Enhanced multi-tuple extraction for materials: integrating pointer networks and augmented attention
Abstract
Extracting reliable, tuple-level information from materials texts is essential for data-driven materials design, yet multi-tuple sentences remain difficult due to intertwined semantics, syntactic complexity, and sparse supervision in higher-density cases. In this study, we address these challenges by formulating information extraction as an integrated process that couples entity extraction with tuple allocation. The framework combines an entity extraction module based on bidirectional encoder representations from transformers (MatSciBERT) with pointer networks and an allocation module that models inter- and intra-entity attention to enforce tuple coherence. Using the mechanical properties of multi-principal element alloys as a case study, we define the target schema and evaluate exact match tuple accuracy. Our experiments demonstrate F1 scores of 0.96, 0.95, 0.85, and 0.75 on datasets containing one to four tuples per sentence, and 0.85 on a randomly curated set. Ablation studies show that the allocation module is most critical, with inter-entity attention contributing more than intra-entity attention. Error analysis attributes the density-related performance decline mainly to semantic overlap and syntactic complexity, with upstream extraction errors more prominent under sparse supervision and allocation errors concentrated in structurally complex templates. This approach delivers precise, structured outputs suitable for downstream analysis and offers a domain-adaptable alternative to prompt-based large models when strict correctness is required.
Keywords
INTRODUCTION
The traditional “trial-and-error” paradigm in the development of new materials is time-consuming and expensive. However, with emerging technologies such as artificial intelligence, a dual-driven scientific artificial intelligence strategy that combines both models and data has been widely applied in the design and development of novel materials[1,2], as well as in elucidating the interrelationships between structure and performance[3-5]. This paradigm has unveiled considerable potential for the efficient design and optimization of materials[6-9]. Undoubtedly, data remain a foundational and critical element in achieving the aforementioned objectives. However, for certain material properties, particularly those pertaining to performance under service conditions, available data remain scarce. The acquisition of high-quality data by labor incurs significant costs, making it difficult to meet the requirements for training models that demand high accuracy and robust performance.
The scientific literature encompasses a vast array of peer-reviewed, high-quality, and relatively reliable data, serving as a crucial resource. However, much of this information exists in unstructured text within diverse sources, such as textbooks, material handbooks, patents, and research articles, rendering it unsuitable for direct use as structured data. Consequently, techniques for the automatic extraction of materials-related data and information present a promising opportunity to construct extensive databases for machine learning and data-driven methodologies.
Natural language processing (NLP) is an interdisciplinary field that integrates linguistics and artificial intelligence. One of its most prevalent applications is the automated extraction of structured information, facilitating the efficient mining of data from semi-structured tables and unstructured texts[10]. Numerous studies have reported various methodologies for mining structured information within specific domains of materials science[11,12], with a general trend moving from generic approaches to specialized techniques. Initially, these processes relied on traditional processing pipelines, generally including named entity recognition (NER) and relation extraction[13], such as ChemDataExtractor[14] for chemical information extraction. Specific techniques in these workflows include look-ups, rule-based methods[13], and machine learning approaches, which have gradually shifted toward more sophisticated deep neural models[15], such as long short-term memory (LSTM) networks[13]. Due to the excellent performance of the pre-training and fine-tuning paradigm, many works now rely on pre-trained models for information extraction. By pre-training on a substantial corpus of materials-related literature, these models enable a deeper analysis of relationships among chemical compositions, structural features, and corresponding properties. Many variations of bidirectional encoder representations from transformers (BERT)[16], such as MatSciBERT[17], MaterialsBERT[18], BioBERT[19], and BatteryBERT[20], have shown improved performance on downstream tasks. Large language models (LLMs), such as Generative Pre-trained Transformer (GPT)-4 and GPT-5[21,22], Large Language Model Meta AI (LLaMA)[23], Pathways Language Model (PaLM)[24], Google’s Gemini model[25], and Fine-tuned LAnguage Net (FLAN)[26], also exhibit great understanding of textual information. When prompted appropriately, LLMs can extract targeted information and data from literature. For example, Dagdelen et al. used multiple LLMs to extract structured information[27].
However, despite extensive research on structured information extraction, effective methodologies for multi-tuple extraction remain notably lacking. A tuple typically consists of a set of related elements that represent structured information, often in the form of (subject, predicate, object) or (entity, attribute, value), although in some cases, tuples may contain more elements. The presence of multiple tuples within text complicates the accurate assignment of extracted entities to their corresponding tuples. This challenge is compounded by contextual ambiguity, diverse modes of expression, and the difficulty of obtaining high-quality annotated datasets. Even though LLMs demonstrate outstanding capabilities in NLP tasks, they still exhibit limitations in understanding multi-tuple scientific texts and may produce hallucinations. Moreover, LLMs typically require substantial computational resources for both training and inference, which may limit their cost-effectiveness and efficiency in practical applications of data mining and knowledge extraction.
In this study, we explore multi-tuple extraction of mechanical properties from alloys, mainly focusing on multi-principal element alloys (MPEAs). We propose a domain-specific framework that integrates an entity extraction model combining MatSciBERT with pointer networks and an entity allocation model employing inter-entity and intra-entity attention mechanisms. To enable principled evaluation across varying linguistic densities, we compile and annotate a sentence-level corpus from the MPEA literature, dividing sentences according to the number of tuples they contain, and assess performance at the tuple level to reflect scientific completeness rather than isolated spans. To begin, we formalize tuple construction as a two-stage problem, consisting of entity extraction followed by allocation. Next, we conduct ablation studies to separate the contributions of the allocation module and its inter- and intra-entity attention, quantifying their roles in precision, pairing stability, and robustness as tuple density increases. Additionally, we perform error analyses to characterize residual failure modes, including cross-assignment between properties and conditions, semantic overlap among entities, and syntax-induced ambiguity, and we relate these errors to specific architectural choices. Finally, we examine substitutes for the domain encoder and overall paradigm by replacing MatSciBERT with SciBERT and BERT and by benchmarking against LLMs under controlled prompts, thereby clarifying the advantages of a domain-specific, scientifically faithful design for precise multi-tuple alignment.
MATERIALS AND METHODS
Workflow overview
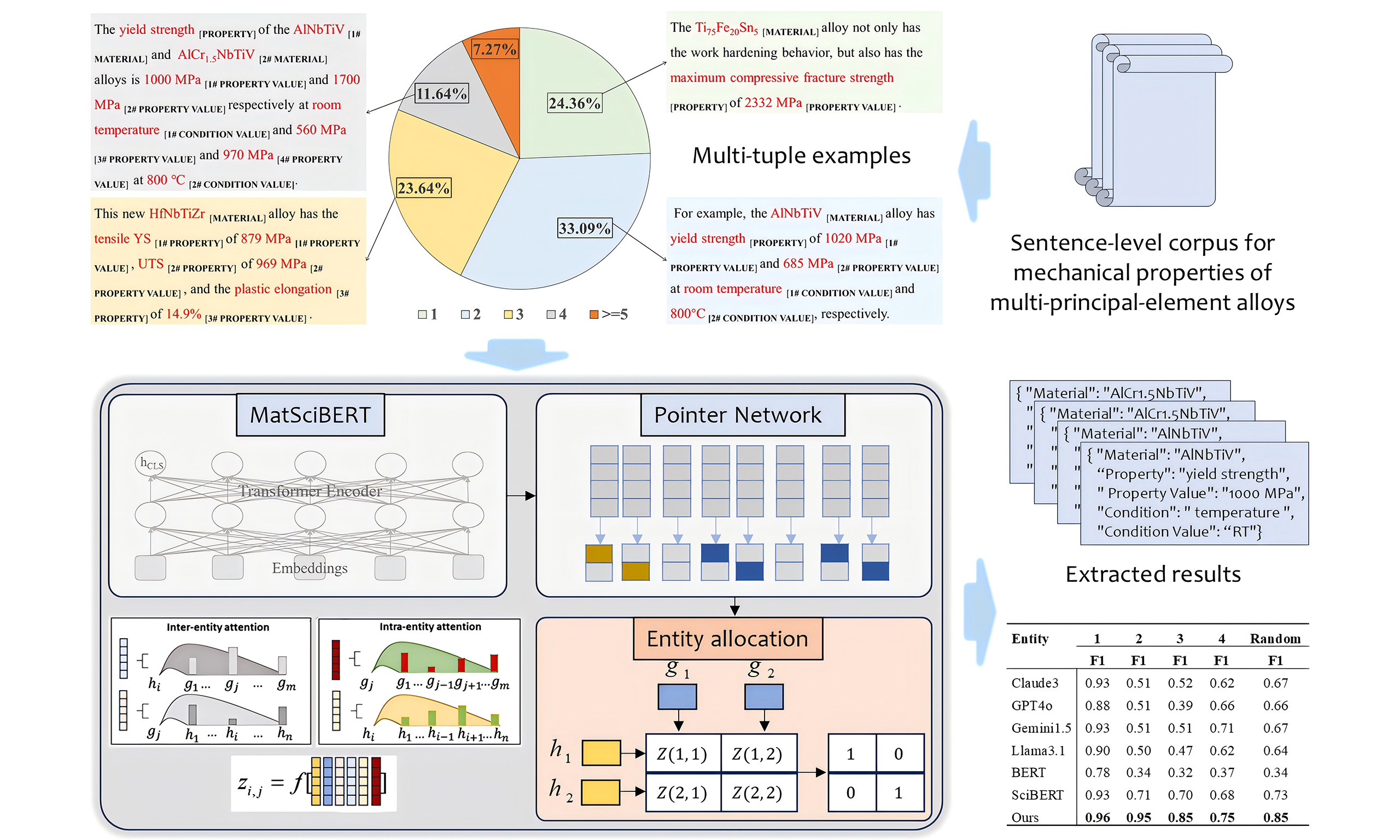
Figure 1 presents the overall workflow of the study. Initially, we constructed a corpus comprising over 200 full-text papers containing the keywords “multi-principal element alloy” and “mechanical properties”. From this corpus, we extracted sentences describing alloy properties and annotated five distinct types of entities within these sentences: MATERIAL, PROPERTY, PROPERTY VALUE, CONDITION, and CONDITION VALUE. This process yielded 255 sentences containing varying numbers of tuples, resulting in a total of 568 tuples stored in a JSON-formatted file. The modeling pipeline proceeds in two stages: the first stage extracts entity spans, and the second stage assigns entities to complete tuples within each sentence. Detailed specifications of the annotation protocol, model architecture, and training procedures are provided in the subsequent subsections.
