fig4
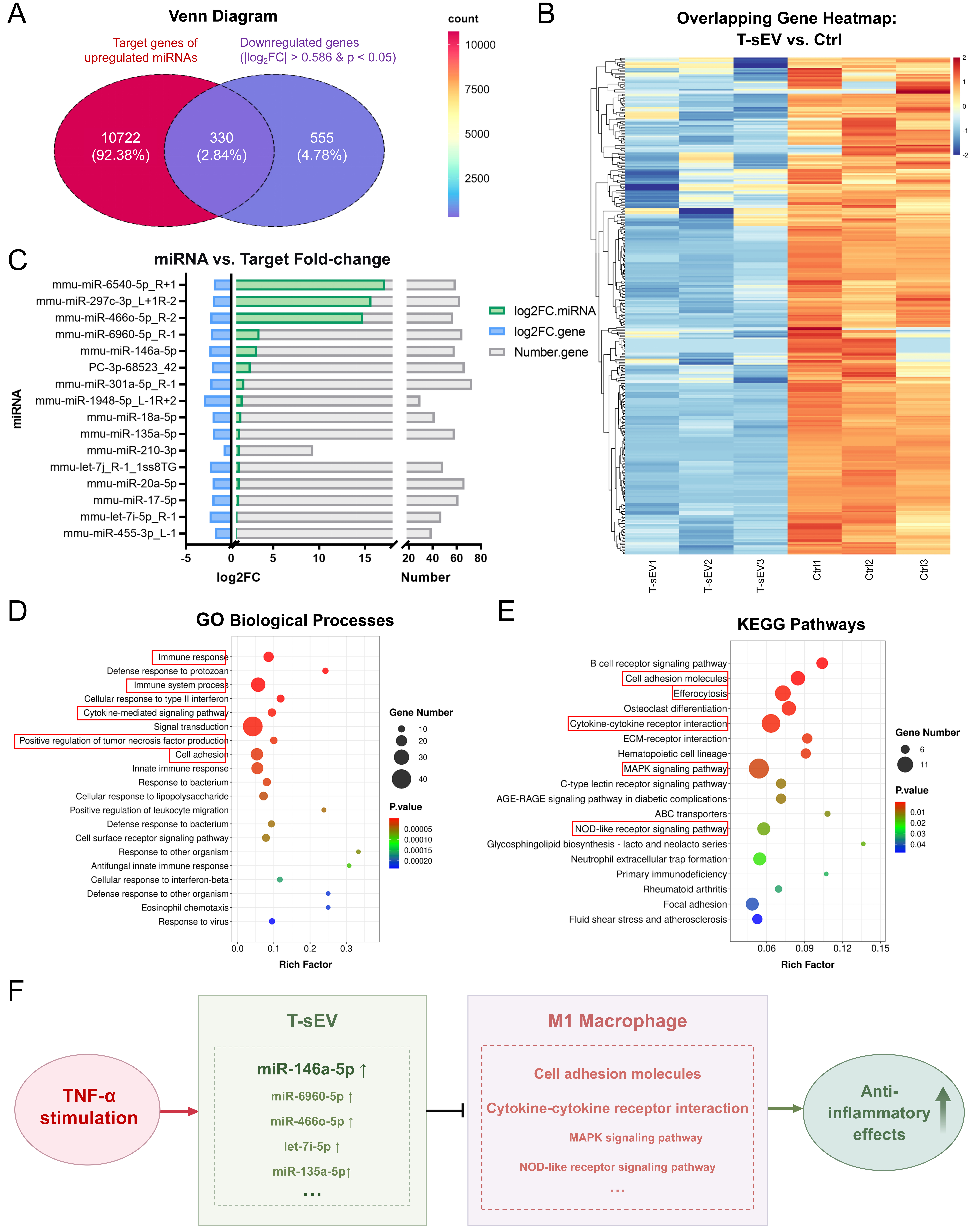
Figure 4. Integrated analysis of T-sEV miRNAs and the transcriptome of recipient macrophages. (A) Venn diagram showing the overlap between the predicted targets of T-sEV-enriched miRNAs (log2FC > 1, P < 0.05) and the experimentally identified downregulated genes in recipient macrophages (log2FC < -0.586, P < 0.05). The intersection identifies 330 high-confidence target genes; (B) Heatmap shows the expression patterns of the overlapping genes from (A) across treatment groups; (C) Bar plot showing the inverse expression patterns of selected highly upregulated miRNAs (green bars) and their corresponding, experimentally downregulated target genes (blue bars) derived from the overlapping region in (A). The gray bars represent the count of these downregulated target genes. The x-axis represents the log2FC or the gene count; (D) Bubble plot displays the top 20 enriched GO biological process terms for the overlapping genes. Red boxes indicate key terms most relevant to the study’s focus; (E) Bubble plot shows 18 selected KEGG pathways from 33 significantly enriched terms (P < 0.05), prioritized based on relevance to immunomodulation (see Methods). Red boxes indicate key pathways most relevant to the study’s focus; (F) Schematic diagram proposes a mechanism by which T-sEV-derived miRNAs regulate macrophage polarization via the key signaling pathways identified above. MSC: Mesenchymal stem cell; sEV: small extracellular vesicles; T-sEV: sEV derived from TNF-α-primed MSCs; FC: fold-change; Ctrl: control; miRNA: microRNA; GO: Gene Ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes.