Decoding the DHX36-m6A-G4 axis: exploring its role in health and disease through epigenetic and molecular insights
Abstract
G-quadruplexes (G4s) are structurally unique, four-stranded nucleic acid formations composed of guanine-rich sequences stabilized by Hoogsteen hydrogen bonds. These highly stable secondary structures have emerged as vital regulatory elements in DNA and RNA metabolism, where they support genomic stability, telomere integrity, and transcriptional control. Their regulation depends on G4 helicases - specialized enzymes that unwind G4 structures using energy derived from nucleoside triphosphate hydrolysis to ensure proper cellular function. Among them, DEAH-box helicase 36 (DHX36) is a principally essential G4 helicase in human cells, capable of resolving G4s in DNA and RNA. Its activity is critical for preserving genomic integrity and regulating broader cellular processes, underscoring its potential impact on human health and disease. This
Keywords
INTRODUCTION
G-quadruplexes (G4s) are highly stable, four-stranded secondary structures formed by tetrads of guanine-rich sequences in DNA and RNA[1]. These distinctive configurations arise through pairing neighboring guanine bases via Hoogsteen hydrogen bonds, which are further stabilized by monovalent cations positioned within a central G4 cavity[2]. G4s are widely distributed in prokaryotic and eukaryotic genomes, playing essential roles in telomere maintenance, nucleic acid metabolism, and genomic integrity preservation[1]. Numerous studies have notably reported the formation of G4 structures common in the promoter region of key regulatory genes such as MYC proto-oncogene (c-MYC), vascular endothelial growth factor (VEGF), and hypoxia-inducible factor 1 subunit alpha (HIF1A), where they are hypothesized to influence gene expression[3,4]. In mitochondrial DNA, transcription initiates in the D-loop, spanning the entire genome and generating an abundance of guanine-enriched non-coding (nc) RNA transcripts that are thus prone to G4 formation[5]. Emerging studies have indicated that these mitochondrial G4 structures, alongside guanine-rich RNA-binding factor 1 sequences, are believed to participate in post-transcriptional regulation within RNA granules, highlighting their potential intracellular regulatory roles[6].
The stability and widespread presence of G4s have garnered interest due to their role in human health and disease. G4s are linked to various conditions, such as cancer, neurodegenerative diseases, and developmental issues, where they can disrupt normal cell functions. As a result, G4-resolving helicases, such as DEAH-Box Helicase 36 (DHX36), are essential for rebuilding cellular structures and possibly influencing disease pathways related to G4s. When DHX36 interacts with messenger RNA (mRNA) transcripts, it causes structural changes, especially in the 3’-untranslated region (3’ UTR) regions rich in N6-methyladenosine (m6A) modifications and closely linked to YT521-B homology (YTH) N6-methyladenosine RNA binding protein F1 (YTHDF1) binding. These structural shifts may help YTHDF1 bind to m6A sites and promote RNA degradation[7].
m6A proteins regulate RNA through m6A, the most prevalent internal modification in eukaryotic RNA. They are categorized as writers (methyltransferases that add m6A), erasers (demethylases that remove it), and readers (proteins that recognize m6A to modulate RNA behavior)[8,9]. These proteins influence gene expression by affecting splicing, stability, translation, and degradation. Writers catalyze the addition of m6A, while erasers remove it, rendering the process reversible. Readers, including YTH domain proteins (YTHDC1, YTHDC2, and YTHDF1-3), Heterogeneous nuclear ribonucleoproteins (HNRNPs), and Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs), bind to m6A-modified RNA to mediate their effects[10]. They influence RNA from synthesis through degradation, especially concerning stability, translation, splicing, and export. Abnormal m6A modifications and dysregulation of these proteins are associated with diseases such as cancer and viral infections.