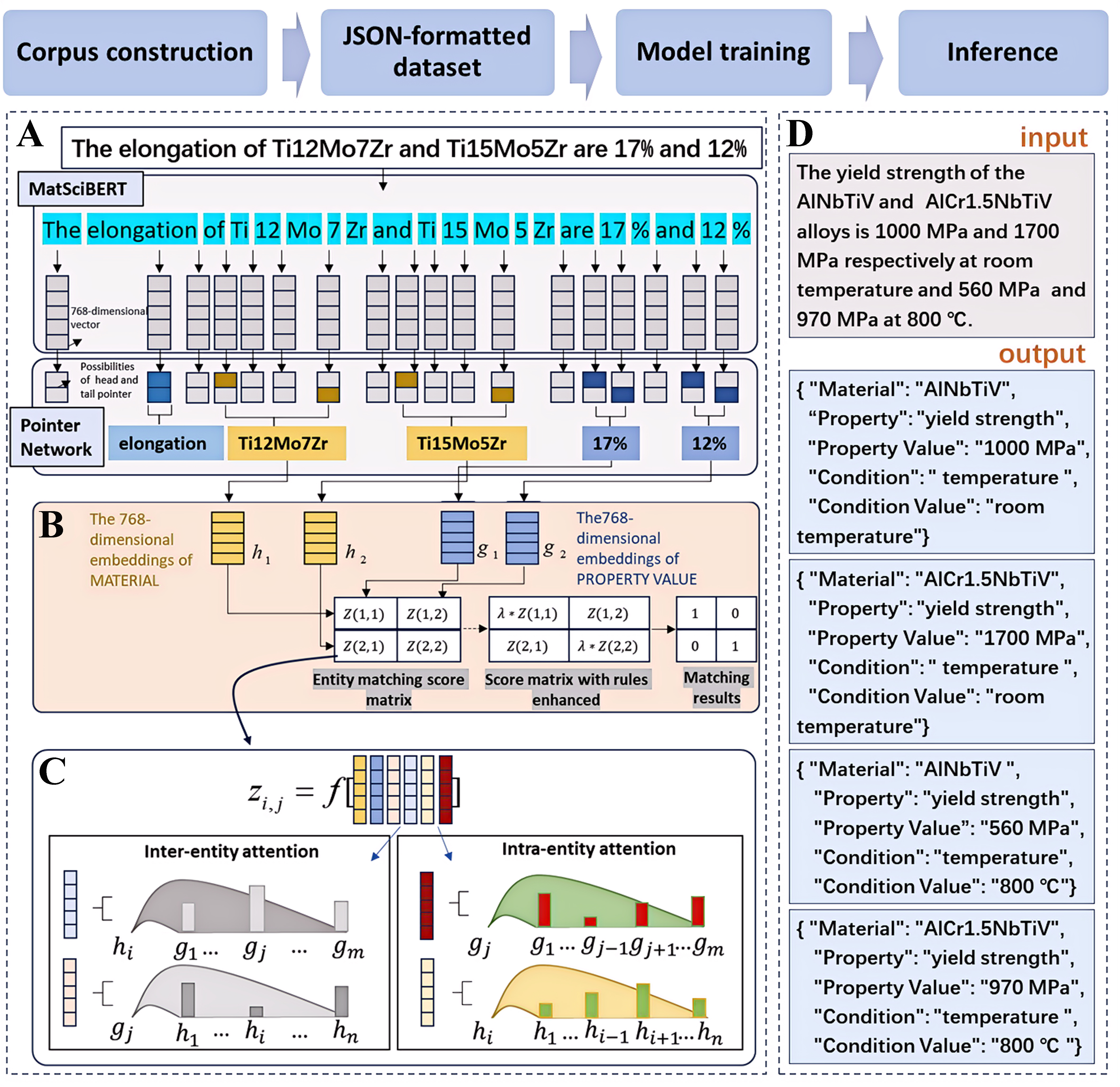
Figure 1. Workflow and framework of the proposed model for extracting and allocating entities. The upper section of the figure presents the workflow, beginning with the retrieval of full-text research articles from Elsevier, followed by the construction of a specialized corpus. Sentences are then extracted and annotated to obtain a JSON-formatted dataset, and the process concludes with model training and inference. The model is primarily composed of two components: entity extraction and entity allocation. (A) Entity Extraction: This component integrates MatSciBERT and a pointer network. MatSciBERT first tokenizes the input sentence and generates vector representations for each token. The pointer network then computes the probability of each token serving as the head or tail of a specific entity, thereby identifying entities based on these probabilities; (B) Entity Allocation: This component assesses whether entities of different types belong to the same tuple using an entity matching score matrix. During model inference, the matching likelihood of entities in corresponding order can be enhanced by multiplying the diagonal elements of the matrix by a parameter; (C) Entity Matching Score Matrix: Each element of the matrix represents a combination of six vectors. The first two vectors correspond to the vector representations of the two entities, while the remaining four vectors are derived from two attention mechanisms: intra-entity and inter-entity attention. Intra-entity attention focuses on the attention distribution among different types of entities, whereas inter-entity attention concentrates on the attention distribution within the same type of entity; (D) Inference: An example of four-tuple extraction is illustrated.
Literature acquisition
The materials science literature used in this study was obtained from Elsevier with publisher consent. This work mainly focuses on high-entropy alloys. A large amount of papers relevant to high-entropy alloys were retrieved, and the top 200 papers were selected and downloaded in HTML/XML format based on their relevance and timeliness. These papers were then converted into text files to form a corpus for subsequent processing.
Preprocessing and annotation
In this stage, sentences describing significant alloy properties were identified from the corpus, and all tuples within these sentences were annotated with scientifically accurate meanings as gold labels. Five entity types were defined.
MATERIAL denotes spans referring to specific alloys, typically compositional formulae (e.g., Ti–30Nb–1Mo–4Sn) or established abbreviations (e.g., C.P. Ti). PROPERTY denotes spans representing a material property, such as elastic modulus, tensile strength, or yield strength. Synonymous labels follow the authors’ original wording within each sentence (e.g., UTS vs. ultimate tensile strength) to preserve textual fidelity. PROPERTY VALUE denotes numerical values with units corresponding to the property, including inequality and range expressions (e.g., over 800 MPa and > 40%). CONDITION denotes measurement conditions that affect the property (e.g., temperature). Its inclusion is essential due to the sensitivity of certain properties to measurement conditions, and its absence would render the entire tuple introduction incomplete. CONDITION VALUE denotes the numerical value and unit for a given condition. It is worth noting that our dataset exhibits varying entity frequencies, particularly for CONDITION and CONDITION VALUE, which occur less frequently than other entity types. This frequency variation is not attributable to subjective factors but reflects the common practice in materials science literature, where most experiments are conducted under standard conditions (room temperature and atmospheric pressure), leading authors to omit these default parameters. Thus, tuple annotation only allows for the absence of CONDITION and CONDITION VALUE. Tuples missing information in the first three entity types are not considered gold labels. Given the stringent requirements and laborious nature of data annotation, a subset of 568 golden tuples was extracted from 255 sentences within the corpus.
Dataset construction and splits
To evaluate performance as tuple density increases, sentences were categorized according to the number of gold tuples per sentence: 1, 2, 3, and ≥ 4. Sentences containing more than four tuples were excluded due to their scarcity. Because categories with 1, 2, or 3 tuples per sentence predominate, 15% of each of these categories was randomly sampled to form their test subsets. The 4-tuple category is smaller; therefore, to preserve sufficient training coverage, 10% was sampled for testing. For the Random dataset, 10% was also sampled following the same principle. The remaining data were split into training and validation sets at a 9:1 ratio after reserving the test portion.
Entity extraction
As shown in Figure 1A, we adopt a pointer-based span extraction method over contextual token embeddings, retaining the original formulation and notation except where clarification is necessary. Let the input sentence be t = [t1, …, tn]. The encoder produces contextual vectors Xt for each token[17]:
where MatSciBERT is a 12-layer, 768-hidden, 12-head transformer pre-trained on materials science literature. SciBERT and BERT-base are interchangeable backbones with identical architectures. Replacement experiments are conducted using the same extraction architecture and hyperparameters. Details about model selection and comparative outcomes are reported in the Results and Discussion section.
After obtaining the vector representations, a pointer network is used to train two binary classifiers per entity type to predict the start and end positions. Specifically, the model calculates the probability that each token, generated by MatSciBERT, corresponds to the start or end of the desired entity. Fully connected layers and activation functions are used to calculate the probabilities of the head and tail pointers[28]:
where Psmaterial(t) and Pematerial(t) are the probabilities that token t is the start and end token of a MATERIAL entity, respectively. Wsmaterial, bs, Wematerial, and be are matrices of learnable parameters specific to the head pointer and tail pointer, respectively. Furthermore, to calculate probabilities for other entity types, new learnable parameters are introduced. In summary, the pointer network essentially trains five sets of binary classifiers to determine whether a token is a head or tail pointer.
After computing the probabilities, tokens with values exceeding predefined thresholds (manually set hyperparameters) are considered pointers. Therefore, a list containing only 0s and 1s is generated, where each position represents whether the corresponding token is a pointer (1) or not (0)[28]:
where βsmaterial and βematerial are thresholds specific to the MATERIAL entity type.
As for training, the objective is to minimize the following cross-entropy loss function[29]:
where T is the set of entity types, t is the token sequence, and Lsτ(t) and Leτ(t) are gold labels indicating whether token t is a head or tail pointer. During inference, text spans corresponding to different entity types are extracted using a simple heuristic method. Specifically, each head pointer is paired with the nearest subsequent tail pointer to form an entity span.
