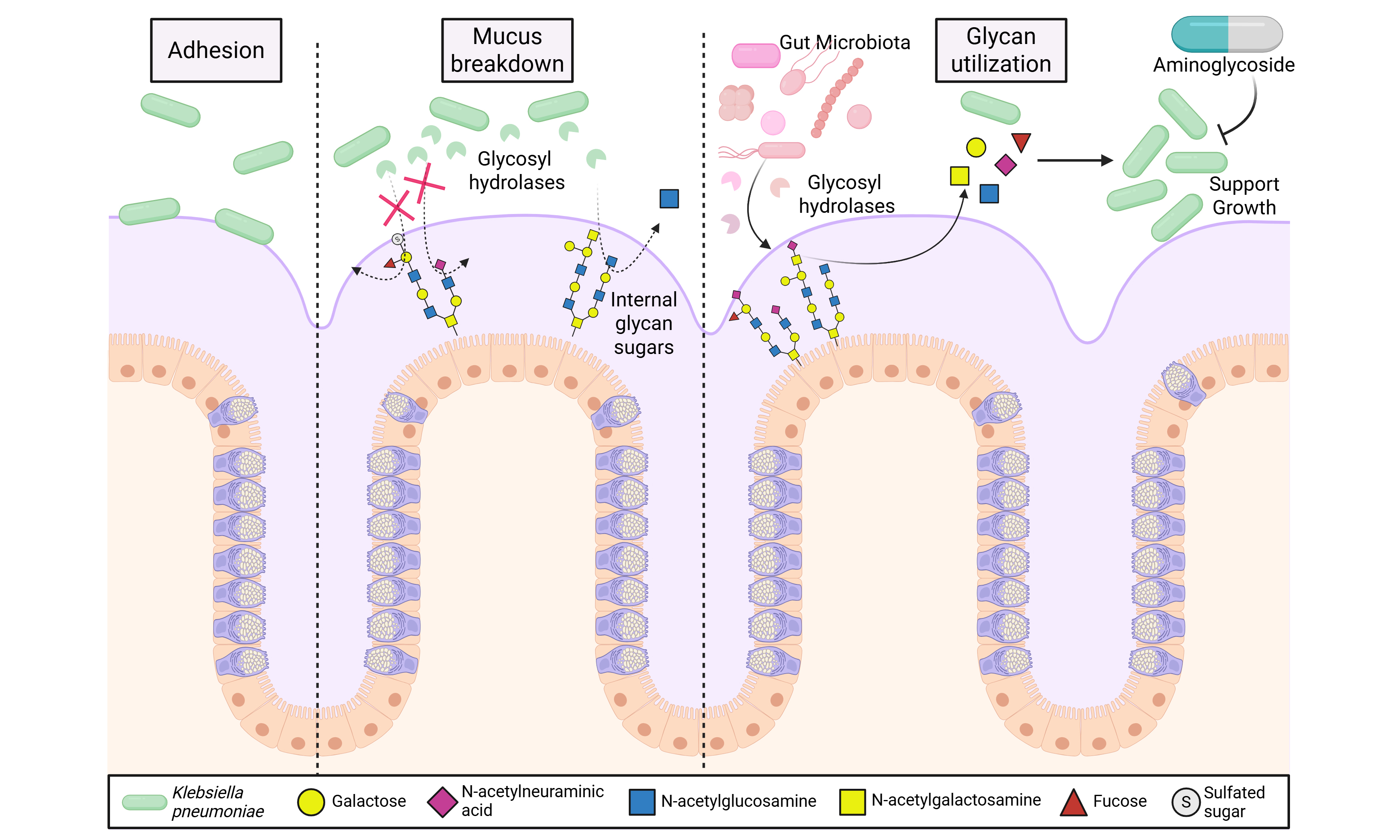
Intestinal mucus acts as a nutrient source and signal for Klebsiella pneumoniae
Abstract
Background: Klebsiella pneumoniae can colonize the gastrointestinal tract, yet its interactions with intestinal mucus remain poorly defined. In this study, we examined the capacity of Klebsiella pneumoniae (K. pneumoniae) to adhere and use intestinal mucus and its associated glycans.
Methods: Multiple commercial and clinical K. pneumoniae isolates were tested for adhesion to porcine and human Mucin 2 (MUC2) using fluorescence-based assays and microscopy. In vivo mucus localization was examined in colonized mice by fluorescent in situ hybridization (FISH). Genomic analyses of K. pneumoniae genomes were performed to identify glycosyl hydrolases and sugar utilization pathways. Growth on mucin-derived monosaccharides or intact mucus was assessed in minimal media. Biofilm formation and aminoglycoside susceptibility were measured in the presence or absence of mucus.
Results: All K. pneumoniae strains adhered robustly to porcine and human MUC2 in vitro and we found K. pneumoniae localized to the murine mucus layer in vivo. Genomic analysis of over 1,000 K. pneumoniae isolates revealed that most strains possess glycosyl hydrolases targeting internal galactose, N-acetyl-glucosamine (GlcNAc), and N-acetyl-galactosamine (GalNAc) glycan sugars, though they lack enzymes to cleave terminal fucose or N-acetyl-neuraminic acid. Consistent with this finding, we found that K. pneumoniae alone could not grow in minimal media with intact mucus as a sole carbon source. However, we found that K. pneumoniae could grow with free mucus glycan-derived sugars galactose, GlcNAc, GalNAc fucose and N-acetyl-neuraminic acid. Mucus did not alter biofilm formation, but it significantly increased sensitivity to gentamicin, kanamycin and streptomycin.
Conclusion: These findings identify mucus as an important modulator of K. pneumoniae colonization and antibiotic responsiveness.
Keywords
INTRODUCTION
Klebsiella pneumoniae (K. pneumoniae) is a Gram-negative opportunistic pathobiont that has garnered attention in recent years due to its antibiotic resistance and capacity to cause severe infections[1]. It has been implicated in a wide range of diseases, including liver abscesses, pneumonia, urinary tract infections, and bloodstream infection[2-5]. The risk of K. pneumoniae infection is highest among immunocompromised individuals, neonates, and the elderly[6-8]. Although K. pneumoniae is not a dominant member of the healthy gut microbiota, it is found at low levels in the gastrointestinal tract and its abundance increases significantly under conditions such as antibiotic exposure, hypertension, prolonged hospitalization, prematurity, and inflammatory bowel disease[9-14]. These findings have led to the theory that the gastrointestinal tract serves as a reservoir for K. pneumoniae, from which the bacterium can translocate across the intestinal epithelium and disseminate to extraintestinal sites[15-19].
A key barrier to microbial translocation is the intestinal mucus layer, which protects the underlying epithelium from potentially harmful microbes. Intestinal mucus is composed largely of the mucin protein Mucin 2 (MUC2), which is decorated with O-linked glycans, such as N-acetyl-neuraminic acid, fucose, galactose, N-acetyl-glucosamine (GlcNAc), and N-acetyl-galactosamine (GalNAc)[20,21]. Certain gut bacteria have evolved the ability to cleave and consume mucin glycans as nutrient sources, either for themselves or through cross-feeding other microbes[20,22]. Additionally, some bacteria use mucin binding proteins to adhere to and colonize the mucus layer[23,24]. K. pneumoniae has been identified in the mucus layer of mice and humans[25,26] and Hudson et al. identified that K. pneumoniae harbors a fucose operon and that L-fucose metabolism is an important factor for K. pneumoniae colonization in vivo[27]. However, apart from these studies, little is known about the relationship between K. pneumoniae and mucus. We hypothesized that K. pneumoniae would be able to adhere to the mucus layer and potentially degrade mucin glycans to support its growth.
METHODS
Bacterial growth conditions
The following commercially available and clinical isolates of Klebsiella pneumoniae were used in this study: American Type Culture Collection (ATCC) 35657, ATCC 700603, CB1, F64162, 91901, and MCFM1-30. K. pneumoniae strains denoted with MCFM were kindly provided by Terri Ellis. All clinical isolates used in this study were fully de-identified bacterial specimens with no associated patient information or protected health identifiers. These isolates were collected as part of routine clinical care, banked, and provided to the investigators without any link to patient identity. In accordance with U.S. federal regulations [45 Code of Federal Regulations (CFR) 46] and Institutional Review Board (IRB) policy, use of de-identified microbial isolates does not constitute human subjects research and therefore did not require IRB review or approval. K. pneumoniae strains were streak plated onto K. pneumoniae specific agar plates (Millipore catalogue # 90925) containing the Klebsiella Selective Supplement (Millipore catalogue # 15821) and then subsequently plated on brain-heart infusion (BHI) agar plates for purity. Single colonies were used to inoculate BHI broth and grown at 37 °C in a shaking at 120 rpm in an aerobic incubator overnight before being used for downstream assays. As a control for mucus adhesion, we also grew Limosilactobacillus reuteri (L. reuteri) ATCC 6475 on De Man Rogosa Sharpe (MRS) agar and single colonies were used to inoculate MRS broth.
Fluorescently-tagging bacteria
K. pneumoniae strains were grown overnight as described above at a total volume of 5 mL. The following day, cultures were washed twice with sterile phosphate-buffered saline (PBS) and resuspended in 10 mL of sterile PBS. Carboxyfluorescein diacetate succinimidyl ester (CFDA-SE, ThermoFisher #C1157) was added to the bacteria in PBS at a final concentration of 50 µg/mL. Cultures were incubated in the dark at 37 °C aerobically for 1 h. Cultures were washed three times with sterile PBS to remove excess dye. To assess potential variability in dye incorporation across strains, baseline fluorescence was measured for each labeled culture and normalized to optical density at 600 nm (OD600). Fluorescence intensity per optical density (OD) unit was comparable across all K. pneumonaie strains tested, indicating minimal differences in CFDA-SE uptake. Labeled bacterial suspensions were subsequently adjusted to the same OD600 prior to use in adhesion assays to ensure equivalent input across conditions.
