms with high efficiency, enabling accurate and fast object detection even for complex datasets. It improves upon its predecessors by incorporating more robust backbone networks, enhanced anchor-free mechanisms, and dynamic label assignment strategies, making it particularly suitable for detecting small-scale molecular features in high-resolution images. Once detection is completed, the results are processed by the “Data Processing” module, which extracts valuable insights such as the identified classes and the average area of detected instances. These insights are crucial for understanding surface nanostructures, molecular self-assembly, and the distribution of reactants and products. The seamless integration of incremental learning ensures that the model evolves continuously as new data become available, maintaining accuracy as experimental conditions change and allowing researchers to adapt their analysis to emerging surface phenomena. This combination of YOLOv9’s advanced detection capabilities and incremental learning provides a powerful framework for analyzing molecular systems.
During training, we found that hyperparameters have a significant impact on the model’s performance. Improper hyperparameter configurations can drastically reduce training efficiency, potentially wasting several hours without yielding satisfactory results. In contrast, an appropriate combination of hyperparameters can accelerate convergence, improve model stability, and significantly enhance overall performance. This effect becomes particularly evident in deep learning tasks, where training is computationally expensive. A poorly tuned learning rate or batch size can either cause divergence or lead to excessively slow convergence, while suboptimal augmentation or regularization parameters may result in underfitting or overfitting, making the training process inefficient and unstable. To address this challenge, it is crucial to adopt a systematic and automated hyperparameter optimization strategy that goes beyond manual trial-and-error. Given the large number of hyperparameters, manually adjusting each value is extremely time-consuming and practically impossible. Therefore, we employed a Bayesian optimization algorithm to search for the optimal combination of hyperparameters. Compared with conventional methods such as grid search and random search, Bayesian optimization can find near-optimal solutions with fewer iterations, especially in high-dimensional hyperparameter spaces. It builds a surrogate model [e.g., Gaussian process (GP)] to predict the behavior of the objective function and leverages an acquisition function [such as expected improvement (EI), probability of improvement, or upper confidence bound] to balance exploration of unknown regions and exploitation of