Entity allocation
To resolve pairing ambiguities in multi-tuple extraction, the second stage defines an entity allocation task [Figure 1B], which determines whether two entities of different types belong to the same tuple. We do not predict multi-class relation labels; instead, we perform binary matching. To reduce the risk that low-frequency types are dominated by parameters tuned for high-frequency types, the extraction stage uses five independent binary pointer heads rather than a single multi-class head. In the allocation stage, different type combinations use separate parameter tensors. This design preserves dedicated capacity for low-frequency entities such as CONDITION and CONDITION VALUE, and maintains interpretability and stability under imbalanced training distributions.
To enable the model to learn both correct and incorrect pairing patterns, we construct an entity matching score matrix that enumerates and scores all candidate pairs for supervised learning. Consider two potentially confusable types within a sentence, with n and m entities, denoted as [h1, h2, ..., hn] and [g1, g2, ..., gm]. When calculating inter-attention representations, we first determine the semantic correlation between the two entity representations. Taking hi and gj as an example, we have[29]:
where Sij is the semantic correlation between hi and gj, σ denotes the dot product operator, and d represents the embedding dimension of hi and gj[29].
Based on Sij, we apply an inter-attention mechanism to generate gj-aware representations for hi and hi-aware representations for gj[29]:
where Aig2h and Ajh2g are the updated representations of hi and gj, respectively. They are also represented by the vectors in the upper box of Figure 1C.
Intra-entity attention allows the model to learn implicit relationships among entities of the same type. For example, among PROPERTY VALUE entities, relationships such as magnitude or unit consistency can help avoid confusion between PROPERTY VALUE and PROPERTY. The calculation methods for the two attention mechanisms are similar. For hi, we have[29]:
where μij is the semantic correlation between hi and hj, σ denotes the dot product operator, d represents the embedding dimension of hi, and Aih2h is the updated representation of hi. For gj, we have[29]:
where vjk is the semantic correlation between gj and gk and Ajg2g is the updated representation of gj.
Finally, for each pair of potentially confusable entities, we concatenate the six resulting vector representations and transform the entity allocation task into a binary classification problem[29]:
where
As for training, the objective is to minimize the following cross-entropy function[29]:
where
To encode common order constraints, when two entity types appear in equal numbers within a sentence and correct alignments typically follow textual order, we add a small diagonal bias λ to the matching matrix during inference. This favors order-consistent matches without overriding the learned scores; λ is tuned on the validation set.
Model and training setup
Training and evaluation are conducted on four NVIDIA RTX 3090 GPUs. Hyperparameters follow established practice for finetuning pre-trained language models for information extraction. For entity extraction, the model is fine-tuned for up to 50 epochs using the AdamW optimizer, with a learning rate of 1e-5 and a batch size of 16. The learning rate follows standard practice for BERT-based models to avoid catastrophic forgetting while enabling effective adaptation. For entity allocation, the model is fine-tuned for up to 50 epochs using the AdamW optimizer, with a learning rate of 1e-5 and a batch size of 8. The reduced batch size accommodates the computational complexity of the allocation module while maintaining training stability.
RESULTS AND DISCUSSION
Multi-tuple analyses
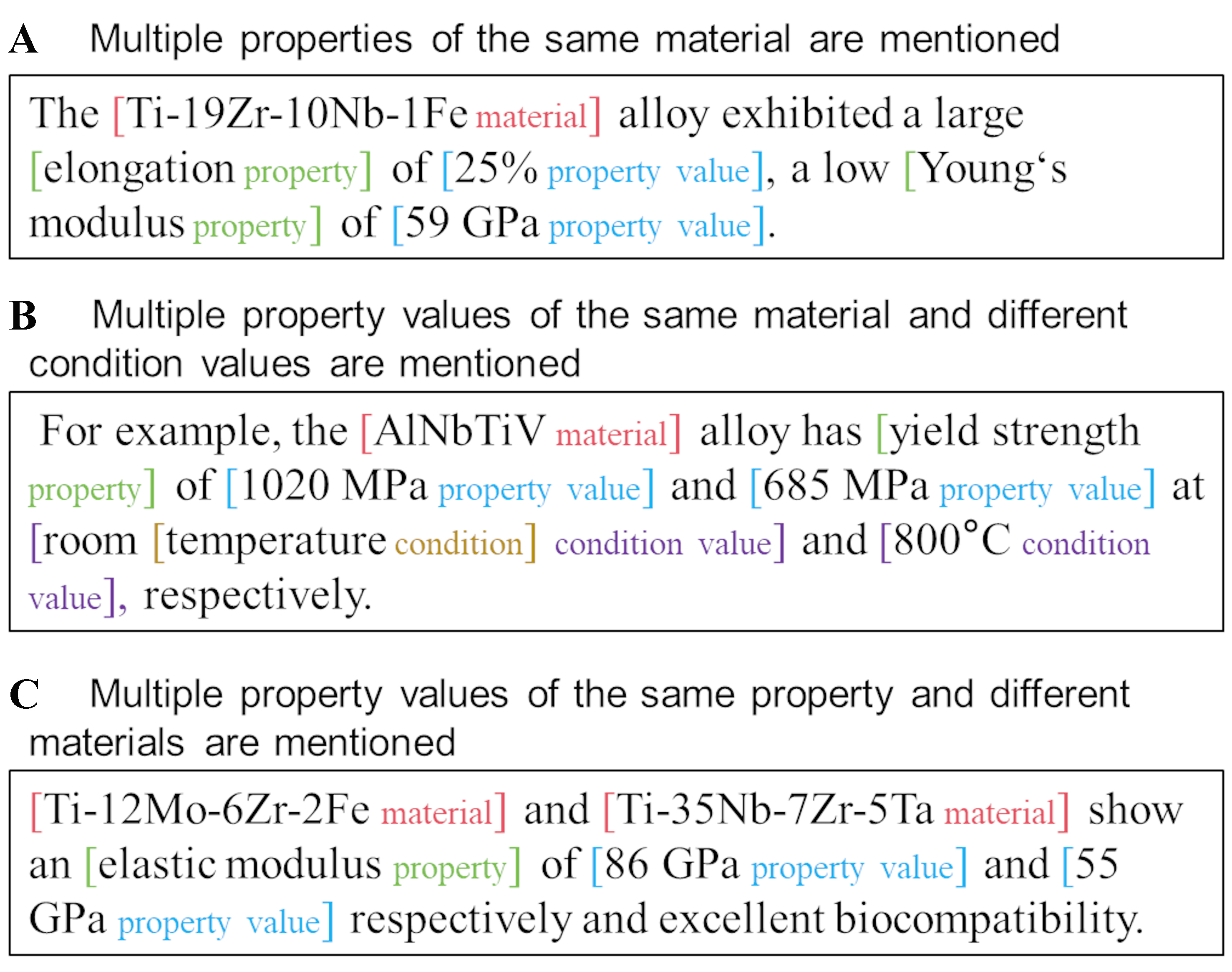
We describe several prevalent multi-tuple patterns observed in the aforementioned scenarios, as illustrated in Figure 2. These patterns include multiple properties of the same material, various property values for the same material under different conditions, and multiple values of the same property across different materials. Furthermore, these common patterns can be interwoven within more complex textual contexts, which are frequently encountered in the literature pertaining to diverse materials. For instance, from the sentence depicted in Figure 2B, two tuples can be extracted: [“AlNbTiV”, “yield strength”, “1,020 MPa”, “temperature”, “room temperature”] and [“AlNbTiV”, “yield strength”, “685 MPa”, “temperature”, “800 °C”]. In this context, the extraction of individual entities alone proves inadequate for conveying the scientific significance embedded within the literature. Instead, complete tuples comprising various types of entities must be extracted.
Figure 2. Common patterns of multiple entities and multiple relations in MPEAs. We use three simple sentences to exemplify three common repetition patterns. However, in real-world scenarios and our dataset, the input text comprises multiple sentences. (A) An example of multiple properties of the same material; (B) An example of multiple property values for the same material under different conditions; (C) An example of multiple property values of the same property across different materials. MPEAs: Multi-principal element alloys.