Mucus preparation and binding assay
Porcine MUC2 (intestinal) mucus (MyBioSource #MBS5311023) was dialyzed in 7,000 MWCO SnakeskinTM Dialysis Tubing (ThermoFisher #68700) against water for three days to remove impurities or unbound glycans. The resulting mucus solution was lyophilized and resuspended in water before downstream use. Human intestinal mucus was collected from the mucus producing cell line HT29-MTX and MUC2 was isolated as previously described[28]. This mucus was also dialyzed and lyophilized. Mucus-coated 96-well plates were prepared by adding 1 mg/mL mucus in Hanks' Balanced Salt Solution (HBSS) (VWR #02-0121-0500) to non-tissue culture treated 96-well plates (VWR; Cat# 10861-562) overnight at 4 °C. As a negative control, plates were coated with 1 mg/mL of methylcellulose (Fisher Scientific #M352-500) in HBSS. After the overnight incubation, the plates were washed twice with sterile PBS. The OD600 of fluorescently tagged bacteria was measured to determine the concentration of bacteria. Tagged-bacteria was put to an OD600 = 1.0 in PBS and 100 µL was plated into respective wells of the mucus-coated 96-well plates. The plates were incubated in the dark at 37 °C for 1 h. The plates were washed twice with sterile PBS to remove unbound bacteria. The fluorescence (excitation: 485 nm; emission: 528 nm) was read to determine adhered bacteria. As confirmation of adhesion, fluorescently tagged K. pneumoniae and a positive control L. reuteri was added at an OD600 =2, 1, 0.5, and 0.1 to generate an adhesion curve. To examine whether K. pneumoniae ATCC 35657 could adhere to human intestinal mucus, fluorescently tagged K. pneumoniae was added at OD600 = 1.0 in PBS to glass coverslips (18 mm) coated with 1 mg/ml human intestinal mucus using 3-aminopropyltriethoxysilane (APTS) as previously described[28,29]. K. pneumoniae was incubated with coverslips at 37 °C for 1 h. After washing, adhesion was examined on a Zeiss Axio microscope with a 60× objective.
Growth and biofilm assay
Bacteria were grown overnight in BHI before being sub-cultured into a M9 minimal medium (ThermoFisher #A 1374401). The M9 medium was supplemented with water, 25 mM D-glucose (FisherSci #A16828-36), L-fucose (FisherSic #A16789-03), N-acetyl-neuraminic acid (Neu5AC, FisherSci #L13950-MC), D-galactose (FisherSci # A12813-0B), GlcNAc, FisherSci # A13047-22, GalNAc, FisherSci # J66095-06, or dialyzed porcine intestinal mucus or human intestinal mucus (1, 0.5 and 0.25 mg/mL) and plated onto a non-tissue culture treated 96-well plate (VWR; Cat# 10861-562). Bacteria were added to the M9 medium at an optical density OD600 of 0.1. Plates were incubated at 37 °C and read on Synergy HT plate reader at OD600 every 10 minutes for 18 hours. For biofilm assays, we coated non-tissue culture treated 96-well plates (VWR; Cat# 10861-562) with 1 mg/mL porcine MUC2 mucus or 1 mg/mL of methylcellulose in HBSS overnight at 4 °C. After washing, we added K. pneumoniae at OD600 of 0.1 in BHI to the plates and incubated them at 37 °C for 48 h. Following the incubation, we washed the plates with PBS and stained with 0.1% crystal violet for 1 h at 37 °C. After washing, the crystal violet was solubilized in alcohol-acetone (80%/20%) solvent and the absorbance values at 570 nm were read on a Synergy H1 plate reader.
Antibiotics resistance curve
Bacteria were grown overnight in BHI before being sub-cultured into a Luria-Bertani (LB) medium (ThermoFisher #A 1374401). The LB medium was supplemented with water or 1 mg/mL human MUC2 mucus and used to generate a serial dilution of the aminoglycoside antibiotics gentamicin, kanamycin, and streptomycin (2,048 μg/mL to 2 μg/mL). K. pneumoniae was added at an OD600 of 0.1. Plates were incubated at 37 °C and read on Synergy HT plate reader at OD600 every 10 min for 18 h. The half maximal inhibitory concentration (IC50) value was calculated for each antibiotic at the 18 h time point.
Quantitative real time polymerase chain reaction assay
K. pneumoniae strains CB1 and ATCC 35657 were grown in a chemically defined minimal medium (CDMM) supplemented with either water or 1 mg/mL dialyzed porcine intestinal mucus and incubated for 16 h at 37 °C in a shaking aerobic incubator. RNA was extracted from the bacteria using the Quick-RNA MiniPrep Kit (Zymo #R1055) per the manufacturer’s instructions with the addition of bead beating. Briefly, bacterial pellets were resuspended in 100 µL STE buffer [100 mM NaCl, 10 mM Tris-HCl, pH 8.0, 1 mM ethylenediaminetetraacetic acid (EDTA)], and transferred to 1.5 mL screw-top tubes containing 0.1 mm glass beads. Bacteria were disrupted on a Bedbug bead homogenizer (Benchmark Scientific) for 20 s at 4.0 m/s, then Zymo Quick RNA lysis buffer was added to each tube, and samples were disrupted for a second time. Following these steps, RNA was isolated according to the manufacturer’s protocol. RNA quality was assessed by Nanodrop (ThermoScientific) and was then used to make cDNA with the Verso cDNA Synthesis Kit (ThermoScientific #AB1453) per the manufacturer’s instructions. Quantitative real time polymerase chain reaction (qPCR) was performed using qPCRBIO SyGreen Mix lo-ROX (PCRBiosystems #PB20.11). The data was analyzed using the ΔddCT method.
Mouse model
All the experiments involving animals were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at the Medical University of South Carolina (protocol #2021-01286). All experimental procedures strictly adhered to the guidelines set forth by the US National Institutes of Health (NIH) Office of Laboratory Animal Welfare (OLAW) and complied with the regulations under the Animal Welfare Act and the Public Health Service (PHS) Policy on Humane Care and Use of Laboratory animals. Male C57BL/6J mice (10 weeks of age) were purchased from Jackson Labs (#000664) and housed under standard specific pathogen-free conditions at controlled temperature and humidity (22 ± 2 °C and 40%-60% humidity) with a 12-h light/dark cycle and ad libitum access to food and water. Mice were acclimated for at least one week prior to experimentation and were randomly assigned to experimental groups. Investigators were not blinded during the experimental procedures, but were blinded during image analysis. For the experiment, the drinking water was switched with antibiotic water that contained 0.4 mg/mL kanamycin, 35 µg/mL gentamicin, 68 µg/mL colistin, 0.215 mg/mL metronidazole, and 45 µg/mL vancomycin. After 5 days, the antibiotic water was switched with standard autoclaved drinking water and mice received a single dose intraperitoneal injection of 0.25 mg of clindamycin in water. Mice were randomly assigned to two groups (n = 5 per group; total of 10 mice in the experiment) and were orally gavaged with sterile PBS (vehicle control) or 107 colony forming units (CFU) K. pneumoniae. After 5 days, mice were humanely euthanized and the colon was fixed in Carnoy’s fixative. Fluorescent in situ hybridization (FISH) staining was performed on 5 µm thick sections of colon on glass slides. Slides were hybridized with the probe [Cy3] CACCTACACACCAGC[30] (Integrated DNA Technologies, IDT) in a humidified dark chamber at 51 °C for 45 min. Slides were then washed in pre-warmed washing buffer and water and and counter-stained with the mucus glycan targeting lectin Ulex europaeus agglutinin I (UEA I-FITC; VectorLabs #FL-1061-2) for 1 h at room temperature. After washing, the slides were then stained with the nuclear stain Hoechst 33424 (FisherSci # 51-17) for 10 min at room temperature. Sections were analyzed by fluorescence microscopy on a Zeiss Axio microscope with a 20× objective.
Data mining
To assess the distribution of mucus-associated glycosyl hydrolases among K. pneumoniae genomes, we queried the Integrated Microbial Genomes (IMG) database v5.0 (http://img.jgi.doe.gov)[31], which is available through the Joint Genomes Institute (JGI) (Version 6.0) on March 26th, 2022. Enzyme Commission (EC) numbers for mucus-degrading enzymes were input into the gene search tool of the IMG database and K. pneumoniae bacterial genomes were binned for analysis. Genomes that harbored at least one gene copy of a specified EC were considered to possess that function. To determine how many K. pneumoniae strains possessed an EC function, we used the following equation[32]:
Statistics and graphs
All graphs were generated using GraphPad Prism software (version 10.0.03) (GraphPad Inc.). Overnight growth curves and biofilm data was analyzed by Two Way ANOVA with the Tukey post-hoc test. Adhesion was analyzed by One Way Analysis of Variance (ANOVA) with the Tukey post-hoc test. IC50 values for antibiotic susceptibility were calculated from a four-parameter logistics curve. Each dot in the graph represents an individual data point. The data are presented as mean ± standard deviation, with P < 0.05 (*) considered statistically significant.
RESULTS
K. pneumoniae adheres to intestinal mucus
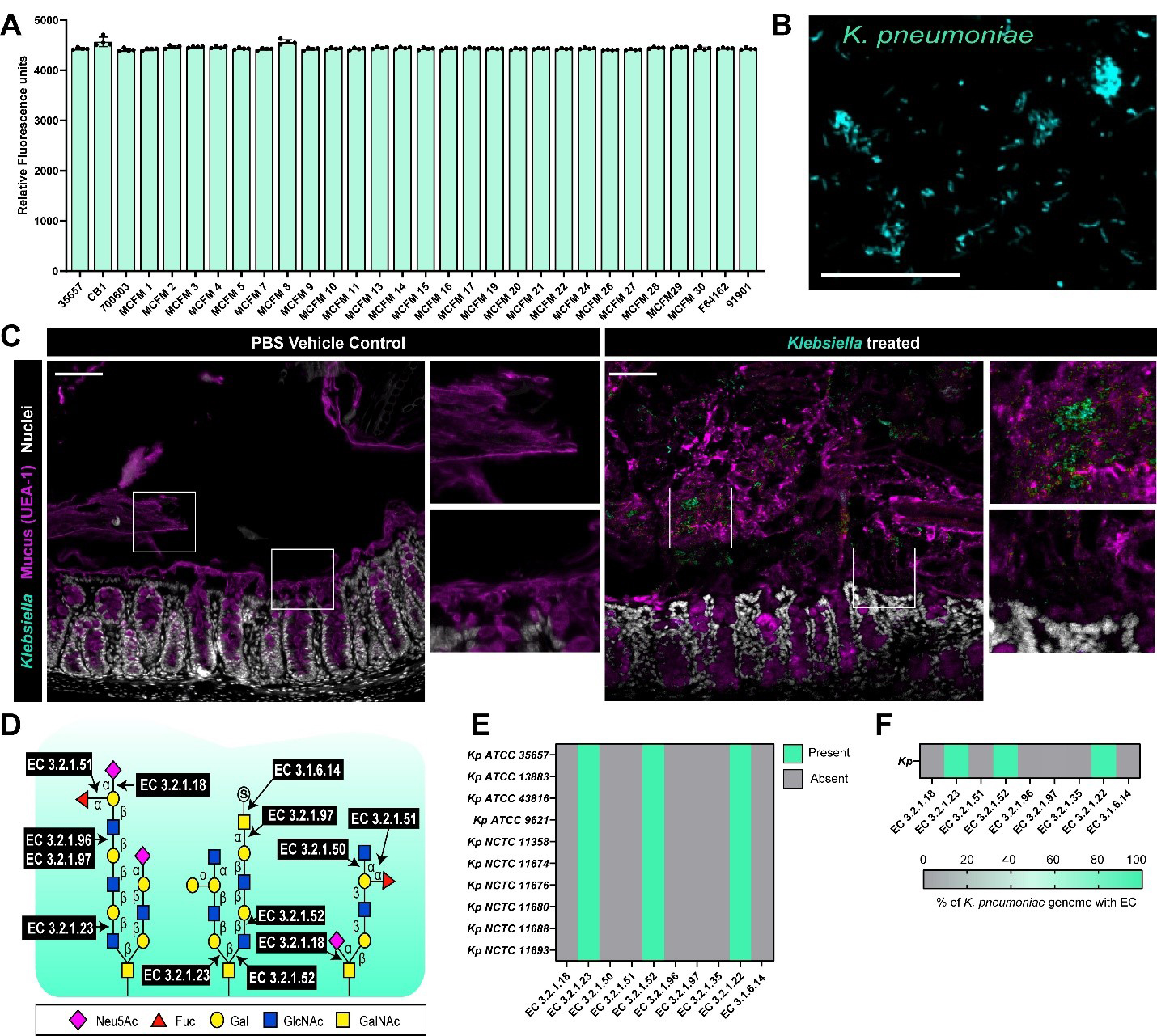
To assess the ability of K. pneumoniae to adhere to intestinal mucus, we fluorescently tagged multiple commercially available (ATCC 35657, ATCC 700603, CB1,) and clinical isolates (MCFM1-30, F64162, 91901) of K. pneumoniae and examined adhesion to porcine MUC2 mucus. We found that all the K. pneumoniae strains adhered to mucus [Figure 1A], suggesting that mucus adhesion is a conserved feature among K. pneumoniae strains. As additional confirmation, we serially diluted K. pneumoniae and a positive control L。 reuteri, which is known to adhere to mucus[33-35], on mucus coated plates. We found that both L. reuteri and K. pneumoniae exhibited a dose-dependent adhesion to porcine MUC2 mucus [Supplementary Figure 1A and B]. As a negative control we also examined the adhesion of K. pneumoniae to methylcellulose; a polymer that is commonly used to mimic the viscosity of mucus, but lacks the glycans for specific adhesion[28]. Although we did note some non-specific adhesion to methylcellulose, the adhesion was far lower than observed for mucus [Supplementary Figure 1C]; suggesting that K. pneumoniae adheres to mucus. To confirm that K. pneumoniae was capable of binding to human intestinal mucus, we added fluorescently tagged K. pneumoniae ATCC 35657 to human MUC2 on glass-coated coverslips [Figure 1B]. Microscopy revealed robust bacterial adhesion, with numerous K. pneumoniae cells bound to the MUC2 mucus. To assess if K. pneumoniae was adept at adhering to mucus in vivo, we oral gavaged adult C57B6/J mice with either PBS (vehicle control) or K. pneumoniae ATCC 35657 and examined mucus adhesion by FISH. As expected, we saw no K. pneumoniae in the PBS treated animals, but we found K. pneumoniae adhered to the mucus layer above the epithelium in the K. pneumoniae treated animals [Figure 1C]. These data indicate that K. pneumoniae adheres to intestinal mucus in vitro and in vivo.
Figure 1. K. pneumoniae can adhere to intestinal mucus. (A) K. pneumoniae strains were fluorescently tagged and added to plates coated with 1 mg/mL dialyzed porcine intestinal mucus and adhesion was examined by a plate reader after 1 h incubation. Data presented as the fluorescence of adhered bacteria to mucus (mean ± standard deviation); (B) Representative fluorescent microscopy image of K. pneumoniae ATCC 35657 adhered to human MUC2-coated coverslips, scale bar = 50 μm; (C) Mice were orally administered with PBS (vehicle control) or K. pneumoniae ATCC 35657 and colons were harvested five days later. K. pneumoniae in the mucus layer was visualized with fluorescent in-situ hybridization (teal), mucus was visualized with UEA-1 staining (purple) and nuclei were identified with Hoechst (white). Scale bar = 50 μm; (D) Schematic of the enzymes required to cleave the bonds attaching mucus-associated sugars; (E) Genome analysis of 10 K. pneumoniae genomes in the IMG database for genes involved in mucus glycan degradation; (F) Genome analysis of the percentage of total K. pneumoniae genomes in the IMG database for genes involved in mucus glycan degradation. Data represents the percentage of genomes with a given EC. Data represents the presence or absence of the target gene. Graphs generated with Graphpad Prism. K. pneumoniae: Klebsiella pneumoniae; PBS: phosphate-buffered saline; EC: Enzyme Commission; ATCC: American Type Culture Collection; MUC2: Mucin 2; IMG: Integrated Microbial Genomes; UEA-1: Ulex europaeus agglutinin I.
K. pneumoniae possesses glycosyl hydrolases necessary for limited mucus degradation
The mucin protein is heavily glycosylated and contains chains of sugars such as galactose, GlcNAc, GalNAc that are terminated by fucose and N-acetyl-neuraminic acid[20]. To identify if K. pneumoniae had the genes necessary for cleaving the glycan linkages in mucus [Figure 1D], we examined 1,052 annotated K. pneumoniae genomes on the Joint Genome Institute’s IMG database and queried the number of genomes that possessed the glycosyl hydrolases required for mucus glycan degradation [Figure 1E]. We first examined the genomes of commercially available K. pneumoniae strains and found that all the strains possessed at least one gene copy of β-galactosidases (EC 3.2.1.23), and α-galactosidases (EC 3.2.1.22); suggesting that these bacteria could remove galactose residues [Figure 1E]. We also found that these K. pneumoniae strains possessed β-N-acetyl-hexosaminidases (EC 3.2.1.52), which remove GlcNAc and GalNAc residues. We found a similar pattern in the > 1,000 annotated K. pneumoniae genomes, with ~ 98% of K. pneumoniae genomes having β-galactosidases (EC 3.2.1.23), α-galactosidases (EC 3.2.1.22), and β-N-acetyl-hexosaminidases (EC 3.2.1.52) [Figure 1F]. Interestingly, none of the K. pneumoniae strains possessed fucosidases (EC 3.2.1.51), neuraminidases (EC 3.2.1.18) or sulfatases (EC 3.1.6.14). These data suggest that K. pneumoniae encode enzymes for cleaving internal glycan sugars such as galactose, GlcNAc and GalNAc residues, but lacks the enzymes necessary to remove terminal fucose, N-acetyl-neuraminic acid, or sulfate groups.
K. pneumoniae possesses glycosyl transferases necessary for the uptake of mucus associated sugars
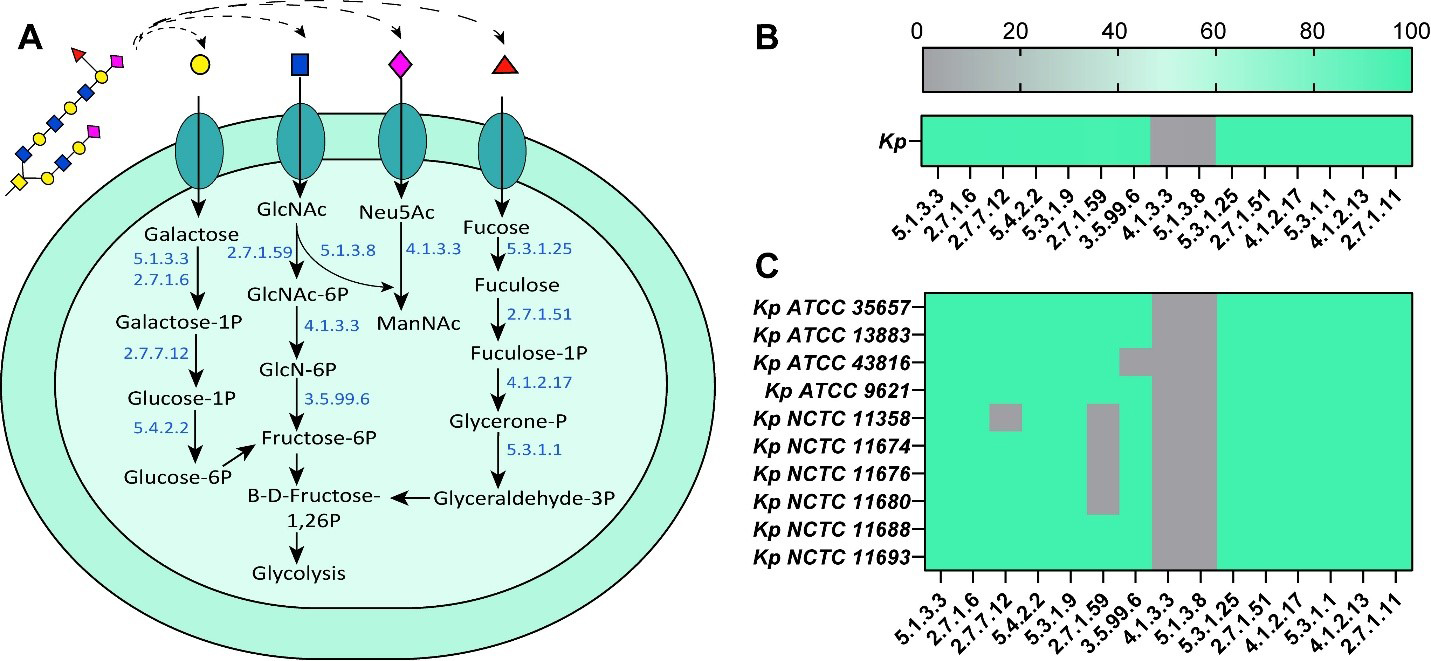
We next examined if K. pneumoniae possessed the downstream pathways required for processing mucin-derived sugars. Using the IMG database, we queried K. pneumoniae genomes for key sugar utilization pathways [Figure 2A]. Overall, the K. pneumoniae genomes encoded numerous enzymes involved in processing galactose, GlcNAc, fructose, and fucose metabolism, including D-galactose-1-phosphotransferase (EC 2.7.1.6), α-D-galactose-1-phosphate uridylyltransferase (EC 2.7.7.12), N-acetyl-D-glucosamine 6-phosphotransferase (EC 2.7.1.59), β-D-fructose-6-phosphate 1-phosphotransferase (EC 2.7.1.11), fructose-bisphosphate aldolase (EC 4.1.2.13), L-fuculose-phosphate aldolase (EC 4.1.2.17), glucose-6-phosphate isomerase (EC 5.3.1.9), and α-D-glucose-1,6-phosphomutase (EC 5.4.2.2) [Figure 2B]. Analysis of the commercially available strains mirrored this pattern, with most strains harboring the same metabolic pathways identified across the > 1,000 genomes [Figure 2C]. At the individual strain level, however, we noted some variation; for example, K. pneumoniae National Collection of Type Cultures (NCTC) 11358 lacked several enzymes required for converting galactose-1-phosphate to glucose-1-phosphate and GlcNAc-6-phosphate to GlcN-6-phosphate [Figure 2C]. None of K. pneumoniae genomes had EC 5.1.3.8 to convert GlcNAc to N-Acetylmannosamine (ManNAc) or EC 4.1.3.3 to convert N-acetyl-neuraminic acid (Neu5Ac) (sialic acid) to ManNAc. Together, these findings indicate that K. pneumoniae broadly encodes the metabolic capacity to utilize liberated mucin-derived sugars, although specific pathway gaps may contribute to strain-level differences in sugar utilization.
Figure 2. K. pneumoniae possesses multiple enzymes necessary for mucus-associated sugar metabolism. (A) Schematic of the necessary genes for the down-stream processing of mucus-associated sugars; (B) Genome analysis of the percent of K. pneumoniae genomes in the IMG database for the presence sugar processing associated genes; (C) Genome analysis of 10 K. pneumoniae genomes in the IMG database for the presence of sugar-processing associated genes. Data represents the presence or absence of the target gene. Graphs generated with Graphpad Prism. GlcNAc: N-acetyl-glucosamine; Neu5Ac: N-acetyl-neuraminic acid; ATCC: American Type Culture Collection; NCTC: National Collection of Type Cultures; K. pneumoniae: Klebsiella pneumoniae; IMG: Integrated Microbial Genomes.
K. pneumoniae can use individual mucin glycans, but not intact mucus, for growth
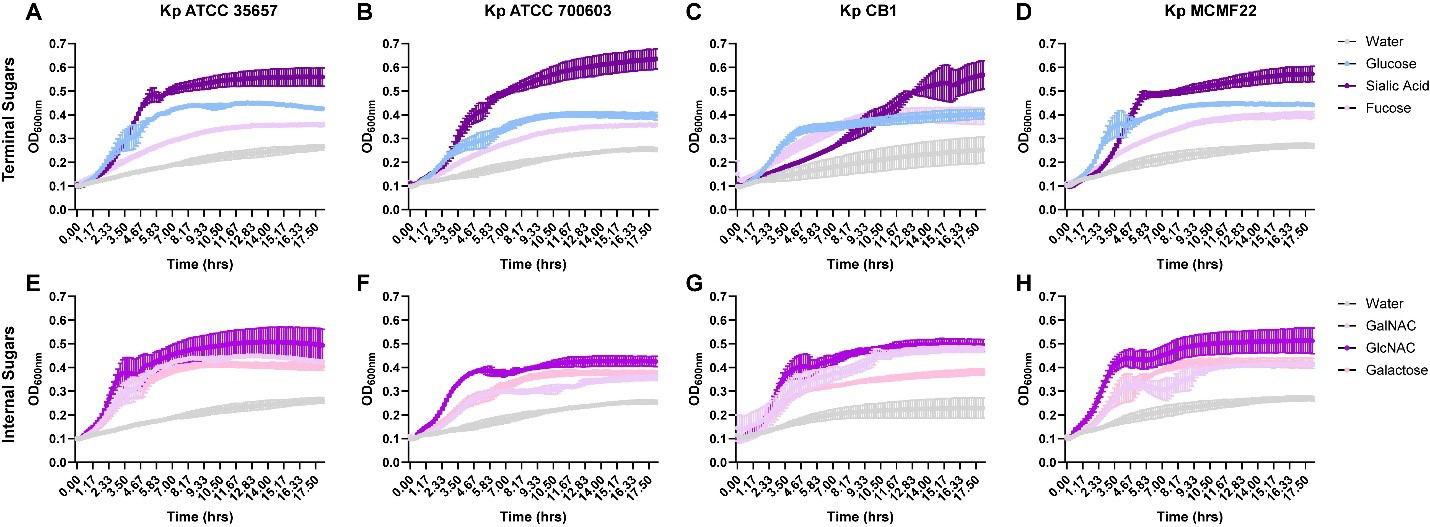
We next tested whether K. pneumoniae could utilize mucus-associated sugars as a fuel source. To do this, we grew K. pneumoniae ATCC 35657, ATCC 700603, CB1, and MCFM22 in M9 minimal medium supplemented with 25 mM of individual mucin-derived monosaccharides and monitored growth over time. As a positive control for carbon utilization, we included 25 mM glucose. As expected, we found that all K. pneumoniae strains had enhanced growth with our positive control glucose [Figure 3A-D and Table 1]. We found that 25 mM of the terminal glycan sugars N-acetyl-neuraminic acid and L-fucose were able to significantly increase K. pneumoniae growth throughout the exponential and lag phase [Figure 3A-D]. We likewise found that 25 mM of the internal glycan sugars galactose, GalNAc and GlcNAc promoted an increase in growth during exponential and lag phase of all our K. pneumoniae strains when compared to the water control [Figure 3E-H and Table 1].
Figure 3. K. pneumoniae can use mucus-associated sugars as a fuel source. K. pneumoniae ATCC 35657 (A and E), ATCC 700603 (B amd F), CB1 (C and G), and MCFM22 (D and H) were grown in a M9 minimal medium supplemented with (A-D) 25 mM D-glucose, N-acetyl-neuraminic acid, L-fucose or (E-H) 25 mM D-galactose, GlcNAc, and GalNAc for 18 h. The OD600 was measured by plate reader to monitor growth over time. Data are represented as mean ± standard deviation. Graphs generated with Graphpad Prism. K. pneumoniae: Klebsiella pneumoniae; ATCC: American Type Culture Collection; GlcNAc: N-acetyl-glucosamine; GalNAc: N-acetyl-galactosamine; OD600: optical density at 600 nm.