G4 helicases are essential for nucleic acid metabolism, mediating processes such as replication, repair, recombination, transcription, and translation[11]. These enzymes aid in actively unwinding G4 structures, positing them as crucial regulators in G4-mediated gene regulation[12,13]. DNA- and RNA-G4s are substrates for members of the Asp-Glu-x-His (DExH) Box resolvase enzyme family, exhibiting high specificity for G4 structures[14]. DHX36 - also known as DDX36, RHAU (RNA helicase associated with AU-rich element), KIAA1488, G4R1, and MLEL1 - has emerged as a central player[15-17]. DHX36 encodes a 1008 amino acid protein with an approximate molecular weight of 114,760 Da, homologous to the Drosophila [maleless (MLE)/MLE-like protein 1 (MLE1)] gene[15]. As with other helicases, DHX36 exhibits adenosine triphosphate (ATP)-dependent G4 unwinding activity in both DNA and RNA. It was initially identified as G4 resolvase-1 due to its specificity for G4 substrates in human cells[12,18]. Structurally, DHX36 was also categorized as a RHAU[19], but later, functional and crystallographic studies further refined its classification to DHX36[20]. These later studies, including those by Chen et al.[21], narrowed their composition to a 440-amino-acid RNA helicase core region and a 105-amino-acid RHAU-specific motif (RSM) that facilitates direct RNA binding[22]. Heddi et al.[23] also described that an 18-amino acid core sequence within the RSM is essential for G4 interaction with G4s.
This comprehensive review will analyze the intricate genetic and epigenetic mechanisms of the DHX36-m6A-G4 axis and its important implications for human health and disease, especially its role in cancer development. The connection between this axis and cancer underscores the urgent need for further research. Moreover, the review aims to offer valuable insights into future research directions for DHX36 and its potential to influence new therapeutic approaches.
G-QUADRUPLEXES IDENTIFICATION AND THEIR SMALL MOLECULE LIGANDS
G-Quadruplexes (G4s) are nucleic acid structures formed by guanine-rich sequences that consist of G-quartets connected by linked loop nucleotides, acting as regulators of nucleic acid metabolism[24,25]. Initially identified in a groundbreaking 1962 study, the formation of G4 tetrameric structures was documented due to their unique composition[26]. Since then, they have become a focus in medicinal chemistry due to their noteworthy structural properties and functional significance in various biological health processes and diseases[27,28]. Each G-tetrad comprises a square-planar arrangement of four guanine bases that form quartets[29] and is stabilized through Hoogsteen-style hydrogen bonding. This facilitates stacking interactions among two or more G-tetrads to form G4s, which are further stabilized by cations such as K+, Na+, and Li+[5,25,30]. Adjacent guanine bases connect via the hydrogen bonds, which occur via the guanine bases’ N1, N7, O6, and N2 atoms. The topological variability of G4s, driven by differences in G-tract orientations and loop arrangements, plays a substantial role in modulating G4-mediated biological functions[30] [Figure 1].
Figure 1. G-quadruplex (G4) structure and topologies. (A) The nucleotide sequence of the G4 motif; (B) Schematic of topologies of G4s.
DNA-G4s: Within the human genome, DNA-G4s are highly prevalent, with early computational analyses estimating an extensive array of potential quadruplex-forming sequences (PQS). The first attempt to compute the number of G4s in the human genome predicted up to 375,000 PQSs via a machine-learning-based algorithm (Huppert and Balasubramanian, 2005). Later, Chambers et al.[31] employed high-throughput G4-sequencing (G4-seq) to analyze purified human DNA with ligand stabilizers, uncovering ~700,000 PQS - an increase of ~350,000 PQS more than prior computational estimates, attributable mainly to non-canonical G4s, such as bulged and long-looped structures, that were previously missed. Moreover, Hänsel-Hertsch et al.[32] employed chromatin immunoprecipitation sequencing (ChIP-seq) with an anti-G4 antibody to map approximately 1,000-10,000 PQS in the human genome. Marsico et al.[33] expanded this mapping to multiple species, revealing that G4 enrichment in gene promoters is distinct from mammals, such as humans and mice.