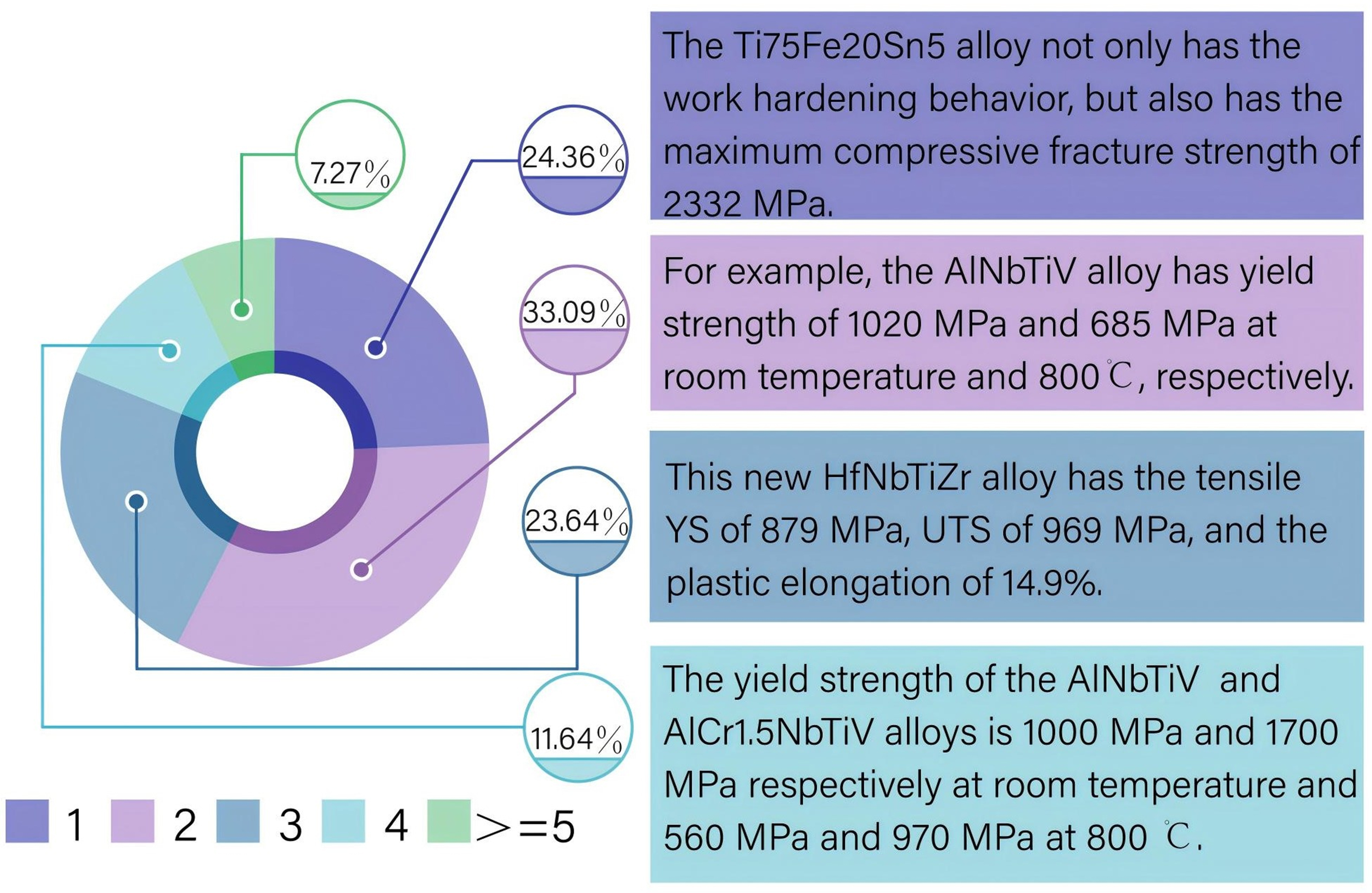
Moreover, the presence of multiple tuples conveying scientific meaning within a single sentence is prevalent in the materials science literature. Our statistical analysis, based on over 100 publications on MPEAs, reveals that sentences describing alloy properties containing only a single tuple account for merely 24.36%. The distribution of sentences containing varying numbers of tuples is illustrated in Figure 3. Nevertheless, there remains a paucity of studies proposing effective and sophisticated solutions to address this issue.
Figure 3. The proportion of sentences containing different numbers of tuples. The numbers below the pie chart indicate the number of tuples represented by each colored segment. On the right side of the pie chart are examples of varying numbers of tuples within one sentence. The proportions, in ascending order, are 24.36%, 33.09%, 23.64%, 11.64%, and 7.27%.
Dataset analyses
Following the protocol detailed in the Methods, the number of sentences and tuples in the test sets and the entire dataset are presented in Tables 1 and 2 (These datasets are denoted as 1, 2, 3, 4, and Random in the tables, and this notation is consistent throughout the subsequent sections).
Number of sentences and tuples in the test sets
| Test dataset | 1 | 2 | 3 | 4 | Random |
| Num of sentences | 40 | 38 | 38 | 22 | 23 |
| Num of tuples | 40 | 76 | 114 | 88 | 50 |
Number of sentences and tuples in the entire dataset
| Dataset | 1 | 2 | 3 | 4 | Total |
| Num of sentences | 67 | 91 | 65 | 32 | 255 |
| Num of tuples | 67 | 182 | 195 | 128 | 568 |
Published materials information extraction datasets differ markedly in domain coverage, task objectives, and annotation granularity, which complicates size-based comparisons. For instance, the superalloy pipeline reports 680 text-extracted instances of γ′ solvus temperature. MSMENTIONS includes 595 synthesis procedures, and SOFCExp covers 45 articles. These datasets target distinct units of annotation, ranging from named entities and pairwise relations to processes and events. Consequently, comparing corpora based solely on relation or instance counts can obscure differences in scope and semantic completeness.
Our contribution is designed at the tuple level with scenario-centric annotation. The 568 tuples in our corpus are drawn from multi-event sentences that explicitly bind at least Material, Property, and Property Value, and frequently consolidate multiple properties for the same material within a single coherent statement. This schema emphasizes semantic fidelity across entities and values within the sentence context, which is critical for downstream materials reasoning.
Model construction rationale
As mentioned earlier, we employed the MatSciBERT model and a pointer network[28,30] for entity extraction from input texts. Our choice of MatSciBERT over general-purpose BERT and SciBERT is grounded in both theoretical considerations and experimental validation. MatSciBERT is obtained by further pre-training SciBERT on approximately 285 million words of materials science literature, with about one-fifth focused on alloy systems. This domain-adaptive pre-training narrows the gap between the pre-training data and downstream tasks, enriching the model’s subword inventory and contextual statistics for materials terminology, synthesis routes, characterization protocols, and property-reporting conventions. Prior evaluations have shown gains over SciBERT and BERT-base on materials-focused NER, relation classification, and abstract classification, indicating that MatSciBERT’s representations capture symbol-heavy expressions and domain-specific collocations more faithfully than general-purpose models. More importantly, MatSciBERT is pre-trained on full-length text sequences, enabling it to handle longer sequences and cross-sentence dependencies. A systematic comparison among the three models is discussed later. The pointer network extracts entities by directly predicting boundary positions from encoder outputs, which reduces the search space compared with sequence tagging and is efficient on well-formatted scientific text. It also accommodates local nesting observed in materials discourse. For instance, “room temperature” contains a CONDITION span “temperature” and a CONDITION VALUE span “room temperature”. Independent heads per type allow recovery of both spans without enforcing mutual exclusivity across tags.
To tackle the challenge of multi-tuple extraction, we propose the task of entity allocation to assign extracted entities of different types to complete tuples and avoid allocation errors caused by entity relationship confusion. We believe that the key to this task is enabling the model to learn both correct and incorrect tuple matching patterns contrastively. Hence, in the second stage, we construct an entity matching score matrix to assess the likelihood of various entities being allocated together, using inter-entity and intra-entity attention to provide representations informed by both the candidate pair and the surrounding entities of the same type. Specifically, inter-entity attention yields representations between different types, and intra-entity attention captures relationships within the same type. This design allows the model to utilize information from the two entities being matched and from all entities of the corresponding types when performing tuple matching. This step is crucial for determining whether the output tuples accurately convey the intended scientific meaning, as any erroneous entity allocation by the model could distort the semantic content of the tuples. It is important to note that we do not directly predict the relationships between captured entities; rather, we determine whether two entities should reside within the same tuple. This approach is justified for two reasons: first, the number of relationships between entities in materials literature is fixed and limited once the types of entities to be extracted are identified, rendering relationship prediction less beneficial for extracting correct tuples. Second, predicting relationships between entities constitutes a multi-class classification task, whereas entity matching is a binary classification task, making the latter less prone to errors. In this stage, the five types of entities mentioned previously are extracted for subsequent assignment to the correct tuples.
Performances of entity extraction
We evaluate entity extraction across five types: MATERIAL, PROPERTY, PROPERTY VALUE, CONDITION, and CONDITION VALUE. Metrics are computed per entity type, reporting precision (P), recall (R), and F1 independently so that performance on rarer categories is visible rather than averaged out. We also report total metrics per dataset, where the denominators of total precision and total recall equal the full count of all entities in the set. This yields an overall score that reflects extraction effectiveness under the empirical frequency distribution. Results are shown in Table 3. The metrics are calculated in the following manner:
Entity extraction performance of the proposed MatSciBERT-based model
| Entity | 1 | 2 | 3 | 4 | Random | ||||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| MAT | 0.96 | 0.95 | 0.97 | 0.97 | 0.94 | 1 | 0.94 | 0.94 | 0.94 | 0.95 | 0.94 | 0.97 | 0.95 | 1 | 0.91 |
| PRO | 1 | 1 | 1 | 0.96 | 0.93 | 1 | 0.93 | 0.89 | 0.97 | 0.91 | 0.93 | 0.89 | 0.95 | 0.95 | 0.95 |
| PRO V | 0.99 | 1 | 0.98 | 1 | 1 | 1 | 0.94 | 0.90 | 0.99 | 1 | 1 | 1 | 0.97 | 0.94 | 1 |
| CON | 1 | 1 | 1 | 1 | 1 | 1 | 0.86 | 0.75 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| CON V | 0.93 | 1 | 0.88 | 1 | 1 | 1 | 0.91 | 0.83 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| TOTAL | 0.98 | 0.97 | 0.98 | 0.96 | 0.98 | 0.95 | 0.92 | 0.88 | 0.97 | 0.97 | 0.95 | 0.99 | 0.98 | 0.99 | 0.97 |
As shown in Table 3, F1 scores for all five entity types exceed 0.9, except for CONDITION on dataset 3, thereby establishing a robust foundation for the subsequent entity allocation task. We highlight the maximum F1 scores achieved for each entity type within the respective datasets. Notably, these peak F1 scores are attained by PROPERTY VALUE, CONDITION, and CONDITION VALUE. However, the F1 scores for the other two entity types exhibit greater fluctuation across different datasets. The minimum F1 scores for CONDITION and CONDITION VALUE are 0.86 and 0.91, respectively, both lower than the minima for the other entity types. This sensitivity arises from their lower frequencies: omissions in the text reduce the number of gold instances, so a few errors can noticeably affect the F1 score. Among the three more common entity types, the model demonstrates the best performance on PROPERTY VALUE, reflecting the regularity of numeric forms with units. MATERIAL remains stable across all test sets, with F1 around 0.96. PROPERTY declines as tuple count increases, from 1.00 to 0.91, which is consistent with the greater heterogeneity of PROPERTY mentions within multi-tuple sentences. As shown in Table 3, the total metric can be lower than the F1 scores for common types. For example, in dataset 3, the total F1 is lower than the F1 scores for the first three entity types, indicating that CONDITION or CONDITION VALUE reduces the overall extraction performance. Reporting both per-type and overall metrics thus captures the effect of imbalance and the contribution of rare categories to the aggregate performance.