Statistics of K. pneumoniae strains grown in M9 minimal medium with 25 mM mucin glycan sugars
| Comparison | Kp ATCC 35657 | Kp ATCC 700603 | Kp CB1 | Kp MCFM22 |
| Water vs. Glucose | < 0.0001 | < 0.0001 | < 0.0001 | < 0.0001 |
| Water vs. Galactose | < 0.0001 | < 0.0001 | < 0.0001 | < 0.0001 |
| Water vs. GalNAc | < 0.0001 | < 0.0001 | < 0.0001 | < 0.0001 |
| Water vs. GlcNAc | < 0.0001 | < 0.0001 | < 0.0001 | < 0.0001 |
| Water vs. Fucose | 0.0009 | 0.0001 | < 0.0001 | < 0.0001 |
| Water vs. Neu5Ac | < 0.0001 | < 0.0001 | < 0.0001 | < 0.0001 |
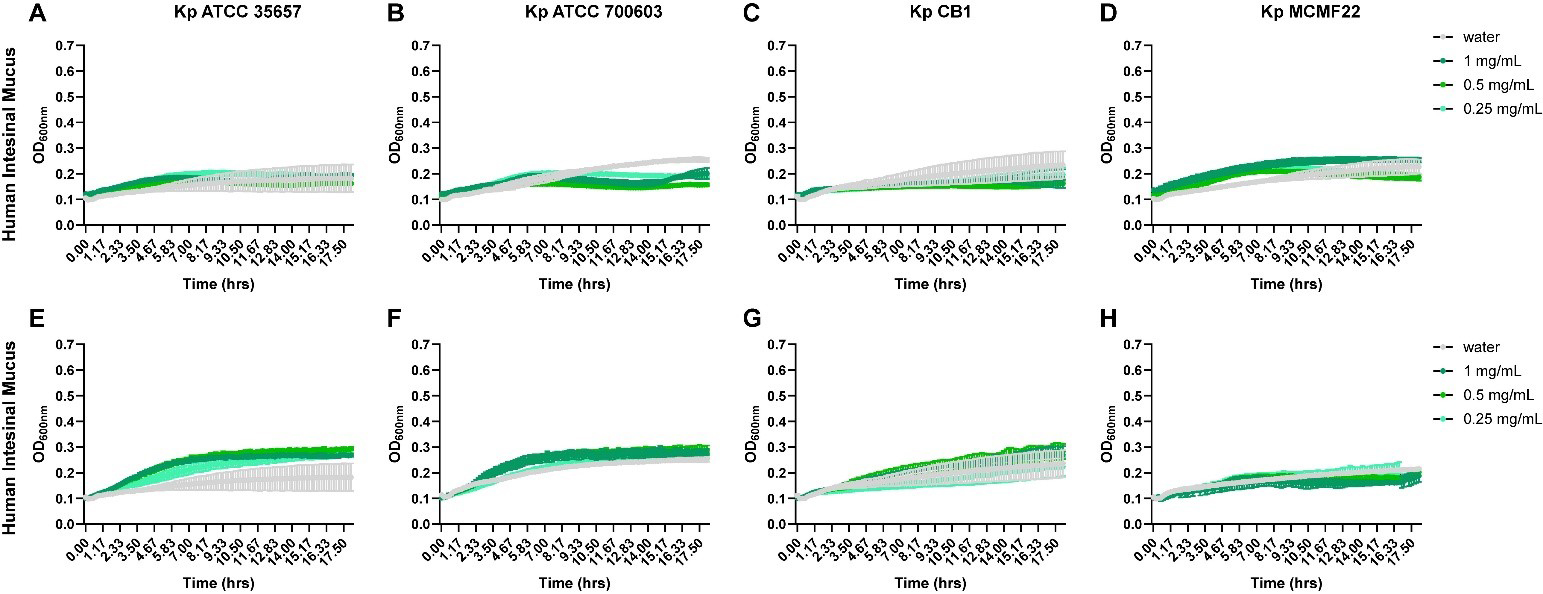
We next tested if K. pneumoniae could utilize intact mucus as a fuel source by growing our four strains of K. pneumoniae in M9 supplemented with different concentrations of porcine MUC2 mucus [Figure 4A-D and Table 2] or human MUC2 mucus [Figure 4E-H and Table 2]. None of the K. pneumoniae strains could use porcine or human MUC2 to support growth [Figure 4]. These data suggests that while K. pneumoniae can metabolize liberated mucin sugars, it lacks the enzymes required to remove terminal glycan residues and thus cannot substantially degrade mucus and consume the released glycans.
Figure 4. K. pneumoniae is unable to use intact mucus as a fuel source. K. pneumoniae ATCC 35657 (A and E), ATCC 700603 (B and F), CB1 (C and G), and MCFM22 (D and H) were grown in a M9 minimal medium supplemented with (A-D) porcine MUC2 (1, 0.5, and 0.25 mg/mL) or (E and H) human MUC2 (1, 0.5, and 0.25 mg/mL) for 18 h. The OD600 was measured by plate reader to monitor growth over time. Data are represented as mean ± standard deviation. Graphs generated with Graphpad Prism. ATCC: American Type Culture Collection; OD600: optical density at 600 nm; K. pneumoniae: Klebsiella pneumoniae; MUC2: Mucin 2.
Statistics of K. pneumoniae strains grown in M9 minimal medium with porcine or human MUC2
| Mucus type | Comparison | Kp ATCC 35657 | Kp ATCC 700603 | Kp CB1 | Kp MCFM22 |
| Porcine MUC2 | Water vs. 1 mg/mL | > 0.9999 | 0.2129 | 0.1137 | 0.9998 |
| Porcine MUC2 | Water vs. 0.5 mg/mL | > 0.9999 | 0.1664 | 0.1001 | 0.9992 |
| Porcine MUC2 | Water vs. 0.25 mg/mL | > 0.9999 | 0.2006 | 0.125 | 0.9994 |
| Human MUC2 | Water vs. 1 mg/mL | 0.9996 | 0.9991 | > 0.9999 | 0.9966 |
| Human MUC2 | Water vs. 0.5 mg/mL | > 0.9999 | > 0.9999 | > 0.9999 | 0.9971 |
| Human MUC2 | Water vs. 0.25 mg/mL | 0.9998 | 0.9996 | 0.9995 | > 0.9999 |
The presence of mucus does not impact biofilm formation or virulence genes, but it does alter K. pneumoniae antibiotic resistance
To determine whether mucus influences K. pneumoniae community behavior, we next examined biofilm formation in the presence and absence of porcine MUC2. Although we observed strain-dependent differences in baseline biofilm production, with notably higher biofilm formation by CB1 and MCFM22 compared to ATCC 35657 and ATCC 700603, the addition of mucus did not alter biofilm levels for any strain [Supplementary Figure 1D]. These findings indicate that while biofilm capacity varies across K. pneumoniae isolates, mucus does not modulate biofilm production under these conditions. We next sought to determine whether mucus alters the expression of well-characterized K. pneumoniae virulence factors. K. pneumoniae CB1 and ATCC 35657 were grown in the presence or absence of porcine MUC2, and gene expression was assessed by qPCR [Supplementary Figure 2A-L]. The presence of mucus did not significantly alter the expression of genes associated with iron acquisition (iroD), urease production (ureA), type 1 fimbriae (fimH), fatty acid biosynthesis (clbA, clbB), or allantoin metabolism (allS). These findings suggest that mucus does not impact canonical virulence pathways. Finally, we examined the impact of mucus on antibiotic resistance. K. pneumoniae is well recognized for its intrinsic resistance to many antibiotics and its capacity to rapidly acquire additional resistance determinants[2,8,36,37]. We examined growth with the antibiotics gentamicin, kanamycin and streptomycin. Across all four K. pneumoniae strains tested, and the presence of human MUC2 consistently lowered the IC50 values for gentamicin, kanamycin and streptomycin, indicating increased antibiotic sensitivity in the presence of mucus [Table 3]. For gentamicin, IC50 values decreased by approximately 45%-65% across strains, with ATCC 35657 dropping from 5.6 to 2.6 µg/mL and MCFM22 from 6.6 to 2.4 µg/mL. A similar trend was observed for kanamycin and streptomycin, where IC50 values declined by ~ 25%-55% and ~ 20%-60% respectively. These results demonstrate that mucus does not protect K. pneumoniae from antibiotic activity; instead, mucus enhances susceptibility to aminoglycosides across diverse strains.
Calculated IC50 values for K. pneumoniae strains grown in LB with water (control) or human MUC2 mucus with the antibiotics gentamicin, kanamycin and streptomycin
| Gentamicin | Kanamycin | Streptomycin | ||||
| Strain | Control | MUC2 | Control | MUC2 | Control | MUC2 |
| ATCC 35657 | 5.64 | 2.56 | 10.80 | 7.97 | 8.21 | 2.43 |
| ATCC 700603 | 7.00 | 2.97 | 18.09 | 10.50 | 3.09 | 1.84 |
| CB1 | 7.57 | 3.78 | 11.16 | 5.13 | 7.98 | 5.05 |
| MCMF22 | 6.63 | 2.41 | 17.95 | 13.66 | 5.01 | 4.45 |
DISCUSSION
In this study, we evaluated how K. pneumoniae strains interact with intestinal mucus and how this interaction influences bacterial behavior. We found that multiple K. pneumoniae strains adhered robustly to porcine and human MUC2, and that K. pneumoniae ATCC 35657 also localized to the mucus layer in the mouse colon in vivo, demonstrating that mucus adhesion is a conserved trait. Genomic analyses of more than 1,000 K. pneumoniae genomes revealed that the species broadly encodes glycosyl hydrolases capable of processing internal mucin-derived sugars. However, K. pneumoniae lacks enzymes required to cleave terminal fucose, N-acetyl-neuraminic acid, and sulfate groups and was unable to grow on intact MUC2 as a sole carbon source, indicating a limited capacity for direct mucin degradation. Consistent with the K. pneumoniae genomic profile, all strains readily used free mucin-derived monosaccharides, including galactose, GlcNAc, GalNAc, fucose, and N-acetyl-neuraminic acid; highlighting a reliance on liberated sugars rather than intact mucin glycans. Although strains varied in their baseline biofilm formation, the presence of mucus did not alter biofilm production for any strain. Unexpectedly, however, exposure to MUC2 increased K. pneumoniae susceptibility to the aminoglycosides gentamicin, kanamycin and streptomycin across all strains tested. Together, these findings establish that while K. pneumoniae efficiently adheres to intestinal mucus and exploits available mucin-derived sugars, intact mucus enhances antibiotic sensitivity, revealing mucus as an important modulatory factor in K. pneumoniae colonization dynamics and treatment response.