RNA-G4s: RNA-G4s are similarly enriched in the human transcriptome, forming diverse structural layers despite their single-stranded nature. Using RNA-G4 sequencing (RNA-G4-seq), extensive in vitro maps of G4 structures have been characterized in RNA from HeLa cells[34]. Eukaryotic systems are equipped with robust machinery that globally unfolds RNA-G4s, whereas some bacterial species exhibit a noticeable depletion of G4-forming sequences[35]. Given the single-stranded nature of RNA, RNA-G4s can form intramolecular or intermolecular interactions, establishing two or more stacked layers of G-quartets[36]. They are involved in essential cellular processes, including telomere maintenance and the regulation of gene expression. Additionally, RNA-G4s modulate splicing and polyadenylation in pre-mRNA processing and translation[37], as stated in Tumor Protein P53 (TP53)[38], NRAS Proto-Oncogene (NRAS)[39], the apoptosis regulator Bcl-2 (BCL2)[40], the estrogen receptor-α[41], and membrane-type matrix metalloproteinases[42].
G4s identification: The structure of G4 and its interaction with small molecules have been explored through computational and experimental methods. Several algorithms, including quadruplex forming G-rich sequences (QGRS) Mapper, QuadParser, and G4 Hunter, predict the potential for G4 formation in DNA. Recent tools, including G4Boost, G4ShapePredictor, and precise exploration of nuclear G4s using interpretable neural networks (PENGUINN), utilize machine learning to enhance prediction accuracy by considering sequence composition, structural topology, and thermodynamic stability. Experimental validation typically compares wild-type G4 sequences with their mutated forms, especially G-to-A substitutions. G4Killer, an online tool, illustrates the combination of computational predictions with experimental design, enabling researchers to reduce G4 propensity while maintaining sequence integrity. These advancements underscore the increasing collaboration between bioinformatics and molecular biology in elucidating the complexities of G4 structures. The computational tools for G4 studies are presented in Supplementary Table 1 and have been thoroughly reviewed by Puig Lombardi and Londoño-Vallejo[43], Kikin et al.[44], and Bedrat et al.[45]. Several biophysical and biochemical methods have validated the computational prediction of G4s. Among these techniques are fluorescence resonance energy transfer (FRET), surface plasmon resonance technology, electrophoretic mobility shift assays (EMSA), circular dichroism (CD), and others, as reviewed by Kwok and Merrick[24] and summarized in Supplementary Table 2.
G4s-small molecule ligands: In recent years, the functional relationship between telomeres, oncogenes,
THE GENETIC AND EPIGENETIC ROLES OF THE DHX36-M6A-G4 AXIS IN HEALTH DEVELOPMENT
DHX36 enzymatically unwinds DNA-G4s and RNA-G4s, thereby regulating genomic stability, telomere maintenance, and nucleic acid metabolism[14]. Through its targeting of G4 structures, DHX36 exhibits significant biological potential and is implicated in various aspects of health development, including spermatogenesis[17,47] and the development of skeletal[48,49], cardiovascular[50], and neurological[51] systems.
The DHX36-m6A-G4 axis is a core RNA post-transcriptional regulation system, which is an amalgamation of RNA architecture and RNA adjustment to regulate gene expression. Instead of a set of loosely coupled pieces, this axis works as a molecular switch that has a special helicase that coordinates the structure of the RNA to allow epigenetic control of mRNA fate. The operational principle of this axis is that it is a sequential and interdependent process where the helicase activity of DHX36 is used to open up the structural form of RNA-G4 structures, thus regulating the stability and eventual fate of target mRNAs by determining the accessibility of m6A epigenetic marks to their reader proteins[7].