Replacement experiments isolate the encoder choice by swapping MatSciBERT, SciBERT, and BERT-base while keeping all other components and hyperparameters fixed. All three models share the same 12-layer, 768-hidden, 12-head architecture, so observed differences can be attributed to pre-training. Tables 4 and 5 list BERT-base and SciBERT results and Table 6 summarizes totals across encoders.
Entity extraction performance of the proposed BERT-based model
| Entity | 1 | 2 | 3 | 4 | Random | ||||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| MAT | 0.74 | 0.87 | 0.65 | 0.76 | 0.81 | 0.71 | 0.55 | 0.67 | 0.47 | 0.88 | 0.90 | 0.85 | 0.68 | 0.74 | 0.63 |
| PRO | 0.82 | 0.86 | 0.76 | 0.78 | 0.87 | 0.7 | 0.82 | 0.87 | 0.77 | 0.85 | 0.90 | 0.81 | 0.80 | 0.96 | 0.68 |
| PRO V | 0.81 | 0.77 | 0.85 | 0.92 | 0.93 | 0.91 | 0.81 | 0.92 | 0.73 | 0.86 | 0.9 | 0.82 | 0.85 | 0.84 | 0.86 |
| CON | 0.8 | 1 | 0.67 | 0.5 | 1 | 0.33 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 |
| CON V | 0.47 | 0.44 | 0.5 | 0.57 | 1 | 0.4 | 0.89 | 1 | 0.8 | 0.62 | 1 | 0.44 | 0 | 0 | 0 |
| TOTAL | 0.78 | 0.81 | 0.74 | 0.82 | 0.89 | 0.75 | 0.78 | 0.86 | 0.72 | 0.85 | 0.90 | 0.80 | 0.80 | 0.84 | 0.77 |
Entity extraction performance of the proposed SciBERT-based model
| Entity | 1 | 2 | 3 | 4 | Random | ||||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| MAT | 0.93 | 0.89 | 0.98 | 0.97 | 0.98 | 0.96 | 0.87 | 0.96 | 0.94 | 0.93 | 1 | 0.88 | 0.94 | 0.97 | 0.91 |
| PRO | 0.93 | 0.91 | 0.95 | 0.88 | 0.89 | 0.88 | 0.93 | 0.96 | 0.89 | 0.89 | 0.93 | 0.96 | 0.96 | 0.97 | 0.95 |
| PRO V | 0.91 | 0.86 | 0.95 | 0.97 | 0.99 | 0.95 | 0.96 | 0.96 | 0.95 | 0.98 | 0.99 | 0.86 | 0.98 | 0.96 | 1 |
| CON | 1 | 1 | 1 | 0.5 | 1 | 0.33 | 0.8 | 1 | 0.67 | 1 | 1 | 1 | 0 | 0 | 0 |
| CON V | 0.92 | 0.88 | 0.95 | 0.94 | 0.95 | 0.93 | 0.93 | 0.95 | 0.91 | 0.95 | 0.98 | 0.93 | 0.96 | 0.95 | 0.97 |
| TOTAL | 0.92 | 0.88 | 0.95 | 0.94 | 0.95 | 0.93 | 0.93 | 0.95 | 0.91 | 0.95 | 0.98 | 0.93 | 0.96 | 0.95 | 0.97 |
Performance comparison of BERT-based model variants for entity extraction: BERT-base, SciBERT, and MatSciBERT
| Entity | 1 | 2 | 3 | 4 | Random | ||||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| BERT-base | 0.78 | 0.81 | 0.74 | 0.82 | 0.89 | 0.75 | 0.78 | 0.86 | 0.72 | 0.85 | 0.90 | 0.80 | 0.80 | 0.84 | 0.77 |
| SciBERT | 0.92 | 0.88 | 0.95 | 0.94 | 0.95 | 0.93 | 0.91 | 0.87 | 0.96 | 0.95 | 0.98 | 0.93 | 0.96 | 0.95 | 0.97 |
| MatSciBERT | 0.98 | 0.97 | 0.98 | 0.96 | 0.98 | 0.95 | 0.92 | 0.88 | 0.97 | 0.97 | 0.95 | 0.99 | 0.98 | 0.99 | 0.97 |
For MATERIAL entities, MatSciBERT attains F1 scores of 0.94-0.97, slightly ahead of SciBERT at 0.93-0.97 and substantially better than BERT-base at 0.55-0.88. For PROPERTY entities, MatSciBERT reaches 0.91-1.00, compared with 0.88-0.96 for SciBERT and 0.78-0.85 for BERT-base. For PROPERTY VALUE entities, MatSciBERT achieves 0.94-1.00, vs. 0.91-0.98 for SciBERT and 0.81-0.92 for BERT-base. For CONDITION entities, MatSciBERT maintains 0.86-1.00, while BERT-base exhibits high variance, ranging from 0.00 to 1.00. For CONDITION VALUE entities, MatSciBERT obtains 0.91-1.00, SciBERT 0.92-0.96, and BERT-base 0.00-0.89. The total performance metrics (as shown in Table 3) corroborate these findings. Overall F1 ranges from 0.92-0.98 for MatSciBERT, 0.91-0.96 for SciBERT, and 0.78-0.85 for BERT-base, with MatSciBERT remaining stable across single-tuple through multi-tuple, and random test sets. SciBERT is steady but consistently behind, while BERT-base degrades in multi-tuple settings, sometimes dropping to 0.78.
These patterns align with expectations for domain-adaptive pre-training in symbol-heavy technical text. PROPERTY VALUE benefits from surface regularity and unit markers. MATERIAL is supported by repeated alloy formula patterns and consistent naming. PROPERTY varies more in phrasing as the tuple count increases, which explains the mild decline. Rare categories amplify noise in totals; presenting disaggregated and aggregated metrics together gives a more accurate account of extraction quality under realistic class imbalance.
Performance of entity allocation
The innovative aspect of the entity allocation task presented in this paper lies in its direct prediction of which entities are assigned to the same tuple, thereby circumventing the need to predict relationships between entities. This approach not only reduces potential errors but also simplifies the process. It is evident that the complexity of the entity allocation task varies across datasets, particularly in sentences containing a diverse number of tuples. When evaluating model performance, we argue that only complete and accurate tuples can contribute to an increase in the total correct count, irrespective of the efficacy of entity extraction. Therefore, the metrics we employ to evaluate entity allocation performance are calculated in a manner similar to those used for entity extraction.
Table 7 shows that entity allocation separates the three encoders more sharply than extraction. MatSciBERT yields F1 scores between 0.75 and 0.96 across tuple complexities, SciBERT ranges from 0.68 to 0.93, and BERT-base from 0.32 to 0.78. The widening performance gaps in the allocation task underscore the benefit of materials-specific pre-training when inference depends on multiple interacting entities.
Performance of the proposed and baseline models
| Entity | 1 | 2 | 3 | 4 | Random | ||||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| Claude 3 | 0.93 | 0.93 | 0.93 | 0.51 | 0.52 | 0.50 | 0.52 | 0.50 | 0.55 | 0.62 | 0.86 | 0.48 | 0.67 | 0.76 | 0.61 |
| GPT-4o | 0.88 | 0.88 | 0.88 | 0.51 | 0.54 | 0.48 | 0.39 | 0.49 | 0.32 | 0.66 | 0.88 | 0.53 | 0.66 | 0.73 | 0.59 |
| Gemini 1.5 | 0.93 | 0.93 | 0.93 | 0.51 | 0.52 | 0.50 | 0.51 | 0.47 | 0.56 | 0.71 | 0.90 | 0.59 | 0.67 | 0.76 | 0.61 |
| Llama 3.1 | 0.90 | 0.90 | 0.90 | 0.50 | 0.50 | 0.50 | 0.47 | 0.43 | 0.53 | 0.62 | 0.91 | 0.47 | 0.64 | 0.71 | 0.57 |
| BERT base | 0.78 | 0.81 | 0.74 | 0.34 | 0.37 | 0.32 | 0.32 | 0.41 | 0.26 | 0.37 | 0.41 | 0.33 | 0.34 | 0.40 | 0.30 |
| SciBERT | 0.93 | 0.89 | 0.98 | 0.71 | 0.71 | 0.70 | 0.70 | 0.71 | 0.69 | 0.68 | 0.69 | 0.67 | 0.73 | 0.71 | 0.74 |
| Ours | 0.96 | 0.95 | 0.98 | 0.95 | 0.95 | 0.95 | 0.85 | 0.89 | 0.81 | 0.75 | 0.75 | 0.75 | 0.85 | 0.83 | 0.88 |
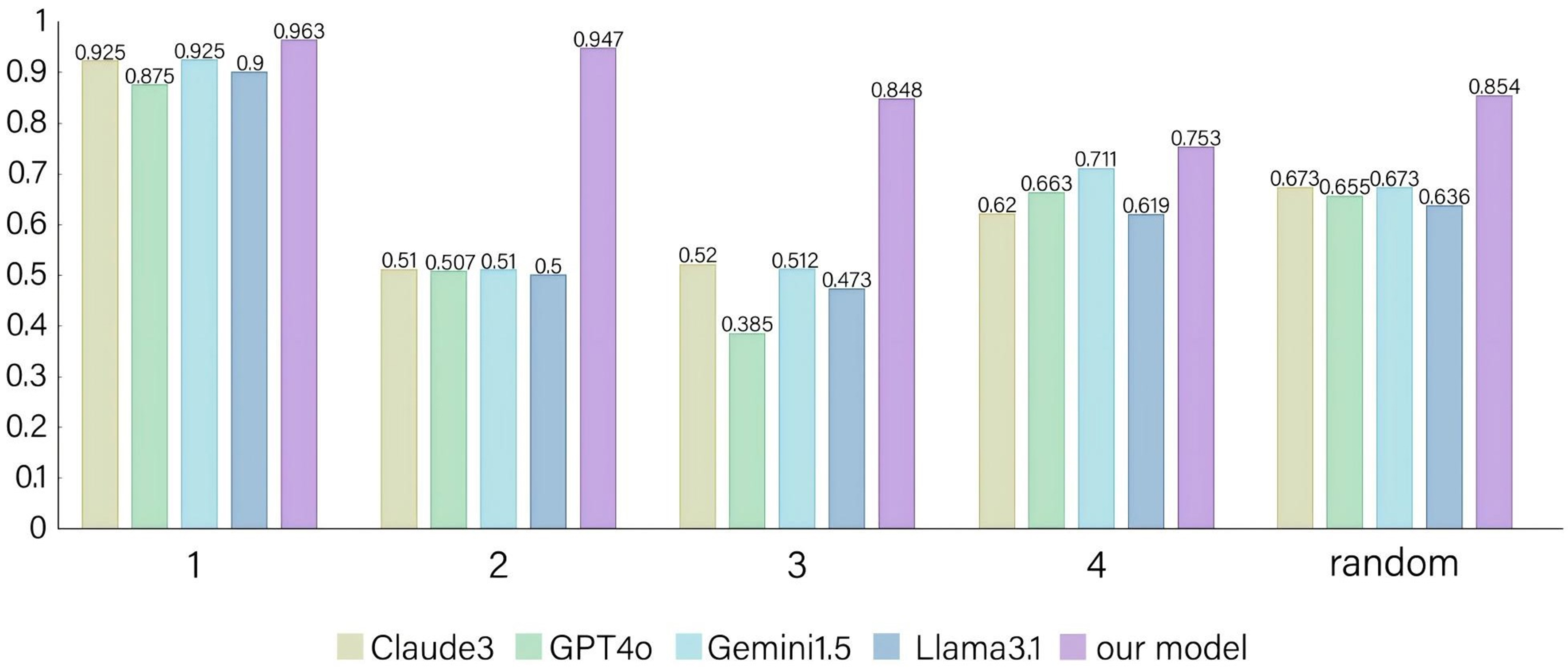
To achieve a comprehensive comparison between our proposed task-specific approach and task-agnostic LLMs, we also employ a pre-training and prompting strategy utilizing four prominent models - Claude 3 Haiku[31], GPT-4o[22], Gemini 1.5[32], and Llama 3.1 (70 B)[33] - as benchmarks. The overall results for both the proposed model and the reference models are presented in Table 7 (for brevity, these models are denoted as Claude 3, GPT-4o, Gemini 1.5, and Llama 3.1, and the highest scores are highlighted in bold for easy identification of the top-performing model), while the F1 scores for all models are illustrated in Figure 4. Our model surpasses all baseline models across all test sets in terms of F1 score. Furthermore, to mitigate the potential issue of hallucination associated with LLMs to some extent, we restrict the entity “CONDITION” to temperature within the prompt. The prompt is defined as “Extract as many tuples of the form [“MATERIAL”, “PROPERTY”, “PROPERTY VALUE”, “TEMPERATURE”, “TEMPERATURE VALUE”] from the input sentences as possible”.