While K. pneumoniae is not classically considered a predominant member of the gut microbiota[38-41], it has been shown to be elevated during different intestinal disease states[16,42-48], such as inflammatory bowel disease (IBD)[49-53] and Clostridioides difficile infection[54,55]. A common feature of these conditions is a disrupted microbial community, and the loss of key commensal organisms likely creates ecological niches that allow opportunistic bacteria such as K. pneumoniae to expand[44,56]. In the absence of microbial competitors, we propose that K. pneumoniae can bind to the intestinal mucus layer and utilize mucus-derived nutrients to support its growth. Despite possessing a limited repertoire of mucin-degrading enzymes, K. pneumoniae may benefit from cross-feeding by classic mucin degraders, such as Bacteroides thetaiotaomicron, Ruminococcus torques, or Akkermansia muciniphila[20]. Such cooperative interactions are well documented in the gut[57,58] and may similarly facilitate K. pneumoniae persistence within the mucus layer. Although this model requires further investigation, these potential interactions have important clinical implications for patients experiencing microbiota disruption.
N-acetyl-neuraminic acid import and utilization is important for a variety of enteric pathogens[58-61]. An interesting observation from our study was that K. pneumoniae did not encode the enzymes required to convert Neu5Ac to ManNAc (EC 4.1.3.3), yet N-acetyl-neuraminic acid still supported bacterial growth. This could be because K. pneumoniae is able to import N-acetyl-neuraminic acid and channel it into alternative metabolic pathways independent of the canonical ManNAc route. Several bacteria, including E. coli, Salmonella enterica, and Vibrio species can import N-acetyl-neuraminic acids through NanT-like transporters[62-66]. Inside bacteria, N-acetyl-neuraminic acid can bind to NanR and influence transcription[67,68]. Although Nan clusters have not been previously reported in K. pneumoniae, it is possible that N-acetyl-neuraminic acid may act as an environmental signal to modulate gene expression in K. pneumoniae.
Biofilm formation is a key virulence factor for K. pneumoniae, contributing to persistence, antibiotic resistance, and host colonization[69-76]. Many environmental cues have been shown to enhance K. pneumoniae including low iron availability, limited nutrients, and low-oxygen conditions[77-80]. Mucus is known to impact biofilm production on diverse microbes. For example, mucus enhances biofilm production by the gut pathobiont Clostridioides difficile[28,81] and decreases biofilm production by the respiratory pathobiont Pseudomonas aeruginosa[82,83]. In this study, we found that mucus did not significantly alter biofilm formation by K. pneumoniae. This suggests that while mucus provides an adhesive substrate, it does not inherently promote biofilm development under the conditions tested. Instead, K. pneumoniae may rely on other environmental cues, host factors, or microbial signals to modulate biofilm architecture within the intestinal environment.
One of the most unexpected observations in this study was that mucus increased K. pneumoniae sensitivity to gentamicin, kanamycin, and streptomycin across all strains tested. Aminoglycoside resistance is a hallmark of K. pneumoniae, driven by intrinsic mechanisms and rapid acquisition of resistance determinants[84-86]. Several compounds that are found in the gut could impact K. pneumoniae’s resistance to antibiotics. Antimicrobial peptides are generated by Paneth and goblet cells in the small intestine and goblet cell in the large intestine[87,88] and antimicrobial peptides have been shown to synergize with antibiotics like chloramphenicol, meropenem, rifampicin, or ceftazidime to effectively kill multidrug resistant K. pneumoniae isolates[89]. Additionally, short chain fatty acids that are produced by commensal microbes can synergize with antibiotics like cefoperazone and sulbactam to inhibit Enterobacteriaceae member Escherichia coli[90]. We found that mucus is another gut factor that can alter antibiotic resistance in K. pneumoniae. We observed a consistent reduction in IC50 values for gentamicin, kanamycin, and streptomycin in the presence of human MUC2, suggesting that mucus alters K. pneumoniae physiology in a way that enhances aminoglycoside efficacy. One possibility is that mucus serves as a colonization-associated signal, prompting K. pneumoniae to shift its gene expression toward adhesion or nutrient acquisition pathways, inadvertently increasing susceptibility to antibiotics. This observation may have therapeutic implications in clinical contexts where mucus could be used to potentially enhance antibiotic susceptibility in K. pneumoniae.
Another potential explanation for the increased aminoglycoside sensitivity observed in the presence of mucus is that mucus alters antibiotic uptake in K. pneumoniae. Aminoglycosides require active transport across the bacterial inner membrane, a process that is highly dependent on the proton motive force and cellular respiration proton motive force[84-86,91,92]. Environmental factors that enhance respiration or membrane potential can increase aminoglycoside uptake and bactericidal activity[91,93,94]. Mucus could shift bacterial metabolism toward a more energetically active state, thereby increasing proton motive force-dependent antibiotic uptake. Additionally, mucin glycans could be impacting antibiotic efflux. K. pneumoniae harbors several multidrug efflux transporters that allow these organisms to evade antibiotics[95-98]. Compounds that target these efflux transporters have been shown to improve antibiotic driven killing of K. pneumoniae[98-101]. It is unclear how mucus may be facilitating this, but it is possible that mucus suppresses efflux transporters and thus promotes antibiotic susceptibility. Future studies examining antibiotic resistance pathways are warranted to better understand this phenomenon.
Several limitations should be considered when interpreting these findings. First, while our in vitro assays provided controlled insights into mucus adhesion, sugar utilization, and antibiotic sensitivity, they cannot fully recapitulate the spatial complexity, dynamic mucus turnover, and host factors present in vivo. Second, we examined a subset of K. pneumoniae strains; although these represented both commercial and clinical isolates, additional genomic diversity, especially hypervirulent or carbapenem-resistant strains, may reveal distinct mucus-interaction phenotypes. Third, mucus composition varies across species, intestinal regions, and disease states, and using purified MUC2 may not fully reflect the heterogeneous mucus environment encountered by bacteria in vivo. Finally, our antibiotic assays focused on aminoglycosides, and it remains unknown whether mucus similarly affects susceptibility to other antibiotic classes. Addressing these limitations in the future will deepen our understanding of how K. pneumoniae interacts with the mucus barrier and how these interactions shape gut colonization and infection dynamics.
In conclusion, our findings provide new insight into the ability of K. pneumoniae to bind to mucus and use mucus associated sugars. These interactions shape key aspects of K. pneumoniae physiology, including susceptibility to aminoglycoside antibiotics. Together, our findings highlight that intestinal mucus is an active modulator of K. pneumoniae behavior, with the capacity to influence both microbial ecology and therapeutic responsiveness. Understanding how mucus-derived signals and microbial community dynamics converge to regulate K. pneumoniae fitness will be critical for developing strategies to prevent gut colonization, limit pathogen expansion during dysbiosis, and enhance antibiotic efficacy in clinical settings in the future.
DECLARATIONS
Acknowledgments
The graphical abstract was created with BioRender (https://BioRender.com/ar497tn).
Authors’ contributions
Conceived and designed research: Ticer TD, Engevik MA
Collected data: Ticer TD, Suresh P, Ghoshal S, Tingler AM, Stubler R, Horvath AE, Glover JS, and Engevik MA
Provided bacterial isolates: Ellis TN
Wrote the manuscript: Ticer TD
Obtained research funding for experiments: Engevik MA
All authors edited and revised the manuscript.
Availability of data and materials
The raw data supporting the conclusions of this manuscript will be made available by the authors upon request.
AI and AI-assisted tools statement:
Not applicable.
Financial support and sponsorship
This work was supported by grants from the National Institute of Health T32DK124191-01A1 (Ticer TD), T32GM132055-05 (Tingler AM), ASIP Summer Research Opportunity Program in Pathology (Horvath AE), T32GM132055-01 (Glover JS), P30DK123704 (Engevik MA), P20GM120457 (Engevik MA), P20GM120457 (Engevik MA).
Conflicts of interest
All authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
All the experiments involving animals were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at the Medical University of South Carolina (protocol #2021-01286) and were performed in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Consent for publication
Not applicable
Copyright
© The Author(s) 2026.
Supplementary Materials
REFERENCES
1. Antibiotic resistance threats in the United States, 2019. Available online: https://www.cdc.gov/antimicrobial-resistance/media/pdfs/2019-ar-threats-report-508.pdf.
2. Chen IR, Lin SN, Wu XN, Chou SH, Wang FD, Lin YT. Clinical and microbiological characteristics of bacteremic pneumonia caused by Klebsiella pneumoniae. Front Cell Infect Microbiol. 2022;12:903682.
3. Guo M, Gao B, Su J, et al. Phenotypic and genetic characterization of hypervirulent Klebsiella pneumoniae in patients with liver abscess and ventilator-associated pneumonia. BMC Microbiol. 2023;23:338.
4. Govindan Nadar R, Chackaravarthy G, Ramachandran G, et al. Isolation and molecular identification of biofilm producing P. aeruginosa and K. pneumoniae from urinary tract infections patient urine sample. J Infect Public Health. 2021;14:1875-80.
5. Roach DJ, Sridhar S, Oliver E, et al. Clinical and genomic characterization of a cohort of patients with Klebsiella pneumoniae bloodstream infection. Clin Infect Dis. 2024;78:31-9.
6. Meatherall BL, Gregson D, Ross T, Pitout JD, Laupland KB. Incidence, risk factors, and outcomes of Klebsiella pneumoniae bacteremia. Am J Med. 2009;122:866-73.
7. Hu Y, Yang Y, Feng Y, et al. Prevalence and clonal diversity of carbapenem-resistant Klebsiella pneumoniae causing neonatal infections: a systematic review of 128 articles across 30 countries. PLoS Med. 2023;20:e1004233.
8. Çölkesen F, Tarakçı A, Eroğlu E, et al. Carbapenem-resistant Klebsiella pneumoniae infection and its risk factors in older adult patients. Clin Interv Aging. 2023;18:1037-45.
9. Lepuschitz S, Hauser K, Schriebl A, et al. Fecal Klebsiella pneumoniae carriage is intermittent and of high clonal diversity. Front Microbiol. 2020;11:581081.