The process begins with the formation of stable RNA-G4 structures in guanine-rich regions of RNA, which act as physical barriers that block other regulatory factors[52]. These four-stranded structures, especially those found in 3’ UTR regions, can block an access site of possible modifications that would otherwise dictate RNA fate. DHX36 is the specific initiator of the above cascade, and its binding and unwinding of RNA-G4 structures have an unusually high affinity. DHX36 breaks these stable conformations using its ATP-dependent helicase activity, which establishes a critical remodeling of the target RNA that results in an open and accessible conformation. This unwinding effect is most intense in the 3’ UTR region of the mRNAs, as the regulatory elements are often located there.
This structural remodeling leads to the critical interdependence of the DHX36 activity and the m6A-mediated regulation. DHX36 unwinding makes m6A modifications accessible in the lapsed RNA structure, which was previously concealed by the folded RNA structure. The exposure facilitates the binding of YTHDF1, which is a major m6A reader protein, to its target sites with a high affinity. In the absence of unwinding by DHX36, YTHDF1 is unable to identify the m6A marks effectively, so the epigenetic modification remains functionally silent even in the presence of the chemical modification. The authoritative selection of RNA fate depends on the binding of YTHDF1 to the now-accessible m6A site, which mainly facilitates the destruction of the target mRNA by recruiting cellular decaying machinery. Thus, DHX36 becomes a molecular switch, the presence of which leads to remodeling of specific mRNAs and their subsequent degradation by YTHDF1, resulting in a decrease in the level of proteins they encode, which directly affects cellular phenotypes and fate choices[7].
This axis in the normal development and cellular activity is a specific tool of post-transcriptional control in a crucial transition. The DHX36-m6A-G4 pathway allows an individual to quickly regulate the stability of particular mRNAs in response to developmental signals, providing a pathway of reversible and dynamic regulation of genes that complements transcriptional control. DHX36 is critical in maintaining the normal regenerative capacity by regulating transcripts important in myogenic differentiation[53,54]. It is this rapid and reversible control that allows this axis to precisely regulate protein synthesis during stem cell activation, proliferation, and differentiation. Similarly, in embryonic development, structural remodeling of mRNA stability provides cells with an effective means of carrying out intricate gene expression programs, with spatial and temporal specificity. The following sections explore these biological functions in depth, summarizing current research implicated with DHX36.
Reproduction and fertility
Spermatogenesis
Spermatogenesis is a highly complex developmental process essential for male fertility, comprising three key stages: mitosis, meiosis, and spermiogenesis[55]. Initially, spermatogonial stem cells undergo proliferation, leading to the formation of diploid spermatocytes. These cells subsequently enter meiosis I and II, resulting in haploid spermatids, which are ultimately transformed into spermatozoa and stored in the epididymis[17]. It has been previously reported that expression of DHX36 is exceptionally high in the human testis[15], suggesting a pivotal role in regulating spermatogenesis.
Recent studies have revealed the specific functions of DHX36 within this developmental context. Zhang et al.[56] postulated that DHX36 promotes the expression of critical proteins involved in spermatogenesis, including SPO11 initiator of meiotic double strand breaks (SPO11), a key initiator of meiotic double-strand breaks, and synaptonemal complex protein 1 (SYCP1), which is essential for the formation of the synaptonemal complex in spermatocytes by unwinding the G4 motifs in their promoters and coding regions. In parallel, Gao et al.[47] proved that DHX36 may bind to DNA-G4 motifs in the KIT proto-oncogene (c-KIT) promoter, facilitating its expression and promoting spermatogonial differentiation [Figure 2]. These findings collectively highlight the critical role of DHX36 in meiotic initiation and progression, with its absence leading to disrupted spermatogenesis and male infertility.