Figure 4. F1 scores of the proposed and baseline models. Four baseline models are employed, all of which are strong LLMs. Among all models, the proposed model achieves the best performance. LLMs: Large language models.
Overall, the performance of the proposed model decreases as the number of tuples contained within a single sentence (denoted as k) increases. Specifically, when there is only one tuple (k = 1), the model is exclusively tasked with entity extraction, and it achieves its optimal performance with an F1 score of 0.963. At k = 2, the model’s precision and recall are equal, as the number of tuples generated by the model matches that of the gold standard labels. Additionally, the number of extracted entities aligns with the true labels. Therefore, the results for k = 1 and k = 2 both underscore the efficacy of the MatSciBERT + Pointer Network approach. The slight decline in F1 to 0.947 at k = 2 suggests the emergence of errors due to confusion during entity allocation.
Moreover, with each increment of k by 1 starting from k = 2, the F1 score decreases by approximately 0.1. Notably, at k = 4, the model’s performance deteriorates significantly to an F1 of 0.753. This decline is partly attributable to the fact that each sentence in the dataset contains, on average, four tuples, resulting in more complex sentence structures that are challenging for the model. Furthermore, as the content volume increases, authors are prone to employ omissions and abbreviations that hinder the model’s learning and prediction capabilities. Additionally, the limited number of sentences containing four tuples and variability in writing styles may further complicate model training.
In the last dataset (with an average k of 2.17), the F1 score is 0.854, slightly higher than that for k = 3 (∆ = +0.006), yet markedly lower than that for k = 2 (∆ = -0.093). Given that a prediction is considered correct only when all five entities are accurately extracted and allocated, an F1 score of 0.854 is regarded as a strong result.
The observed decline as tuple density increases reflects the growing difficulty of maintaining correct matches as more tuples appear within a sentence, compounded by syntactic complexity and semantic overlap. Authors also tend to use abbreviations or omit contextual information in multituple settings, reducing the available signal. Annotation sparsity for certain entity types and upstream extraction errors further reduce endtoend tuple accuracy. Detailed analyses follow. Across encoders, MatSciBERT remains the most stable as tuple density increases, SciBERT performs consistently but at a slightly lower level, and BERTbase degrades in multituple settings. These results are consistent with those observed in the extraction stage and support the view that domain-adaptive pre-training mitigates distributional mismatch in symbol-heavy, convention-rich materials science text, thereby benefiting entity allocation.
On the other hand, the performance of LLMs remains relatively consistent across datasets. Notably, Gemini 1.5 demonstrates superior efficacy, slightly trailing Claude 3 on the third dataset (∆ = -0.008) while significantly outpacing it on the fourth dataset (∆ = +0.091). For the remaining datasets, both models achieve identical F1 scores. At k = 1, F1 scores for the large models consistently exceed 0.9, indicating their proficiency in the entity extraction task. However, in contrast to the proposed model, LLMs exhibit diminished performance on the second and third datasets (average F1 = 0.507 for k = 2 and average F1 = 0.473 for k = 3). Although their performance improves on the fourth dataset (average F1 = 0.653 for k = 4), it remains inferior to that of the proposed model.
Our analysis suggests that large models are prone to extract excessive irrelevant content as entities, often misidentifying the concentration of a chemical element within an alloy or fabrication methods as CONDITION entities. Additionally, these models tend to generate entities containing extraneous information rather than extracting precise entities based on the input sentence, resulting in reduced performance. Conversely, on the fourth test set, where sentences focus more on tuple-related content with minimal extraneous information, the precision of LLMs improves significantly (average P = 0.89 for k = 4), even surpassing that of the proposed model (P = 0.75 for k = 4). However, their recall remains lower and the overall F1 score still lags behind.
Ablation studies
Since our work uses several novel components, especially attention mechanisms, for the task of entity allocation, we conducted a series of ablation experiments to verify the importance of these components for structural information extraction in materials science literature. In particular, after removing selected components from the model, we retrain it and evaluate its performance on the test datasets. The examined components include entity allocation, intra-entity attention, and inter-entity attention (These settings are denoted as without allocation, without intra, and without inter in Table 8). We start the ablation experiments from dataset 2, as dataset 1 does not involve entity allocation. The overall results are shown in Table 8. The best F1, precision (P), and recall (R) values for each test set are highlighted in bold.
Results of the ablation study
| Dataset | Our model | Without allocation | Without intra | Without inter | ||||||||
| F1 | P | R | F1 | P | R | F1 | P | R | F1 | P | R | |
| 2 | 0.95 | 0.95 | 0.95 | 0.56 | 0.40 | 0.95 | 0.87 | 0.87 | 0.87 | 0.86 | 0.86 | 0.86 |
| 3 | 0.85 | 0.89 | 0.81 | 0.45 | 0.31 | 0.83 | 0.81 | 0.85 | 0.77 | 0.79 | 0.84 | 0.75 |
| 4 | 0.75 | 0.75 | 0.75 | 0.34 | 0.21 | 0.85 | 0.73 | 0.73 | 0.73 | 0.69 | 0.69 | 0.69 |
| Random | 0.85 | 0.83 | 0.88 | 0.56 | 0.41 | 0.90 | 0.84 | 0.81 | 0.86 | 0.82 | 0.79 | 0.84 |
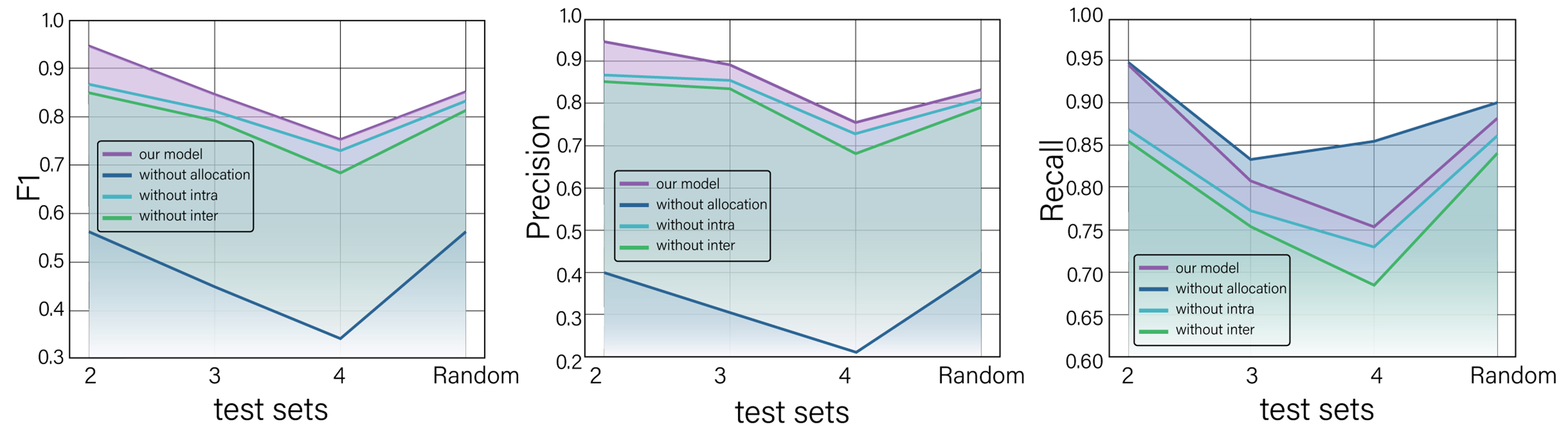
To enhance interpretability, the results are visualized in Figure 5. Overall, the proposed model with all components achieves the best F1 scores across all datasets, indicating that entity allocation, intra-entity attention, and inter-entity attention all contribute positively to model performance. Our ablation results show that removing any of these three components degrades semantic accuracy in distinct ways, with direct implications for user trust.