promising areas, thereby avoiding inefficient or blind searches. Unlike grid search, which exhaustively evaluates every point in a predefined parameter grid, or random search, which samples configurations uniformly at random, Bayesian optimization intelligently selects the next point to evaluate based on prior knowledge from past observations. This approach is particularly efficient when each model evaluation is computationally expensive, as in our training pipeline, where a single training run can take several hours.
The surrogate model we utilized, GP, offers a probabilistic interpretation of the function being optimized. GP estimates not only the mean performance for any hyperparameter configuration but also the associated uncertainty, enabling the acquisition function to determine whether to explore new regions or exploit areas that are already known to yield favorable results. The EI criterion, in particular, evaluates the expected gain over the current best-known performance, guiding the search toward configurations with the highest probability of improving the model metric (
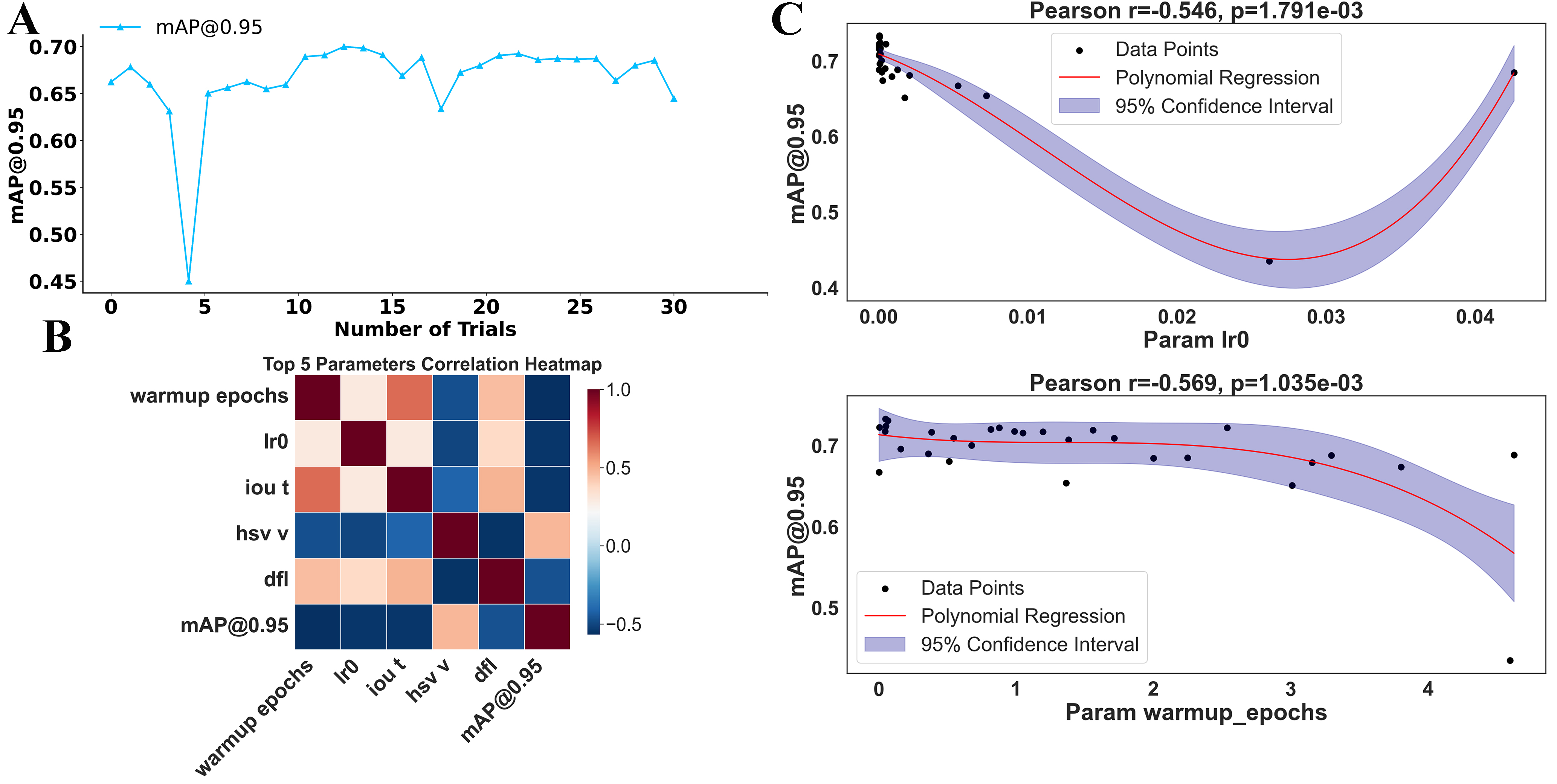
We constructed a search space from the selected hyperparameters and performed 30 iterations of Bayesian optimization. The selected hyperparameters[30] were chosen based on their relevance to model performance and training stability. From Figure 2A, we observed that the optimal configuration was achieved at the 13th iteration, where
Figure 2. Hyperparameter optimization and its impact on model performance. (A) Model performance (
In our program, we specifically incorporated the feature of incremental learning, and among the various incremental learning methods, we selected the most stable and convenienta replay mechanism[31]. The replay mechanism works by saving a portion of historical data during the training process and periodically retraining the model with both the historical data and new data in subsequent training sessions. This approach allows the model to retain knowledge of previously seen data while learning new information, thereby mitigating catastrophic forgetting.
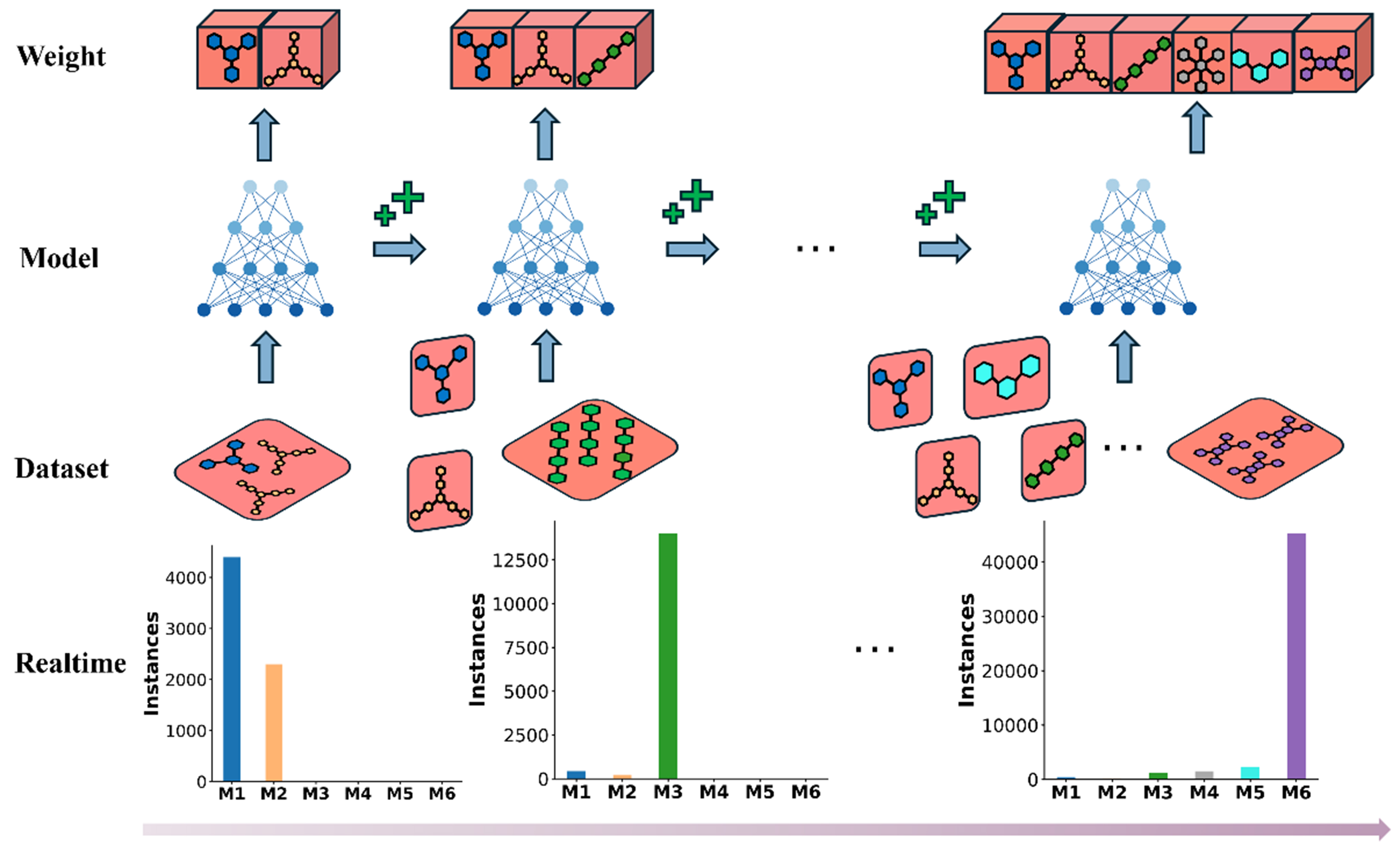
From Figure 3, the replay mechanism can be seen to select a small batch of historical data and combine it with the current training data for model training. This process not only enables the model to continuously adapt to new data but also ensures training stability, as the model “reviews” previously learned samples. As a result, the model avoids overfitting to the current samples while retaining knowledge acquired from earlier data. In this way, the model can gradually enhance its generalization ability and robustness during the incremental learning process, leading to more effective knowledge updates and applications.
