fig10
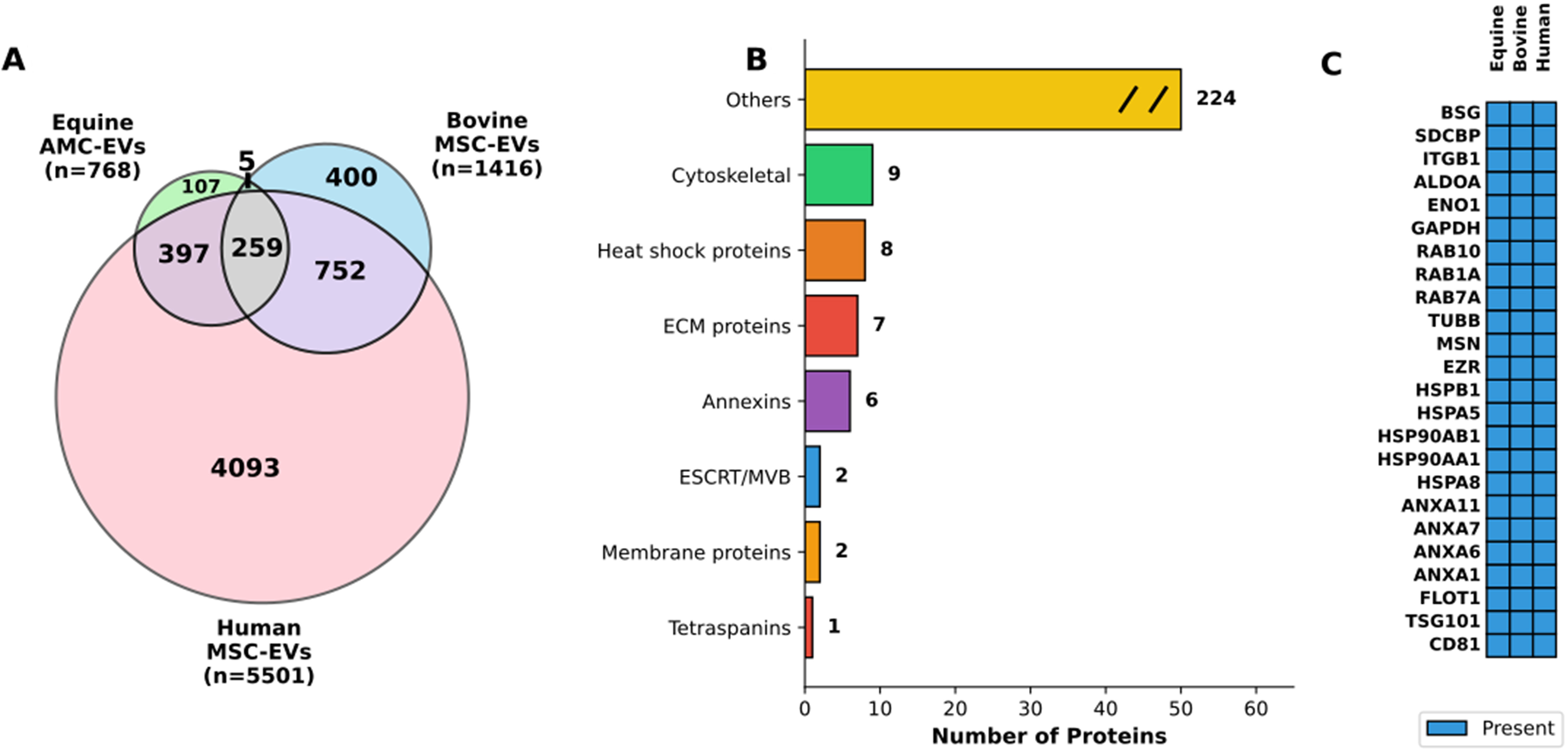
Figure 10. Cross-species comparative analysis of extracellular vesicle proteomes. (A) Venn diagram showing protein overlap across species. The diagram illustrates the distribution of proteins identified in EVs derived from equine AMCs (Equine AMC-EVs, n = 768), bovine mesenchymal stem cells (Bovine MSC-EVs, n = 1,416), and human mesenchymal stem cells (Human MSC-EVs, n = 5,501). The equine dataset comprises both proteins shared with eAMCs (n = 697) and proteins exclusive to EVs (n = 71). Numbers indicate unique and shared proteins between species, with 259 proteins conserved across all three species, representing 33.7% of the equine EV proteome; (B) Functional categorization of conserved proteins. Horizontal bar chart displaying the functional classification of the 259 proteins conserved across all three species. Categories include: Cytoskeletal proteins (n = 9), Heat shock proteins (n = 8), ECM proteins (n = 7), Annexins (n = 6), ESCRT/MVB machinery (n = 2), Membrane proteins (n = 2), Tetraspanins (n = 1), and Others (n = 224). The “Others” category is displayed with a break symbol (//) to indicate axis truncation; (C) Top conserved EV protein markers across species. Heatmap displaying 24 representative proteins conserved in EVs from all three species, organized by functional category. Blue boxes indicate presence of the protein. The panel includes: classical EV markers (CD81, TSG101, FLOT1), Annexins (ANXA1, ANXA6, ANXA7, ANXA11), heat shock proteins (HSPA8, HSPA5, HSP90AA1, HSP90AB1, HSPB1), cytoskeletal proteins (EZR, MSN, TUBB), RAB proteins (RAB7A, RAB1A, RAB10), metabolic enzymes (GAPDH, ENO1, ALDOA), and membrane proteins (ITGB1, SDCBP, BSG). EVs: Extracellular vesicles; EV: extracellular vesicle; AMCs: amniotic-derived mesenchymal stromal cells; MSC: mesenchymal stromal cell; eAMCs: equine amniotic mesenchymal cell; ECM: extracellular matrix; ESCRT: endosomal sorting complexes required for transport; MVB: multivesicular body.