Figure 5. F1 scores of the proposed complete model and its ablated variants. The proposed model achieves the best performance among all configurations.
The model without entity allocation consistently shows the lowest F1 and precision scores, but the highest recall. The F1 score drops to 0.56 on the random dataset and as low as 0.34 for datasets with higher tuple density, with precision decreasing substantially while recall remains unchanged. This pattern arises because the model without entity allocation generates all possible combinations of entities as output tuples. As a result, the model performs poorly in balancing precision and recall, resulting in the lowest F1 scores. Moreover, removing entity allocation disrupts cross-mention coherence when multiple materials appear in the same passage. The model then mixes entities across materials, yielding tuples that pair a property value from one alloy with the name of another. Such cross-assignment produces scientifically invalid combinations that would mislead downstream analyses. This underscores the significance of entity allocation in our work.
When entity allocation is retained, the F1 scores (0.86, 0.79, 0.69, and 0.82, respectively) of the model without inter-attention are consistently lower than those (0.868, 0.81, 0.73, and 0.84, respectively) of the model without intra-entity attention, proving that inter-entity attention is more important than intra-entity attention for extracting structured information. This suggests that correctly modeling cross-type relationships is more important than capturing relationships within entities of the same type for accurate tuple extraction. Specifically, removing inter-entity attention disrupts semantic alignment across entities, increasing mismatches between properties and their appropriate units or scales, such as matching “yield strength” with “Hv” units instead of “MPa”. This mechanism helps maintain value-unit associations and handle modifiers within entity boundaries, creating a false sense of correctness. In contrast, removing intra-entity attention weakens internal consistency within the same entity. The resulting outputs may appear syntactically valid but violate domain constraints. Without it, the model frequently outputs incomplete or fragmented value-unit pairs, which distorts quantitative meaning in materials design tasks.
Regardless of the model settings, the F1 score decreases as the number of tuples to be extracted in the test set increases. Nevertheless, the complete model retains the highest scores at each level of complexity. Notably, the complete model also achieves the highest recall (0.95) on dataset 2, indicating that it successfully identifies all correct tuples during the entity allocation phase without introducing incorrect ones.
Overall, the performance drops observed in the ablation study directly translate into an increased likelihood of scientifically invalid extractions. Without these components, users are more likely to encounter cases where extracted tuples appear syntactically correct but violate domain knowledge, requiring additional verification steps and ultimately reducing trust in automated materials data extraction for practical applications.
Error analyses
To pinpoint the factors behind the observed decline as tuple density increases, we performed fine-grained error analyses across all datasets and examined mispredictions. Four error classes account for most of the degradation, often interacting within the same sentence. Table 9 presents a quantitative breakdown of error causes across datasets.
Proportion of error causes for each dataset
| Dataset | 2 | 3 | 4 |
| Syntactic complexity | 20% | 10.5% | 28.6% |
| Annotation sparsity | 0 | 10.5% | 9.5% |
| Semantic overlap | 40% | 10.5% | 47.6% |
| Entity extraction error | 40% | 68.4% | 14.3% |
Syntactic complexity is a recurrent source of error. Sentences with layered grammatical structures, embedded clauses, long-distance dependencies, or unusual word order increase ambiguity in boundary detection and argument assignment. For example, in the sentence “Among the developed Ti-xCr-3Sn alloys, Ti-6Cr-3Sn alloy showed maximum ductility (41% fracture strain), maximum (58%) recovery ratio due to shape memory effect and maximum pseudoelastic response due to stress induced martensitic transformation and twinning deformation mode”, the model incorrectly extracted “Ti-xCr-3Sn” as a specific alloy name rather than recognizing it as a compositional family, which in turn yields an incorrect tuple. As tuples accumulate within a sentence, the same syntactic devices that improve narrative flow introduce competing local cues, increasing the incidence of such errors.
Annotation sparsity constitutes a second, more prosaic limitation. Training examples featuring multituple configurations and rare entity types are less common, reducing the model’s exposure to the very patterns that dominate highdensity sentences. This limits the learned decision boundaries for complex extraction cases and reduces generalization to unseen yet systematic configurations.
Semantic overlap represents the most significant challenge when sentences contain several tuples. Shared lexical items, repeated subjects, or reused contextual frames can lead to conflicts in boundary assignment and entity pairing. A typical pattern involves multiple properties reported for a single material under distinct conditions, where the model may struggle to correctly associate each property-value pair with its corresponding experimental condition.
Entity extraction errors constitute the fourth class and act as upstream error amplifiers. For instance, in the sentence “The calculated elastic constants C11, elastic constants C12 and elastic constants C44 for the Ti20Zr20Hf20Nb20Ta20 single crystalline disordered alloy yielded 160.2, 124.4 and 62.4 GPa, respectively”, if “elastic constants” is misrecognized due to its rarity in the training data, all three resulting tuples will contain incorrect property names.
For dataset 2, semantic overlap and entity extraction each account for 40% of errors, with syntactic complexity contributing 20%. In dataset 3, entity extraction dominates at 68.4%, while the remaining categories each contribute 10.5%. In dataset 4, semantic overlap is the primary source of error at 47.6%, followed by syntactic complexity at 28.6%. Although entity extraction errors constitute the largest overall proportion, no clear linear relationship is observed between error type distribution and tuple density. Importantly, these categories often cooccur in dense sentences, where semantic overlap and syntactic complexity combine to create boundary ambiguities that are difficult even for human annotators. When focusing specifically on the entity allocation task (excluding upstream entity extraction errors), semantic overlap and syntactic complexity emerge as the principal bottlenecks, particularly in dataset 4. This observation aligns with the intuition that allocation is most sensitive to reused constituents and partial token reuse across tuples, both of which increase with tuple density. In addition, we analyze potential concentrated vulnerabilities across entity-type combinations and specific sentence structures, with detailed discussion provided in the Supplementary Materials. Although no such vulnerability is observed for entity-type combinations, dataset 4 shows a higher error rate associated with increased structural diversity, with errors concentrated in the most complex template featuring parallel quantification and cross-cutting condition mentions [Supplementary Figure 1 and Supplementary Table 1].
By examining MPEAs as a case study, our work distinguishes itself from traditional approaches in two key aspects. First, we prioritize the accuracy of the structured information extracted by the model, rather than focusing solely on entity extraction performance. Second, instead of using the conventional named entity recognition and relation extraction (NERRE) framework, we employ an integrated methodology that combines entity extraction and matching to generate structured information. Isolated entities often lack the capacity to convey complete scientific meaning; therefore, accurate multidimensional tuples are essential for representing structured information in the literature. Additionally, we note that it is sufficient to identify which entities belong to the same tuple without explicitly predicting the relationships between individual entities. When a sentence contains multiple tuples, our integrated entity extraction and matching approach proves to be more precise and efficient compared to the NERRE method, which is more susceptible to errors and incurs unnecessary computational overhead.
LLMs employ a pre-training-plus-prompting approach, where users construct prompts and provide inputs to guide the model in performing specific tasks. In contrast, our methodology adopts a pre-training-plus-fine-tuning framework, which involves the deliberate design of deep learning networks and systematic data annotation. This approach not only enhances model capability but also significantly improves performance in extracting multiple tuples, as demonstrated in this study. Although LLMs offer a more user-friendly interface, their outputs are influenced by commercial access limits and weak control over error modes. Another notable drawback of LLMs is the phenomenon of “hallucination”[34,35], in which models generate plausible but misleading information not grounded in factual data, such as misinterpreting a specific element within an alloy as a CONDITION entity. Although some of these outputs may seem reasonable, they ultimately compromise the accuracy required for successful entity extraction. Our approach, on the other hand, as a specialized method designed for this task, offers greater precision compared to the broader but less accurate capabilities of LLMs and can be easily adapted to other materials domains given adequate high-quality data for fine-tuning. Nonetheless, this does not imply that LLMs are inherently inadequate. With appropriate fine-tuning, their extraction performance can improve, and they remain strong in broader tasks such as language understanding, coding, and reasoning.
Our approach has associated costs. Fine-tuning requires high-quality annotations at the tuple level, typically on the order of hundreds of verified tuples per domain to stabilize allocation performance. Data scarcity becomes more pronounced as tuple density increases, where abbreviations, omissions, and overlapping semantics raise ambiguity. We acknowledge the practical constraint that generating high-quality labels from raw scientific texts requires substantial time and domain expertise, which can hinder rapid scalability. Looking ahead, we plan to incrementally expand the corpus in future work while reducing reliance on extensive manual annotation through transfer learning and data augmentation. A separate study in our recent work[36] preliminarily shows that transfer across related materials tasks improves performance under data scarcity. Active learning could further optimize annotation efforts by prioritizing samples where the model exhibits high uncertainty, particularly when entity allocation confidence scores fall below predefined thresholds. This ensures that limited resources are focused on the most informative examples for model improvement.
The current work focuses on narrative text. We observe that some data are presented in tables and figures, especially in comparative studies across materials. However, in most cases, authors also describe the semantic content of figures in the surrounding text, including the variables compared, conditions, and principal findings. Hence, for the purposes of the present work, text-based extraction is sufficient for most applications. Nevertheless, there remain cases in which key values exist only in plots or highly structured tables. Addressing these scenarios requires computer vision techniques for chart and plot understanding, including axis calibration, legend and marker parsing, and mapping graphical encodings to numerical series with associated metadata such as units, conditions, and uncertainty. A rigorous treatment demands careful handling of multi-panel layouts, non-linear axes, overplotting, and figure-specific conventions in materials science journals.