Figure 2. Proposed role of DHX36 and key regulators in spermatogenesis. (A) Schematic overview of spermatogenesis, progressing from spermatogonia through primary and secondary spermatocytes to spermatids and mature spermatozoa. Arrows indicate developmental progression. Key regulators act at distinct stages: c-KIT supports stem cell maintenance in spermatogonia; SPO11 initiates meiotic recombination in primary spermatocytes; and SYCP1 contributes to synaptonemal complex formation in secondary spermatocytes; (B) Mechanistic representation of DHX36 activity. DHX36 binds to G-quadruplex (G4) structures in m6A-modified RNA, promoting RNA remodeling and facilitating translational regulation during meiosis; (C) Summary of critical regulators involved in spermatogenesis. SPO11 is required for meiotic double-strand breaks and recombination; SYCP1 is essential for synaptonemal complex assembly; and c-KIT mediates proliferation and differentiation of spermatogonial stem cells. Together with DHX36, these regulators coordinate proper meiotic progression and germ cell maturation. DHX36: DEAH-box helicase 36; SYCP1: synaptonemal complex protein 1; c-KIT: KIT proto-oncogene; SPO11: SPO11 initiator of meiotic double strand breaks.
Several genes crucial for spermatogenesis have been identified with significant expression in testicular tissues. These include testis-specific protein Y-linked 4 (TSPY4), leucine zipper protein 4 (LUZP4), anomalous homeobox (ANHX), MAGE family member A4 (MAGEA4), and fibroblast growth factor receptor 3 (FGFR3), as revealed by total RNA sequencing (RNA-seq) analyses conducted by Siebert-Kuss et al.[57]. Further significant genes were recognized in spermatocytes, including ferritin heavy chain-like 17 (FTHL17), Synaptogyrin 4 (SYNGR4), aurora kinase A (AURKA), and ovo-like transcriptional repressor 1 (OVOL1). Other key players in the spermatogenesis process include transition protein 1 (TNP1), protamine 1 (PRM1), high mobility group box 4 (HMGB4), Cation channel sperm associated 3 (CATSPER3), and tektin2 (TEKT2)[17,58].
This section highlights the role of DHX36 in spermatogenesis, particularly in regulating meiotic and spermatogenic genes through G4 interactions, as studied by Zhang et al.[56] and Gao et al.[47]. These studies show DHX36 boosts the expression of SPO11, SYCP1, and c-KIT, but its regulatory mechanisms are not fully understood. RNA-seq identified sperm-related genes such as TSPY4, LUZP4, FTHL17, and TNP1, though DHX36’s influence via G4 structures remains unclear. Its effects on chromatin, transcription, and post-transcriptional processes across spermatogenic stages are poorly known. Future research should map DHX36-G4 interactions with G4-seq and clustered regularly interspaced short palindromic repeats (CRISPR). In vivo models, including DHX36 knockout mice, are crucial for examining stage-specific impacts on meiosis and spermiogenesis, especially AURKA and CATSPER3, related to chromosomal segregation and sperm motility. Multi-omics approaches, including epigenomics and proteomics, are essential for understanding DHX36’s regulatory role. Addressing these gaps will enhance insights into DHX36’s function in spermatogenesis and male reproductive health.
During spermatogenesis, genes such as SPO11, SYCP1, and c-KIT are regulated by m6A at various stages, especially in pachytene, diplotene spermatocytes, and round spermatids. High-throughput methods have identified many m6A-targeted transcripts in male germ cells[59]. Another research issue is whether m6A modifies SPO11, SYCP1, and c-KIT, affecting DHX36’s binding and activity. A thorough investigation is needed.