10. Lu B, Zhou H, Zhang X, Qu M, Huang Y, Wang Q. Molecular characterization of Klebsiella pneumoniae isolates from stool specimens of outpatients in sentinel hospitals Beijing, China, 2010-2015. Gut Pathog. 2017;9:39.
11. Raffelsberger N, Hetland MAK, Svendsen K, et al. Gastrointestinal carriage of Klebsiella pneumoniae in a general adult population: a cross-sectional study of risk factors and bacterial genomic diversity. Gut Microbes. 2021;13:1939599.
12. Li J, Zhao F, Wang Y, et al. Gut microbiota dysbiosis contributes to the development of hypertension. Microbiome. 2017;5:14.
13. Seki D, Mayer M, Hausmann B, et al. Aberrant gut-microbiota-immune-brain axis development in premature neonates with brain damage. Cell Host Microbe. 2021;29:1558-1572.e6.
14. Yang J, Li Y, Tang N, et al. The human gut serves as a reservoir of hypervirulent Klebsiella pneumoniae. Gut Microbes. 2022;14:2114739.
15. Bray AS, Zafar MA. Deciphering the gastrointestinal carriage of Klebsiella pneumoniae. Infect Immun. 2024;92:e0048223.
16. Hecht AL, Harling LC, Friedman ES, et al. Dietary carbohydrates regulate intestinal colonization and dissemination of Klebsiella pneumoniae. J Clin Invest. 2024;134:e174726.
17. Hyeon JY, Choi YJ, Jung MJ, Lee DH, Song CS, Kim JH. Multidrug-resistant CTX-M-15-positive Klebsiella pneumoniae ST 307 causing bacteremia via gut translocation in a dog. Front Vet Sci. 2023;10:1275822.
18. Hsu CR, Pan YJ, Liu JY, Chen CT, Lin TL, Wang JT. Klebsiella pneumoniae translocates across the intestinal epithelium via Rho GTPase- and phosphatidylinositol 3-kinase/Akt-dependent cell invasion. Infect Immun. 2015;83:769-79.
19. Fung CP, Lin YT, Lin JC, et al. Klebsiella pneumoniae in gastrointestinal tract and pyogenic liver abscess. Emerg Infect Dis. 2012;18:1322-5.
20. Glover JS, Ticer TD, Engevik MA. Characterizing the mucin-degrading capacity of the human gut microbiota. Sci Rep. 2022;12:8456.
21. Bergstrom K, Shan X, Casero D, et al. Proximal colon-derived O-glycosylated mucus encapsulates and modulates the microbiota. Science. 2020;370:467-72.
22. Shuoker B, Pichler MJ, Jin C, et al. Sialidases and fucosidases of Akkermansia muciniphila are crucial for growth on mucin and nutrient sharing with mucus-associated gut bacteria. Nat Commun. 2023;14:1833.
23. Elzinga J, Narimatsu Y, de Haan N, Clausen H, de Vos WM, Tytgat HLP. Binding of Akkermansia muciniphila to mucin is O-glycan specific. Nat Commun. 2024;15:4582.
25. Favre-Bonté S, Licht TR, Forestier C, Krogfelt KA. Klebsiella pneumoniae capsule expression is necessary for colonization of large intestines of streptomycin-treated mice. Infect Immun. 1999;67:6152-6.
26. Caballero S, Carter R, Ke X, et al. Distinct but spatially overlapping intestinal niches for vancomycin-resistant Enterococcus faecium and carbapenem-resistant Klebsiella pneumoniae. PLoS Pathog. 2015;11:e1005132.
27. Hudson AW, Barnes AJ, Bray AS, Ornelles DA, Zafar MA. Klebsiella pneumoniae l-fucose metabolism promotes gastrointestinal colonization and modulates its virulence determinants. Infect Immun. 2022;90:e0020622.
28. Engevik MA, Danhof HA, Auchtung J, et al. Fusobacteriumnucleatum Adheres to Clostridioides difficile via the RadD adhesin to enhance biofilm formation in intestinal mucus. Gastroenterology. 2021;160:1301-1314.e8.
29. Landry RM, An D, Hupp JT, Singh PK, Parsek MR. Mucin-Pseudomonas aeruginosa interactions promote biofilm formation and antibiotic resistance. Mol Microbiol. 2006;59:142-51.
30. Søgaard M, Hansen DS, Fiandaca MJ, Stender H, Schønheyder HC. Peptide nucleic acid fluorescence in situ hybridization for rapid detection of Klebsiella pneumoniae from positive blood cultures. J Med Microbiol. 2007;56:914-7.
31. Chen IA, Chu K, Palaniappan K, et al. The IMG/M data management and analysis system v.6.0: new tools and advanced capabilities. Nucleic Acids Res. 2021;49:D751-63.
32. Engevik KA, Hazzard A, Puckett B, et al. Phylogenetically diverse bacterial species produce histamine. Syst Appl Microbiol. 2024;47:126539.
33. Muscariello L, De Siena B, Marasco R. Lactobacillus cell surface proteins involved in interaction with mucus and extracellular matrix components. Curr Microbiol. 2020;77:3831-41.
34. Engevik MA, Ruan W, Esparza M, et al. Immunomodulation of dendritic cells by Lactobacillus reuteri surface components and metabolites. Physiol Rep. 2021;9:e14719.
35. Jensen H, Roos S, Jonsson H, et al. Role of Lactobacillus reuteri cell and mucus-binding protein A (CmbA) in adhesion to intestinal epithelial cells and mucus in vitro. Microbiology (Reading). 2014;160:671-81.
36. Ernst CM, Braxton JR, Rodriguez-Osorio CA, et al. Adaptive evolution of virulence and persistence in carbapenem-resistant Klebsiella pneumoniae. Nat Med. 2020;26:705-11.
37. Taitt CR, Leski TA, Erwin DP, et al. Antimicrobial resistance of Klebsiella pneumoniae stool isolates circulating in Kenya. PLoS ONE. 2017;12:e0178880.
38. Zheng Y, Ding Y, Xu M, et al. Gut microbiota contributes to host defense against Klebsiella pneumoniae-induced liver abscess. J Inflamm Res. 2021;14:5215-25.
39. Bray AS, Broberg CA, Hudson AW, et al. Klebsiella pneumoniae employs a type VI secretion system to overcome microbiota-mediated colonization resistance. Nat Commun. 2025;16:940.
40. Yang Y, van Schaik W, McNally A, Zong Z. Microbiome-mediated colonization resistance to and countermeasures of Klebsiella pneumoniae. Nat Commun. 2026;17:1761.
41. Mccartney AL, Hoyles L. Host interactions of bioactive molecules produced by Klebsiella spp. Microbiota and Host. 2025;3:e240011.
42. Vornhagen J, Rao K, Bachman MA. Gut community structure as a risk factor for infection in Klebsiella pneumoniae-colonized patients. mSystems. 2024;9:e0078624.
43. Liao CH, Kao KL, Wu SI, Yang CJ. Stool microbiota analysis for abundance of genus Klebsiella among adults and children in endemic area for community Klebsiella pneumoniae infection. J Microbiol Immunol Infect. 2024;57:470-9.
44. Wu T, Xu F, Su C, et al. Alterations in the gut microbiome and cecal metabolome during Klebsiella pneumoniae-induced pneumosepsis. Front Immunol. 2020;11:1331.
45. Calderon-Gonzalez R, Lee A, Lopez-Campos G, et al. Modelling the gastrointestinal carriage of Klebsiella pneumoniae infections. mBio. 2023;14:e0312122.
46. Vornhagen J, Rao K, Bachman MA. Gut community structure as a risk factor for infection in Klebsiell-colonized patients. medRxiv 2023. Available online: https://doi.org/10.1101/2023.04.18.23288742. [accessed 17 April 2026].
47. Moreira de Gouveia MI, Bernalier-Donadille A, Jubelin G. Enterobacteriaceae in the human gut: dynamics and ecological roles in health and disease. Biology (Basel). 2024;13:142.
48. Vega L, Birchenall-Jiménez CI, Aponte A, et al. Contrasting the gut microbiome in Colombian patients with diarrhea: a comparative metagenomic study in hospitalization and emergency room services. Gut Pathog. 2026;18:20.
49. Federici S, Kredo-Russo S, Valdés-Mas R, et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell. 2022;185:2879-2898.e24.
50. Atarashi K, Suda W, Luo C, et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science. 2017;358:359-65.
51. Kharaghani AA, Harzandi N, Khorsand B, Rajabnia M, Kharaghani AA, Houri H. High prevalence of mucosa-associated extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae among iranain patients with inflammatory bowel disease (IBD). Ann Clin Microbiol Antimicrob. 2023;22:86.
52. Qiu P, Ishimoto T, Fu L, Zhang J, Zhang Z, Liu Y. The gut microbiota in inflammatory bowel disease. Front Cell Infect Microbiol. 2022;12:733992.
53. Olamafar E, Yadegar A, Shahrokh S, Ebrahimi MT, Houri H. Pathogenic potential of invasive mucosa-associated Klebsiella pneumoniae isolates in ulcerative colitis: characterization of virulence factors and inflammatory response. Immun Inflamm Dis. 2025;13:e70285.
54. Hong W, Yang J, Cheng Y, et al. Bacteria co-colonizing with Clostridioides difficile in two asymptomatic patients. Open Life Sci. 2019;14:628-37.
55. Wang L, Chen X, Pollock NR, et al. Metagenomic analysis reveals distinct patterns of gut microbiota features with diversified functions in C. difficile infection (CDI), asymptomatic carriage and non-CDI diarrhea. Gut Microbes. 2025;17:2505269.
56. Jiang Q, Xu Q, Kenéz Á, Chen S, Yang G. Klebsiella pneumoniae infection is associated with alterations in the gut microbiome and lung metabolome. Microbiol Res. 2022;263:127139.