Figure 3. Illustration of the replay mechanism in incremental learning. The top row (Weight) represents weight adjustments as more data are included. The second row (Model) illustrates the model training steps. The third row (Dataset) shows the dataset replay strategy, where historical data are incorporated into the training process. The bottom row (Realtime) presents the change in the number of instances from each category (M1 to M6) participating in training as a result of applying the replay strategy.
The advantage of the replay mechanism is that it allows the model to be continuously updated and optimized as new data become available, without the need for complete retraining from scratch. This greatly improves training efficiency and the flexibility of data handling. Specifically, for atomic-scale STM image analysis, the replay mechanism offers distinct advantages over other incremental learning strategies, such as knowledge distillation. STM images contain highly detailed pixel-level information - including fine variations in local contrast, adsorption geometry, and tip-induced electronic effects, which are physically meaningful and critical for molecular recognition. Distillation-based methods, which transfer feature distributions between teacher and student models, often compress or smooth out these subtle spatial and intensity variations, leading to a partial loss of nanoscale structural information. In contrast, replay revisits a small portion of real historical STM data during training, thereby directly preserving the original pixel intensity distribution and spatial correlations. This approach enables the model to maintain sensitivity to atomic-scale morphological and electronic features while learning new molecular systems, effectively mitigating catastrophic forgetting. As a result, the replay strategy not only aligns with the physical nature of STM imaging but also achieves superior stability and accuracy in nanoscale feature recognition.
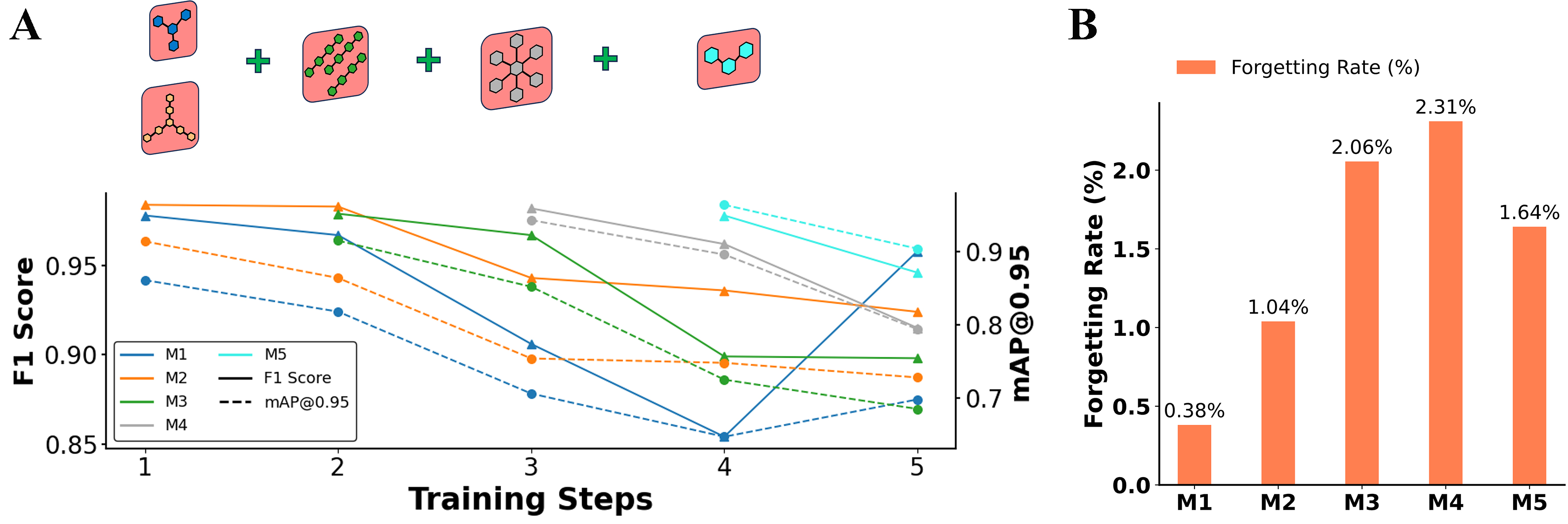
As shown in Figures 4 and 5A, five molecular categories (M1-M5) were selected to evaluate model performance under the replay mechanism. The model underwent five rounds of training, during which new categories of molecular image data were continuously introduced. Throughout this process, changes in F1 scores and
Figure 4. Model performance and forgetting rate across training steps. (A) Model performances over five training steps, represented by F1 scores and
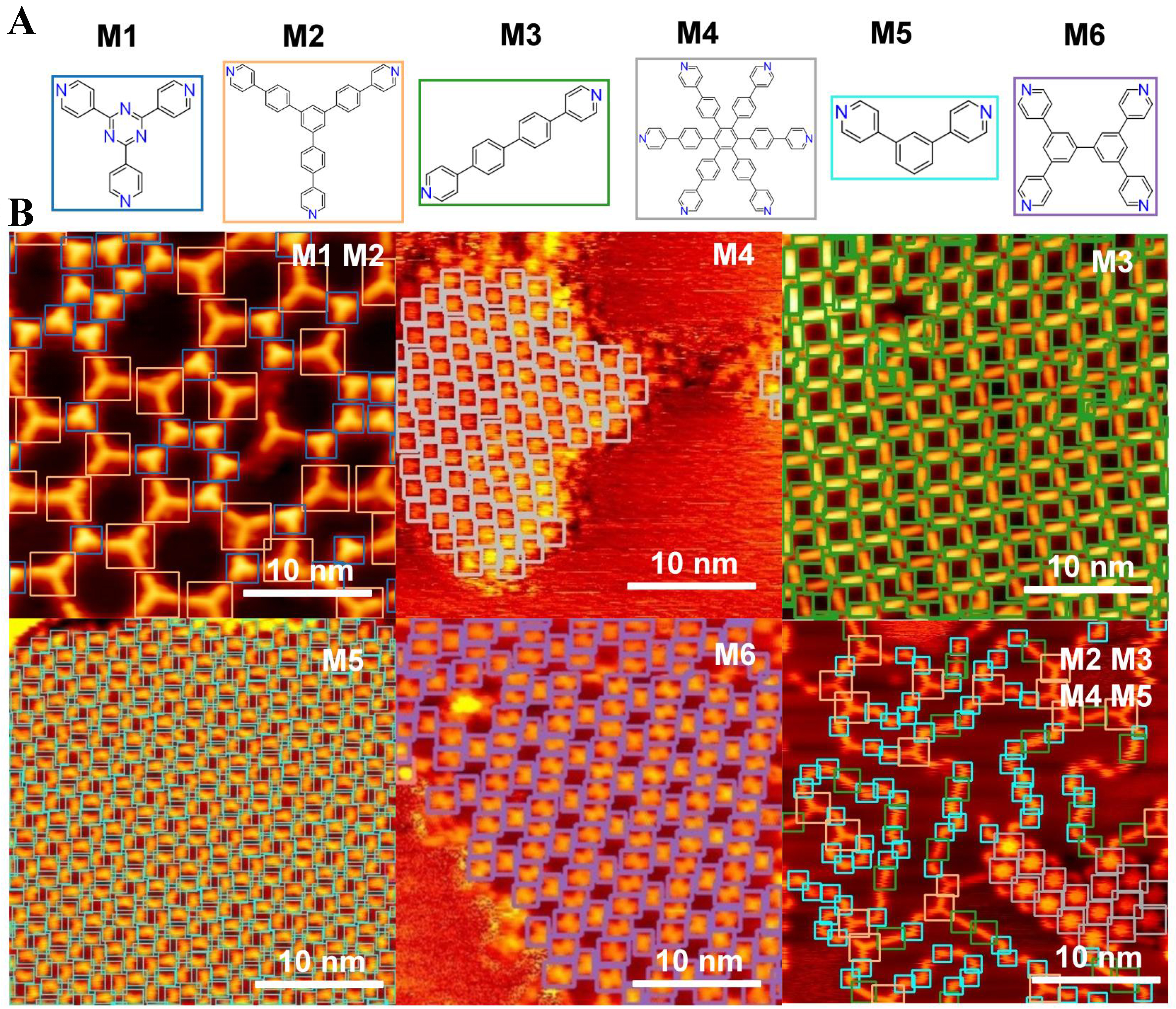
Figure 5. STM molecular images and prediction results. (A) Chemical structures of six molecular categories (M1-M6); (B) STM images and prediction results obtained using the well-trained model. The detected molecules are highlighted with color-coded markers: blue squares denote molecule M1, orange squares denote M2, green squares denote M3, grey squares denote M4, cyan squares denote M5, and purple squares denote M6. STM: Scanning tunneling microscopy.
where Pinitial represents the model performance on a specific category or task before incremental training (e.g., F1 score), and Pfinal represents the performance on the same category or task after training is completed.
