fig2
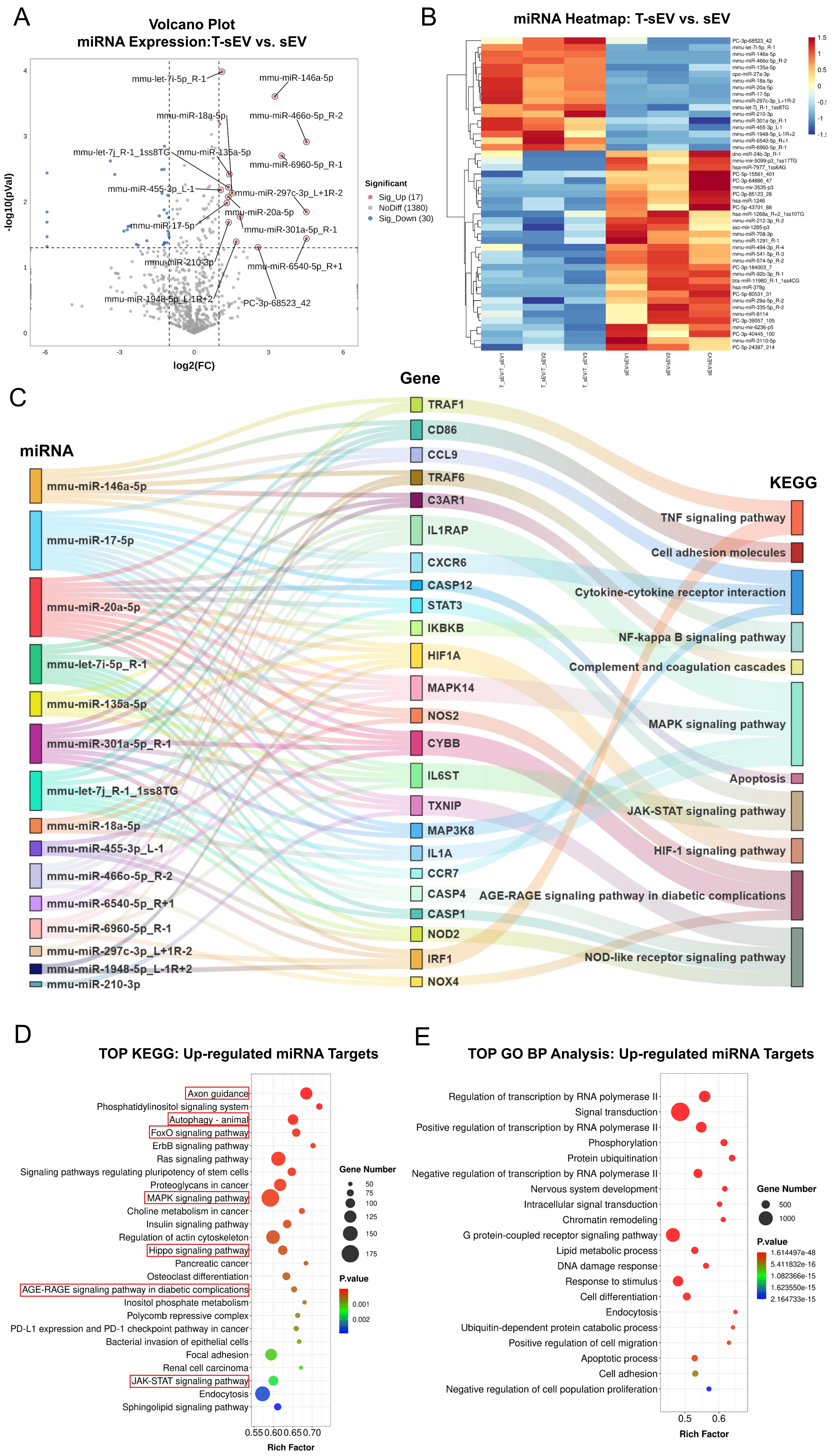
Figure 2. Functional annotation of T-sEV miRNAs through integrated bioinformatic analysis. (A) Volcano plot of differentially expressed miRNAs between T-sEV and sEV. Significantly dysregulated miRNAs were identified using a threshold of |log2FC| > 1 and P-value < 0.05. Upregulated and downregulated miRNAs are highlighted in red and blue, respectively; (B) Heatmap of significantly dysregulated miRNAs identified in (A), showing hierarchical clustering of miRNAs across samples; (C) Sankey diagram illustrating the network from key upregulated miRNAs in T-sEV (left) to their predicted target genes (center) and subsequently to the significantly enriched KEGG pathways (right); (D) Bubble plot of KEGG pathway enrichment analysis for predicted targets of upregulated T-sEV miRNAs. The top 25 enriched pathways are displayed, selected from the top 30 most significant results (P-value < 0.05) by excluding 5 pathways with overly broad functions (see Methods for criteria). The rich factor (X-axis), gene count (bubble size), and statistical significance (color) are indicated; (E) Bubble plot of the top 20 significantly enriched GO biological process terms for the predicted target genes of upregulated T-sEV miRNAs. MSC: Mesenchymal stem cell; sEV: small extracellular vesicles; T-sEV: sEV derived from TNF-α-primed MSCs; FC: fold-change; miRNA: microRNA; KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: Gene Ontology.