CONCLUSIONS
In conclusion, to capture structured information in materials science literature, particularly within complex corpora containing multiple scientific tuples, we propose an approach that combines a MatSciBERT-based pointer extractor with an allocation module that enforces coherence through inter- and intra-entity attention. Applied to the mechanical properties of MPEAs, exact match evaluation yields F1 scores of 0.96, 0.95, 0.85, and 0.75 for sentences containing one to four tuples, respectively, and 0.85 on a randomly curated set, confirming strong and consistent performance as tuple complexity increases. Ablation studies indicate that the allocation module is the key driver of precision, with inter-entity attention contributing more than intra-entity attention to tuple correctness. Substituting SciBERT or BERT for MatSciBERT degrades performance, with the gap widening as tuple density increases. Prompted large models exhibit inconsistent tuple assembly, leading to lower exact match reliability in structurally complex sentences. These findings establish the proposed method as a simpler, more effective, and accurate alternative to LLMs in specialized tasks where precise multi-tuple alignment and scientific correctness are required for downstream modeling. Future work will emphasize data efficiency and portability through transfer learning and active learning.
DECLARATIONS
Authors’ contributions
Data curation, model construction, and writing - original draft: Hei, M.
Analysis, methodology, and writing - review and editing: Zhang, Z.
Supervision: Liu, Q.; Zhao, X.; Ye, Y.
Data curation and visualization: Pan, Y.; Peng, Y.
Supervision and methodology: Zhang, X.
Supervision and funding acquisition: Bai, S.
All authors have reviewed and approved the manuscript.
Availability of data and materials
The JSON-formatted dataset and all code used in this work are available at https://github.com/HEI-MENGZHE/Material_Tuple.
AI and AI-assisted tools statement
During the preparation of this manuscript, the AI tool DeepSeek (version V3.2, released 2025-09-06) was used solely for language editing. The tool did not influence the study design, data collection, analysis, interpretation, or the scientific content of the work. All authors take full responsibility for the accuracy, integrity, and final content of the manuscript.
Financial support and sponsorship
This work was supported by the National Natural Science Foundation of China (Grant Nos. 62102431 and U20A20231).
Conflicts of interest
All authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
Supplementary Materials
REFERENCES
1. Gormley, A. J.; bookb, M. A. Machine learning in combinatorial polymer chemistry. Nat. Rev. Mater. 2021, 6, 642-4.
2. Wang, S.; Yue, H.; Yuan, X. Accelerating polymer discovery with uncertainty-guided PGCNN: explainable AI for predicting properties and mechanistic insights. J. Chem. Inf. Model. 2024, 64, 5500-9.
3. Carvalho, R. P.; Marchiori, C. F. N.; Brandell, D.; Araujo, C. M. Artificial intelligence driven in-silico discovery of novel organic lithium-ion battery cathodes. Energy. Storage. Mater. 2022, 44, 313-25.
4. Bhowmik, A.; Berecibar, M.; Casas‐Cabanas, M.; et al. Implications of the BATTERY 2030+ AI‐assisted toolkit on future low‐TRL battery discoveries and chemistries. Adv. Energy. Mater. 2021, 12, 2102698.
5. Zhou, Z.; Shang, Y.; Liu, X.; Yang, Y. A generative deep learning framework for inverse design of compositionally complex bulk metallic glasses. npj. Comput. Mater. 2023, 9, 15.
6. Basu, B.; Gowtham, N. H.; Xiao, Y.; Kalidindi, S. R.; Leong, K. W. Biomaterialomics: data science-driven pathways to develop fourth-generation biomaterials. Acta. Biomater. 2022, 143, 1-25.
7. Singh, A. V.; Rosenkranz, D.; Ansari, M. H. D.; et al. Artificial intelligence and machine learning empower advanced biomedical material design to toxicity prediction. Adv. Intell. Syst. 2020, 2, 2000084.
8. Debnath, A.; Krajewski, A. M.; Sun, H.; et al. Generative deep learning as a tool for inverse design of high entropy refractory alloys. J. Mater. Inf. 2021, 1, 3.
9. Hart, G. L. W.; Mueller, T.; Toher, C.; Curtarolo, S. Machine learning for alloys. Nat. Rev. Mater. 2021, 6, 730-55.
10. Kononova, O.; He, T.; Huo, H.; Trewartha, A.; Olivetti, E. A.; Ceder, G. Opportunities and challenges of text mining in aterials research. iScience 2021, 24, 102155.
11. Sierepeklis, O.; Cole, J. M. A thermoelectric materials database auto-generated from the scientific literature using ChemDataExtractor. Sci. Data. 2022, 9, 648.
12. Kumar, P.; Kabra, S.; Cole, J. M. Auto-generating databases of Yield Strength and Grain Size using ChemDataExtractor. Sci. Data. 2022, 9, 292.
13. Wang, W.; Jiang, X.; Tian, S.; et al. Automated pipeline for superalloy data by text mining. npj. Comput. Mater. 2022, 8, 9.
14. Swain, M. C.; Cole, J. M. ChemDataExtractor: a toolkit for automated extraction of chemical information from the scientific literature. J. Chem. Inf. Model. 2016, 56, 1894-904.
15. Widiastuti, N. I. Convolution neural network for text mining and natural language processing. IOP. Conf. Ser. Mater. Sci. Eng. 2019, 662, 052010.
16. Devlin, J.; Chang, M. W.; Lee, K.; Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. In Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Minneapolis, USA, June 2, 2019; Association for Computational Linguistics; Vol. 1. pp. 4171-86.
17. Gupta, T.; Zaki, M.; Krishnan, N. M. A.; Mausam,
18. Shetty, P.; Rajan, A. C.; Kuenneth, C.; et al. A general-purpose material property data extraction pipeline from large polymer corpora using natural language processing. npj. Comput. Mater. 2023, 9, 52.
19. Lee, J.; Yoon, W.; Kim, S.; et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2020, 36, 1234-40.
20. Huang, S.; Cole, J. M. BatteryBERT: a pretrained language model for battery database enhancement. J. Chem. Inf. Model. 2022, 62, 6365-77.
21. Brown, T. B.; Mann, B.; Ryder, N.; et al. Language models are few-shot learners. arXiv 2020, arXiv:2005.14165. Available online: https://doi.org/10.48550/arXiv.2005.14165. (accessed 23 Mar 2026).
22. OpenAI, Achiam, J.; Adler, S.; et al. GPT-4 technical report. arXiv 2023, arXiv:2303.08774. Available online: https://doi.org/10.48550/arXiv.2303.08774. (accessed 23 Mar 2026).
23. Touvron, H.; Lavril, T.; Izacard, G.; et al. LLaMA: open and efficient foundation language models. arXiv 2023, arXiv:2302.13971. Available online: https://doi.org/10.48550/arXiv.2302.13971. (accessed 23 Mar 2026).
24. Chowdhery, A.; Narang, S.; Devlin, J.; et al. PaLM: scaling language modeling with pathways. arXiv 2022, arXiv:2204.02311. Available online: https://doi.org/10.48550/arXiv.2204.02311. (accessed 23 Mar 2026).
25. Team, G.; Anil, R.; Borgeaud, S.; et al. Gemini: a family of highly capable multimodal models. arXiv 2023, arXiv:2312.11805. Available online: https://doi.org/10.48550/arXiv.2312.11805. (accessed 23 Mar 2026).
26. Wei, J.; Bosma, M.; Zhao, V. Y.; et al. Finetuned language models are zero-shot learners. arXiv 2021, arXiv:2109.01652. Available online: https://doi.org/10.48550/arXiv.2109.01652. (accessed 23 Mar 2026).
27. Dagdelen, J.; Dunn, A.; Lee, S.; et al. Structured information extraction from scientific text with large language models. Nat. Commun. 2024, 15, 1418.
28. Vinyals, O.; Fortunato, M.; Jaitly, N. Pointer networks. In NIPS'15: Proceedings of the 29th International Conference on Neural Information Processing Systems, Montreal, Canada, December 7-12, 2015; MIT Press: Cambridge, Massachusetts, United States, 2015; Vol. 28. pp. 2692-700.
29. Vaswani, A.; Shazeer, N.; Parmar, N.; et al. Attention is all you need. arXiv 2017, arXiv:1706.03762. Available online: https://doi.org/10.48550/arXiv.1706.03762. (accessed 23 Mar 2026).
30. Sun, F.; Jiang, P.; Sun, H.; Pei, C.; Ou, W.; Wang, X. Multi-source pointer network for product title summarization. arXiv 2018, arXiv:1808.06885. Available online: https://doi.org/10.48550/arXiv.1808.06885. (accessed 23 Mar 2026).
31. Anthropic. The Claude 3 model family: opus, sonnet, haiku. https://www-cdn.anthropic.com/de8ba9b01c9ab7cbabf5c33b80b7bbc618857627/Model_Card_Claude_3.pdf. (accessed 2026-03-23).
32. Gemini Team Google. Gemini 1.5: unlocking multimodal understanding across millions of tokens of context. arXiv 2024, arXiv:2403.05530. Available online: https://doi.org/10.48550/arXiv.2403.05530. (accessed 23 Mar 2026).
33. Grattafiori, A.; Dubey, A.; Jauhri, A.; et al. The Llama 3 herd of models. arXiv 2024, arXiv:2407.21783 Available online: https://doi.org/10.48550/arXiv.2407.21783. (accessed 23 Mar 2026).
34. Ji, Z.; Lee, N.; Frieske, R.; et al. Survey of hallucination in natural language generation. ACM. Comput. Surv. 2023, 55, 1-38.
35. Farquhar, S.; Kossen, J.; Kuhn, L.; Gal, Y. Detecting hallucinations in large language models using semantic entropy. Nature 2024, 630, 625-30.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data


















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].