As shown in Figure 4B, the forgetting rates for all categories are very low, with all values below 2.5%, demonstrating the effectiveness of the replay mechanism in mitigating catastrophic forgetting while maintaining the model performance across multiple categories. These results indicate that the training strategy effectively preserves previously learned knowledge, as reflected by the minimal performance degradation.
Figure 5 provides a more intuitive demonstration of the effectiveness of the incremental learning approach. These results are based on the model after five rounds of incremental learning, where the trained weights integrate the learned representations of six molecular categories (M1 to M6). Figure 5A displays six representative molecular categories (M1-M6), each corresponding to a specific precursor or coordination unit used in STM studies of on-surface molecular self-assembly, i.e., M1: 2,4,6-tri(pyridin-4-yl)-1,3,5-triazine; M2: 4,4’-(5’-(4-(pyridin-4-yl)phenyl)-[1,1’:3’,1’’-terphenyl]-4,4’’-diyl)dipyridine; M3: 4,4’-di(pyridin-4-yl)-1,1’-biphenyl; M4: hexakis[4-(4’-pyridyl)phenyl]benzene; M5: 4,4’-(1,3-phenylene)bis-pyridine; and M6: 3,3’,5,5’-tetra(pyridin-4-yl)-1,1’-biphenyl.
Using the trained weights, a single prediction was performed, and the results shown in Figure 5B demonstrate clear and accurate identification of the corresponding molecular structures. The predicted patterns align well with the molecular structures in Figure 5A, indicating that the model effectively captures and retains the features of all six categories without significant performance degradation. To further quantify this performance, the evaluation metrics indicate a precision of 0.959, recall of 0.931,
These findings further highlight the strength of the incremental learning strategy, which enables the model to continuously learn new molecular categories while retaining knowledge of previously learned ones. The predictions indicate that the model can handle the complexity of molecular assemblies and accurately detect and classify molecules, thus illustrating its robustness and generalization capability after incremental learning. The clear patterns and accurate bounding boxes in the predictions reflect the model’s ability to integrate diverse molecular features, making it a powerful tool for analyzing high-resolution images of complex molecular arrangements.
Batch effects arising from different STM imaging environments, such as variations in tunneling current stability, tip geometry, or detector sensitivity, may introduce distributional bias into the image data. To mitigate this issue, the dataset includes STM images collected under diverse experimental conditions on Au(111), Ag(111), and Cu(111) substrates. By incrementally introducing data from these different imaging environments while replaying earlier samples, the model continuously calibrates itself across multiple acquisition domains, thereby reducing the risk of bias. In addition to the experiments discussed above, the model’s performance was further evaluated on images of the same molecular species acquired on three different substrates, namely Au(111), Ag(111), and Cu(111) [Figure 6]. The results show that the incremental learning strategy not only allows the model to continuously adapt to new data but also exhibits strong generalization across diverse imaging conditions. Despite variations in surface structures, noise levels, and contrast among different substrates, the model successfully identified the target molecule with no noticeable degradation in detection accuracy. This outcome indicates that the replay mechanism effectively preserves the essential features of the learned molecule, enabling robust recognition across heterogeneous datasets. The ability to adapt to different substrates highlights the strong transferability and generalization capability of the incremental learning framework. In STM imaging, the substrate type plays a critical role in determining the overall image characteristics. For example, Au(111), Ag(111), and Cu(111) surfaces differ in lattice constants and electronic structures, leading to significant variations in background textures, contrast levels, and noise patterns in the resulting images. Conventional deep learning detection models are often sensitive to such differences; when trained exclusively on data from a single substrate, their performance typically declines significantly when tested on another[32]. To overcome this issue, the aforementioned incremental learning strategy combined with a replay mechanism was adopted.