Female fertility
Oocytes with an intact nuclear membrane are called germinal vesicles, known for active transcription of mRNA and ribosomal RNA (rRNA). As the oocyte grows, transcription declines as chromatin condenses. Typical nucleoli are replaced by atypical nucleolus-like bodies, which decrease during development[60]. The role of G4 in the growth and development of oocytes remains unexamined and requires further investigation. A recent study by Jiao et al.[61] explored the physiological role of DHX36 in oocyte and early embryo development using an oocyte-specific conditional knockout mouse model, along with RNA-seq and Assay for transposase-accessible chromatin using sequencing (ATAC-seq) analyses, as well as fluorescence imaging of G4 probe and anti-G4 antibody. G4 structures have accumulated within chromatin-accessible regions in the mRNA and pre-rRNA transcription of knockout oocytes. This study highlighted the physiological significance of DHX36 in maintaining female fertility, underscoring its critical role in rRNA homeostasis
m6A is vital for oocyte and follicle development. Regulators such as METTL3 aid maturation, prevent DNA damage, and ensure correct chromosome alignment during meiosis. Abnormal m6A can impair development, cause early ovarian aging, and lead to female infertility, emphasizing its importance in reproductive health[63]. Search snippets indicate that METTL3 and DHX36 have distinct roles in RNA regulation, but their direct interaction remains unclear and requires further investigation.
Cardiac development and regeneration
Post-transcriptional regulation of mRNA translation and stability is essential for maintaining proper heart function[64]. This regulation is achieved by involving RNA-binding proteins, particularly G4 helicases, which have been established as key players in modulating gene expression. However, the specific mechanisms by which G4 resolvase activity influences cardiac function remain unexplored[65]. Research by Jiang et al. shows that deleting DHX36 in postnatal mouse cardiomyocytes impairs heart function. They found DHX36 binds to the 5’ and 3’ UTRs of yes1-associated transcriptional regulator (YAP1) and HEXIM P-TEFb complex subunit 1 (HEXIM1), destabilizing these transcripts but increasing their translation[66]. Reduced levels of YAP1 and HEXIM1 due to DHX36 deletion highlight DHX36’s role in postnatal heart development and function.
In addition to its role in mRNA stability and translation, DHX36 regulates ventricular chamber development through a combination of transcriptional and post-transcriptional mechanisms. Huang et al.[50] reported that DHX36 modulates the expression of NK2 homeobox 5 (NKX2-5) and Hes-related family BHLH transcription factor with YRPW motif 2 (HEY2), two critical transcriptional factors in cardiac development. By interacting with the G4 structures within the mRNAs of these genes, DHX36 concomitantly facilitates their expression, thereby contributing to the proper development of ventricular chambers [Figure 3]. The studies above indicate that DHX36 is crucial for postnatal cardiomyocyte function by modulating YAP1 and HEXIM1. YAP1, a key player in the Hippo signaling pathway, plays a significant role in cardiomyocyte growth and repair. The destabilization of its mRNA after DHX36 knockdown reveals a new aspect of post-transcriptional regulation in this pathway. Lower HEXIM1 levels imply DHX36 may affect transcription in cardiomyocytes. It’s unclear whether DHX36’s effects are solely from G4 resolution or involve other RNA-protein interactions. Future research should explore whether pharmacologically targeting DHX36 can improve cardiac repair, especially in ischemic heart disease or heart failure. Studying DHX36 in patient-derived cardiomyocytes or engineered tissues may connect mouse findings with clinical applications.
Skeletal muscle development and regeneration
The development and regeneration of skeletal muscle tissues are intricately linked to muscle stem cells, commonly referred to as satellite cells (SCs), which are associated with myofibers beneath the basal lamina[54]. The myogenesis process relies on the activation of SCs, which can differentiate into new muscle fibers[53]. DHX36 contributes to the activation of the SCs through the regulation of guanine nucleotide-binding protein G(I) subunit alpha-2 (GNAI2) mRNA translation by unwinding its RNA-G4 structures at the 5’ UTR region[48] [Figure 3]. This facilitates the translation of GNAI2, a key downstream effector in activating SCs and skeletal muscle development. This mechanistic insight highlights the regulatory role of DHX36 in maintaining muscle tissue homeostasis.