57. Engevik MA, Engevik AC, Engevik KA, et al. Mucin-degrading microbes release monosaccharides that chemoattract Clostridioides difficile and facilitate colonization of the human intestinal mucus layer. ACS Infect Dis. 2021;7:1126-42.
58. Ng KM, Ferreyra JA, Higginbottom SK, et al. Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature. 2013;502:96-9.
59. Chang DE, Smalley DJ, Tucker DL, et al. Carbon nutrition of Escherichia coli in the mouse intestine. Proc Natl Acad Sci U S A. 2004;101:7427-32.
60. Jeong HG, Oh MH, Kim BS, Lee MY, Han HJ, Choi SH. The capability of catabolic utilization of N-acetylneuraminic acid, a sialic acid, is essential for Vibrio vulnificus pathogenesis. Infect Immun. 2009;77:3209-17.
61. Pezzicoli A, Ruggiero P, Amerighi F, Telford JL, Soriani M. Exogenous sialic acid transport contributes to group B streptococcus infection of mucosal surfaces. J Infect Dis. 2012;206:924-31.
62. North RA, Horne CR, Davies JS, et al. “Just a spoonful of sugar...”: import of sialic acid across bacterial cell membranes. Biophys Rev. 2018;10:219-27.
63. Vimr ER, Troy FA. Identification of an inducible catabolic system for sialic acids (nan) in Escherichia coli. J Bacteriol. 1985;164:845-53.
64. Severi E, Hosie AH, Hawkhead JA, Thomas GH. Characterization of a novel sialic acid transporter of the sodium solute symporter (SSS) family and in vivo comparison with known bacterial sialic acid transporters. FEMS Microbiol Lett. 2010;304:47-54.
65. Huang YL, Chassard C, Hausmann M, von Itzstein M, Hennet T. Sialic acid catabolism drives intestinal inflammation and microbial dysbiosis in mice. Nat Commun. 2015;6:8141.
66. Almagro-Moreno S, Boyd EF. Insights into the evolution of sialic acid catabolism among bacteria. BMC Evol Biol. 2009;9:118.
67. Horne CR, Venugopal H, Panjikar S, et al. Mechanism of NanR gene repression and allosteric induction of bacterial sialic acid metabolism. Nat Commun. 2021;12:1988.
68. Afzal M, Shafeeq S, Ahmed H, Kuipers OP. Sialic acid-mediated gene expression in Streptococcus pneumoniae and role of NanR as a transcriptional activator of the nan gene cluster. Appl Environ Microbiol. 2015;81:3121-31.
69. Ramakrishnan R, Nair AV, Parmar K, Rajmani RS, Chakravortty D, Das D. Combating biofilm-associated Klebsiella pneumoniae infections using a bovine microbial enzyme. NPJ Biofilms Microbiomes. 2024;10:119.
70. Guerra MES, Destro G, Vieira B, et al. Klebsiella pneumoniae biofilms and their role in disease pathogenesis. Front Cell Infect Microbiol. 2022;12:877995.
71. Li Y, Ni M. Regulation of biofilm formation in Klebsiella pneumoniae. Front Microbiol. 2023;14:1238482.
72. Centeleghe I, Norville P, Hughes L, Maillard JY. Klebsiella pneumoniae survives on surfaces as a dry biofilm. Am J Infect Control. 2023;51:1157-62.
73. Piperaki ET, Syrogiannopoulos GA, Tzouvelekis LS, Daikos GL. Klebsiella pneumoniae: virulence, biofilm and antimicrobial resistance. Pediatr Infect Dis J. 2017;36:1002-5.
74. Zaborskytė G, Coelho P, Wrande M, Sandegren L. Rapid evolution of Klebsiella pneumoniae biofilms in vitro delineates adaptive changes selected during infection. bioRxiv 2024. Available online: https://www.biorxiv.org/content/10.1101/2024.03.16.585345v1.full.pdf.
75. Gomes AÉI, Pacheco T, Dos Santos CDS, et al. Functional insights from KpfR, a new transcriptional regulator of fimbrial expression that is crucial for Klebsiella pneumoniae pathogenicity. Front Microbiol. 2020;11:601921.
76. Beckman RL 4th, Cella E, Azarian T, Rendueles O, Fleeman RM. Diverse polysaccharide production and biofilm formation abilities of clinical Klebsiella pneumoniae. NPJ Biofilms Microbiomes. 2024;10:151.
77. Guilhen C, Charbonnel N, Parisot N, et al. Transcriptional profiling of Klebsiella pneumoniae defines signatures for planktonic, sessile and biofilm-dispersed cells. BMC Genomics. 2016;17:237.
78. Liu K, Tan S, Ye W, Hou L, Fang B. Low-concentration iron promotes Klebsiella pneumoniae biofilm formation by suppressing succinic acid. BMC Microbiol. 2022;22:95.
79. Anderl JN, Zahller J, Roe F, Stewart PS. Role of nutrient limitation and stationary-phase existence in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob Agents Chemother. 2003;47:1251-6.
80. Pacheco T, Gomes AÉI, Siqueira NMG, et al. SdiA, a quorum-sensing regulator, suppresses fimbriae expression, biofilm formation, and quorum-sensing signaling molecules production in Klebsiella pneumoniae. Front Microbiol. 2021;12:597735.
81. Furtado KL, Plott L, Markovetz M, et al. Clostridioides difficile-mucus interactions encompass shifts in gene expression, metabolism, and biofilm formation. mSphere. 2024;9:e0008124.
82. Co JY, Cárcamo-Oyarce G, Billings N, et al. Mucins trigger dispersal of Pseudomonas aeruginosa biofilms. NPJ Biofilms Microbiomes. 2018;4:23.
83. Yang S, Boboltz A, Stern A, Rathi V, Duncan G. Synthetic mucus biomaterials synergize with antibiofilm agents to combat Pseudomonas aeruginosa biofilms. Adv Ther. 2025;8:2400359.
84. Zhang X, Li Q, Lin H, et al. High-Level Aminoglycoside resistance in human clinical Klebsiella pneumoniae complex isolates and characteristics of armA-carrying IncHI5 plasmids. Front Microbiol. 2021;12:636396.
85. Ahmed OB, Asghar AH, Bamaga M, Bahwerth FS, Ibrahim ME. Characterization of aminoglycoside resistance genes in multidrug-resistant Klebsiella pneumoniae collected from tertiary hospitals during the COVID-19 pandemic. PLoS ONE. 2023;18:e0289359.
86. Foudraine DE, Strepis N, Stingl C, et al. Exploring antimicrobial resistance to beta-lactams, aminoglycosides and fluoroquinolones in E. coli and K. pneumoniae using proteogenomics. Sci Rep. 2021;11:12472.
87. Bevins CL, Salzman NH. Paneth cells, antimicrobial peptides and maintenance of intestinal homeostasis. Nat Rev Microbiol. 2011;9:356-68.
88. Tonetti FR, Eguileor A, Llorente C. Goblet cells: guardians of gut immunity and their role in gastrointestinal diseases. eGastroenterology. 2024;2:e100098.
89. Chatupheeraphat C, Peamchai J, Luk-In S, Eiamphungporn W. Synergistic effect and antibiofilm activity of the antimicrobial peptide K11 with conventional antibiotics against multidrug-resistant and extensively drug-resistant Klebsiella pneumoniae. Front Cell Infect Microbiol. 2023;13:1153868.
90. Kadry AA, El-Antrawy MA, El-Ganiny AM. Impact of short chain fatty acids (SCFAs) on antimicrobial activity of new β-lactam/β-lactamase inhibitor combinations and on virulence of Escherichia coli isolates. J Antibiot (Tokyo). 2023;76:225-35.
91. Lang M, Carvalho A, Baharoglu Z, Mazel D. Aminoglycoside uptake, stress, and potentiation in Gram-negative bacteria: new therapies with old molecules. Microbiol Mol Biol Rev. 2023;87:e0003622.
92. Farha MA, Verschoor CP, Bowdish D, Brown ED. Collapsing the proton motive force to identify synergistic combinations against staphylococcus aureus. Chem Biol. 2013;20:1168-78.
93. Webster CM, Shepherd M. A mini-review: environmental and metabolic factors affecting aminoglycoside efficacy. World J Microbiol Biotechnol. 2022;39:7.
94. Taber HW, Mueller JP, Miller PF, Arrow AS. Bacterial uptake of aminoglycoside antibiotics. Microbiol Rev. 1987;51:439-57.
95. Zdarska V, Arcari G, Kolar M, Mlynarcik P. Antibiotic resistance in Klebsiella pneumoniae and related enterobacterales: molecular mechanisms, mobile elements, and therapeutic challenges. Antibiotics (Basel). 2026;15:37.
96. Li Y, Kumar S, Zhang L. Mechanisms of antibiotic resistance and developments in therapeutic strategies to combat Klebsiella pneumoniae infection. Infect Drug Resist. 2024;17:1107-19.
97. Sharma R, Yeshwante S, Bui NM, et al. A novel role for colistin as an efflux pump inhibitor in multidrug-resistant Klebsiella pneumoniae. Sci Rep. 2025;15:34612.
98. Devi BT, Aditya V, Kini S, Kumar BK, Deekshit VK. Synergistic effects of antibiotics and efflux pump inhibitors on multidrug-resistant Escherichia coli and Klebsiella pneumoniae. J Appl Microbiol. 2025;136:lxaf169.
99. Huy TXN. Overcoming Klebsiella pneumoniae antibiotic resistance: new insights into mechanisms and drug discovery. Beni-Suef Univ J Basic Appl Sci. 2024;13:13.
100. Filgona J, Banerjee T, Anupurba S. Role of efflux pumps inhibitor in decreasing antibiotic resistance of Klebsiella pneumoniae in a tertiary hospital in North India. J Infect Dev Ctries. 2015;9:815-20.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Copyright
Data & Comments
Data


















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].