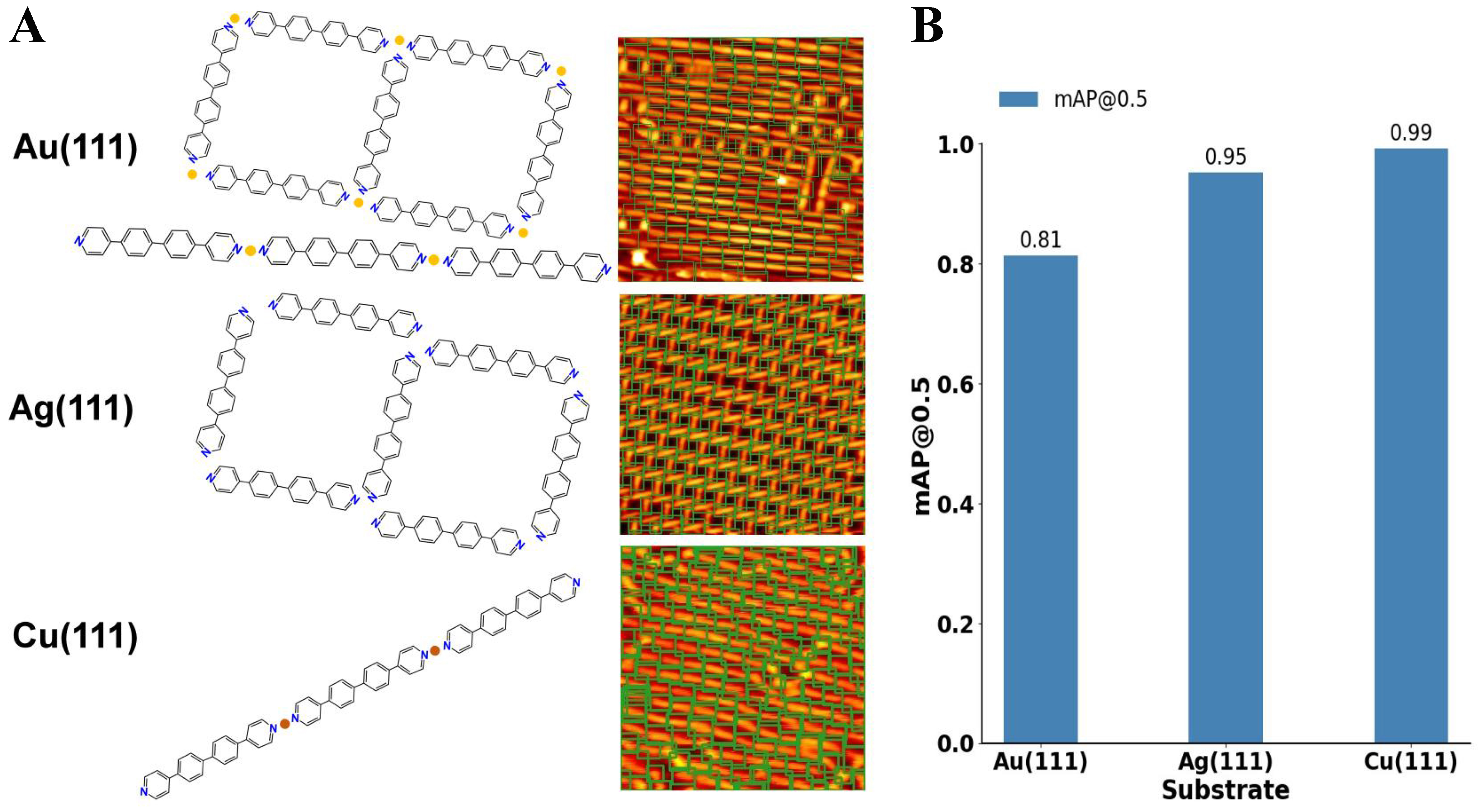
Figure 6. Performance of the incremental learning framework on the sample molecules across multiple metal substrates. (A) Visualization of molecular detection results for 4,4’-di(pyridin-4-yl)-1,1’-biphenyl in STM images acquired from Au(111), Ag(111), and Cu(111) surfaces. All STM images span 20 nm × 20 nm. Bounding boxes highlight the successfully identified molecules. In the model visualization, yellow and orange represent gold (Au) and copper (Cu) atoms, respectively; (B) Quantitative performance comparison across substrates using
In our experiments, we progressively introduced STM images of the target molecule obtained on Au(111), Ag(111), and Cu(111). At the initial stage, the model was trained solely on Au(111) data and achieved high detection accuracy. We then incrementally incorporated images from Ag(111) and Cu(111) substrates while simultaneously replaying Au(111) images to reinforce the retention of previously acquired knowledge. This strategy ensured that the model maintained strong recognition performance for Au(111) molecules while adapting to the unique imaging characteristics of Ag(111) and Cu(111). The final evaluation results demonstrate that the model maintains consistently high accuracy across all three substrates (with only minor variations in
As shown in Figure 6, the model demonstrates robust molecular recognition even when the substrate type changes, which typically introduces differences in background texture, surface electronic structure, and imaging noise. Despite these challenges, the incremental learning approach enables the model to retain the discriminative features of the molecule while adapting to substrate-induced variations. This robustness is largely attributed to the fact that the incremental training strategy does not simply overwrite previously learned weights with new data, but instead uses a balanced training approach in which data from both old and new substrates are sampled in each training epoch, with the old dataset comprising 10% of the training data. This design allows the model to maintain equilibrium between previously learned features and new information, resulting in stable recognition performance under diverse imaging conditions.
Our experiments confirm that even when Cu(111) images present higher noise levels and lower contrast, the model is still able to accurately detect and classify the molecule based on its essential structural features. Furthermore, variations in substrate properties not only affect image noise and contrast but can also influence the self-assembly behavior of molecules on the surface. For instance, 4,4’-di(pyridin-4-yl)-1,1’-biphenyl molecules may exhibit slightly different arrangements on Au(111), Ag(111), and Cu(111) due to differences in adsorption energy and intermolecular interactions, which in turn affect molecular packing density and orientation[33,34]. By incorporating data from multiple substrates, the incremental learning model learns to recognize key molecular structures and features regardless of such variations.
Lastly, we note an additional advantage of the incremental learning framework: improved training efficiency. Traditional approaches typically require training a separate model for each substrate, which is computationally costly and time-consuming. In contrast, our incremental approach dynamically updates a single set of model parameters, achieving a “train-once, adapt-to-many” capability. The replay mechanism also reinforces the retention of key features learned in earlier stages, ensuring that the model does not forget previously learned datasets while adapting to new conditions. This approach not only reduces computational overhead but also streamlines the entire training pipeline. These results emphasize the transferability of the learned representations and confirm that the model can accurately detect and classify molecules across heterogeneous environments without significant performance loss.