Zhang et al.[7] investigated the DHX36 binding specificity and its effect on RNA structures using RNA Bind-n-Seq (a high-throughput method for identifying the RNA-binding proteins and qualitatively defining the relative dissociation constants) with the RHAU53 [RNA helicase associated with AU-rich element (RHAU)53] peptide and RNA Structure-seq in wild-type and DHX36-knockout C2C12 mouse myoblast cells. Their findings revealed that DHX36 binding sites are abundant in m6A-modified regions and are associated with YTHDF1 binding. This study highlights how DHX36 binding affects mRNA structures and proposes a mechanism by which changes in RNA secondary structure contribute to post-transcriptional regulation by facilitating YTHDF1 binding. Recently, Zhang et al. introduced a tool that disrupts DHX36’s binding to RNA-G4 using RNA-G4-based proteolytic targeting chimeras. This causes DHX36 degradation via the proteasome, promoting RNA-G4 formation in mRNA. The tool inhibited RNA-G4-mediated amyloid beta precursor protein (APP) expression and affected skeletal muscle stem cell proliferation by downregulating GNAI2, impairing muscle development[67].
In a 2022 study, the same team identified the role of Lockd, a long non-coding RNA (lncRNA), as a significant player in SC activation and muscle regeneration[49]. Lockd enhances the stabilization and binding of DHX36 to eukaryotic translation initiation factor 3 subunit B (EIF3B) protein, forming a complex that promotes the synthesis of acidic nuclear phosphoprotein 32 family member E (ANP32E) protein, a protein essential for myoblast proliferation. This regulatory axis - Lockd/DHX36/ANP32E - promotes myoblast proliferation and skeletal muscle regeneration, underscoring the collaborative interplay between RNA molecules and helicases in promoting muscle regeneration.
DHX36 interacts with RNA entities such as long non-coding RNA (SmaRT) and Spire1 G4 structures. Its depletion reduces Spire1 expression, highlighting its role in promoting skeletal muscle differentiation. lnc-SmaRT can repress Spire1 by base pairing with its G4. Ultimately, DHX36 boosts Spire1 expression, supporting muscle differentiation[68]. These findings highlight DHX36’s multifaceted roles in muscle development and regeneration, emphasizing its potential as a therapeutic target for improving muscle repair.
Neurological development
The regulation of DNA-G4s during learning and memory is crucial in neurobiology. DNA-G4s temporarily accumulate in activated neurons within milliseconds to minutes in response to new experiences[51]. In the neurobiological context of fear extinction - a process in which learned fear responses diminish without reinforcement[69], enabling adaptation to changing environments - G4s play an essential role. Marshall et al.[51] utilized CRISPR-based gene editing in male C57 black 6 (C57/BL6) mice to investigate how DNA-G4s dynamically modulate gene expression in neurons, uncovering a mechanism in which DHX36 helicase is selectively recruited to G4 sites. DNA-G4s accumulate in neurons and dynamically control the activation and repression of genes underlying long-term memory formation; the underlying mechanism of DNA-G4 regulation in the brain involves site-directed deposition of DHX36. They revealed that this targeted recruitment of DHX36 supports timely gene activation and repression linked to long-term memory, showing helicases such as DHX36 can modulate complex cognitive processes. However, the roles of DHX36, and other helicases in neurodevelopment are largely unknown. Because DHX36 impacts gene regulation in memory pathways and its disruption relates to neurological impairments, further research is needed to better understand its role in memory disorders, cognitive issues, and neurodegenerative diseases.
m6A modification is crucial for regulating gene expression and cell differentiation in radial keratinocytes and adult neural stem cells during mammalian brain neurogenesis. It prolongs the cell cycle and delays differentiation by promoting the decay of key mRNAs[70]. m6A also affects self-renewal, differentiation, and lineage decisions of various stem cells. This methylation is enriched in conserved motifs of the aNSC transcriptome, partly supporting neural stem cell proliferation and the development of newborn neurons in the adult brain[71].