CONCLUSIONS
In this study, we developed a comprehensive YOLOv9-based program for object detection and instance segmentation of molecular STM images, specifically addressing the challenges associated with manual molecular image analysis in surface nanostructures. The program integrates several advanced features, including data labeling, dataset creation, model training, detection, and analysis, with a strong focus on supporting incremental learning. The inclusion of a replay mechanism for incremental learning has proven highly effective in mitigating catastrophic forgetting, allowing the model to retain previously learned information while adapting to new data. By replaying a small portion of the old dataset, the system reduces the rate of forgetting while maintaining efficiency. Additionally, Bayesian optimization was employed to fine-tune hyperparameters, further enhancing model performance. The results demonstrate that our approach provides a scalable, efficient, and robust solution for the analysis of high-resolution molecular STM images, enabling researchers to accurately interpret complex surface reactions beyond the limitations of manual analysis. To support the practical implementation of this workflow, a software framework was developed, as described in Supplementary Section 5. Overall, the integration of machine learning techniques, including incremental learning and hyperparameter optimization, offers promising improvements in the accuracy, consistency, and scalability of molecular image analysis, paving the way for further innovations in the field of surface chemistry.
DECLARATIONS
Acknowledgments
The authors would like to thank the developers of Bgolearn (https://doi.org/10.48550/arXiv.2601.06820) for providing the Bayesian global optimization framework used to optimize the hyperparameters during the SciBERT model training process.
Authors’ contributions
Research conception and study design, data analysis and interpretation, and software framework development and machine learning model implementation: Yuan, S.; Zhu, Z.
Data acquisition, annotation pipeline development, and result visualization: Lu, J.
Manuscript drafting, critical revision, and review of important intellectual content: Yuan, S.; Cai, L.
Administrative, technical, and material support, project supervision, and funding acquisition: Sun, Q.
Availability of data and materials
The code and trained models supporting the findings of this study are publicly available in a GitHub repository: https://github.com/yuanke75/yolo_program_for_spm. All datasets generated and analyzed during this study are included in the repository as processed files and annotation scripts.
AI and AI-assisted tools statement
During the preparation of this manuscript, the AI tool ChatGPT (GPT-4o, released 2024-05-13) was used solely for language editing. The tool did not influence the study design, data collection, analysis, interpretation, or the scientific content of the work. All authors take full responsibility for the accuracy, integrity, and final content of the manuscript.
Financial support and sponsorship
This work was supported by the National Natural Science Foundation of China (No. 22302120).
Conflicts of interest
All authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
Supplementary Materials
REFERENCES
1. Lo, S.; Baird, S. G.; Schrier, J.; et al. Review of low-cost self-driving laboratories in chemistry and materials science: the “frugal twin” concept. Digit. Discover. 2024, 3, 842-68.
2. Wang, Z.; Qin, M.; Zhang, P.; et al. High throughput characterization method of electrical and phonon properties by dielectric resonant spectroscopy. Mater. Genome. Eng. Adv. 2025, 3, e70010.
3. Xiang, J.; Li, Y.; Zhang, X.; He, Y.; Sun, Q. Local large language model-assisted literature mining for on-surface reactions. Mater. Genome. Eng. Adv. 2025, 3, e88.
4. Li, Y.; Liu, X.; Wang, X.; Xie, W.; Qiu, D.; Yang, J. Effect of material properties on the thermal responses of the carbonization and pyrolysis layers of polymer matrix composites for charring-ablators. J. Mater. Inf. 2025, 5, 31.
5. Wang, W. Y.; Zhang, S.; Li, G.; et al. Artificial intelligence enabled smart design and manufacturing of advanced materials: the endless Frontier in AI+ era. Mater. Genome. Eng. Adv. 2024, 2, e56.
6. Jung, T. A.; Schlittler, R. R.; Gimzewski, J. K. Conformational identification of individual adsorbed molecules with the STM. Nature 1997, 386, 696-8.
7. Meng, T.; Lu, Y.; Lei, P.; et al. Self-assembly of triphenylamine macrocycles and co-assembly with guest molecules at the liquid-solid interface studied by STM: influence of different side chains on host-guest interaction. Langmuir 2022, 38, 3568-74.
8. Wyrick, J.; Wang, X.; Namboodiri, P.; et al. Enhanced atomic precision fabrication by adsorption of phosphine into engineered dangling bonds on H-Si using STM and DFT. ACS. Nano. 2022, 16, 19114-23.
9. Wang, L.; Xia, Y.; Ho, W. Atomic-scale quantum sensing based on the ultrafast coherence of an H2 molecule in an STM cavity. Science 2022, 376, 401-5.
10. Moreno, D.; Parreiras, S. O.; Urgel, J. I.; et al. Engineering periodic dinuclear lanthanide-directed networks featuring tunable energy level alignment and magnetic anisotropy by metal exchange. Small 2022, 18, e2107073.
11. Lyu, C. K.; Gao, Y. F.; Gao, Z. A.; et al. Synthesis of single-layer two-dimensional metal–organic frameworks M3(HAT)2 (M=Ni, Fe, Co, HAT=1,4,5,8,9,12-hexaazatriphenylene) using an on-surface reaction. Angew. Chem. Int. Ed. Engl. 2022, 61, e202204528.
12. Liu, J.; Li, J.; Xu, Z.; et al. On-surface preparation of coordinated lanthanide-transition-metal clusters. Nat. Commun. 2021, 12, 1619.
13. Di Giovannantonio, M.; Fasel, R. On-surface synthesis and atomic scale characterization of unprotected indenofluorene polymers. J. Polym. Sci. 2022, 60, 1814-26.
14. Wang, J.; Niu, K.; Xu, C.; et al. Influence of molecular configurations on the desulfonylation reactions on metal surfaces. J. Am. Chem. Soc. 2022, 144, 21596-605.
15. Kinikar, A.; Di Giovannantonio, M.; Urgel, J. I.; et al. On-surface polyarylene synthesis by cycloaromatization of isopropyl substituents. Nat. Synth. 2022, 1, 289-96.
16. Liu, L.; Klaasen, H.; Witteler, M. C.; et al. Polymerization of silanes through dehydrogenative Si-Si bond formation on metal surfaces. Nature. Chem. 2021, 13, 350-7.
17. Mallada, B.; de la Torre, B.; Mendieta-Moreno, J. I.; et al. On-surface strain-driven synthesis of nonalternant non-benzenoid aromatic compounds containing four- to eight-membered rings. J. Am. Chem. Soc. 2021, 143, 14694-702.
18. Zhu, X.; Liu, Y.; Pu, W.; et al. On-surface synthesis of C144 hexagonal coronoid with zigzag edges. ACS. Nano. 2022, 16, 10600-7.
19. Hellerstedt, J.; Cahlík, A.; Švec, M.; Stetsovych, O.; Hennen, T. Counting molecules: python based scheme for automated enumeration and categorization of molecules in scanning tunneling microscopy images. Softw. Impacts. 2022, 12, 100301.
20. Krull, A.; Hirsch, P.; Rother, C.; Schiffrin, A.; Krull, C. Artificial-intelligence-driven scanning probe microscopy. Commun. Phys. 2020, 3, 54.
21. Milošević, D.; Vodanović, M.; Galić, I.; Subašić, M. Automated estimation of chronological age from panoramic dental X-ray images using deep learning. Expert. Syst. Appl. 2022, 189, 116038.
22. Li, J.; Telychko, M.; Yin, J.; et al. Machine vision automated chiral molecule detection and classification in molecular imaging. J. Am. Chem. Soc. 2021, 143, 10177-88.
23. Gordon, O. M.; Hodgkinson, J. E. A.; Farley, S. M.; Hunsicker, E. L.; Moriarty, P. J. Automated searching and identification of self-organized nanostructures. Nano. Lett. 2020, 20, 7688-93.
24. Kang, J.; Yoo, Y. J.; Park, J. H.; et al. DeepGT: deep learning-based quantification of nanosized bioparticles in bright-field micrographs of gires-tournois biosensor. Nano. Today. 2023, 52, 101968.
25. Faraz, K.; Grenier, T.; Ducottet, C.; Epicier, T. Deep learning detection of nanoparticles and multiple object tracking of their dynamic evolution during in situ ETEM studies. Sci. Rep. 2022, 12, 2484.
26. Newby, J. M.; Schaefer, A. M.; Lee, P. T.; Forest, M. G.; Lai, S. K. Convolutional neural networks automate detection for tracking of submicron-scale particles in 2D and 3D. Proc. Natl. Acad. Sci. U. S. A. 2018, 115, 9026-31.
27. Zhu, Z.; Yuan, S.; Yang, Q.; et al. Autonomous scanning tunneling microscopy imaging via deep learning. J. Am. Chem. Soc. 2024, 146, 29199-206.
28. Seifert, T. J.; Stritzke, M.; Kasten, P.; et al. Chirality detection in scanning tunneling microscopy data using artificial intelligence. Small. Methods. 2024, 8, e2400549.
29. Wang, C. Y.; Yeh, I. H.; Liao, H. Y. M. YOLOv9: learning what you want to learn using programmable gradient information. arXiv 2024, arXiv:2402.13616. Available online: https://doi.org/10.48550/arXiv.2402.13616. (accessed 27 Mar 2026).
30. Gao, T.; Gao, J.; Zhang, J.; Song, B.; Zhang, L. Development of an accurate “composition-process-properties” dataset for SLMed Al-Si-(Mg) alloys and its application in alloy design. J. Mater. Inf. 2023, 3, 6.
31. Zhou, D. W.; Wang, Q. W.; Qi, Z. H.; Ye, H. J.; Zhan, D. C.; Liu, Z. Class-incremental learning: a survey. IEEE. Trans. Pattern. Anal. Mach. Intell. 2024, 46, 9851-73.
32. Pan, S. J.; Yang, Q. A survey on transfer learning. IEEE. Trans. Knowl. Data. Eng. 2010, 22, 1345-59.
33. Otero, R.; Gallego, J. M.; de Parga, A. L.; Martín, N.; Miranda, R. Molecular self-assembly at solid surfaces. Adv. Mater. 2011, 23, 5148-76.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data
















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].