While DHX36’s role in resolving G4s for gene regulation linked to learning and memory is recognized, and m6A’s importance in neural stem cell fate is known, their interaction during neurogenesis is largely unexplored. Studying the DHX36-m6A-G4 pathway could reveal new regulatory layers affecting brain development and disorders.
DHX36-m6A-G4 axis in RNA regulation
Research suggests the relationship is a complex interplay, not one-way. An m6A modification near or within a G4 structure may act as a switch to control DHX36’s binding or unwinding activity[72]. This modification could alter the electrostatic landscape or the precise geometry of the G4 structure, making it a more or less optimal substrate for DHX36. Alternatively, m6A reader proteins (such as YTHDF proteins or YTHDC1) could bind first and either recruit or hinder DHX36’s access to the adjacent G4[73]. The second DHX36-m6A-G4 model suggests DHX36 unwinds G4s, changing RNA structure to expose or hide m6A sites and regulate their function. When DHX36 unwinds G4, the RNA becomes accessible [Figure 4]. This can expose a previously hidden m6A site, allowing it to be recognised by reader proteins such as YTHDF1, a well-characterized translational enhancer[7,73]. Functionally, this unwinding simultaneously removes the structural barrier to translation and exposes the m6A site. The m6A site is then bound by YTHDF1, which directly recruits translation initiation factors, leading to the potent activation of protein synthesis. This creates a powerful “on” switch for genes that need rapid and robust expression (e.g., key transcription factors during differentiation, oncogenes in cancer)[74]. If an unwound G4 exposes an m6A site bound by a decay-promoting reader such as YTHDF2 [which recruits the carbon catabolite repression 4 (CCR4)-negative on TATA-less (NOT) complex], it could target mRNA for degradation. The outcome depends on the proteins present and the cellular context. Many oncogenes and pro-survival mRNAs are rich in 5’ UTR G4s and regulated by m6A[75]. The DHX36-m6A axis may be a key mechanism for the dysregulated, high-level expression of specific genes in cancers. This makes the axis a promising therapeutic target.
Figure 4. DHX36-m6A-G-quadruplex (G4) axis. Arrows showed the effect of m6A proteins and their regulation of RNA through m6A, whereas writers (methyltransferases that add m6A), erasers (demethylases that remove it), and readers (proteins that recognize m6A to modulate RNA behavior), which finally regulate G-quadruplex. DHX36: DEAH-box helicase 36.
THE GENETIC AND EPIGENETIC ROLES OF THE DHX36-M6A-G4 AXIS IN DISEASES
The malfunction of DHX36-m6A-G4 results in disease, and a cohesive model can be used to describe how disturbance at the level of DHX36 activity predetermines the appearance of foreseeable effects in a variety of pathological conditions. In the disease conditions associated with the downregulation, mutation, or inactivation of DHX36, the axis is impaired in a way that the RNA-G4 structures on the most important regulating mRNAs remain uncleaved[7,65]. As a result, these mRNAs are abnormally stabilized since YTHDF1 is unable to be recruited to facilitate their degradation, resulting in excessive protein products. It is specifically this dysregulation that is involved in cardiac hypertrophy, whereby DHX36 downregulation results in a pathological gene expression program supporting the disease progression[52]. The identical principle of mechanisms is probably applicable to various settings: any illness that entails mishandling of genes having target RNA-G4 structures in the neighborhood of functional m6A sites might well be caused by or worsened by DHX36 malfunction.
Cancer is especially a compelling environment with respect to this cohesive model of dysregulation. Altered RNA metabolism and post-transcriptional control are common in tumors, and the DHX36-m6A-G4 axis provides a mechanistic description of the manner in which these changes occur[76,77]. In case one of the tumor suppressor mRNAs was a target in this axis, its pathological stabilization under the influence of DHX36 dysfunction may cause unregulated growth and survival of cells. Besides, hyperactivity or overexpression of DHX36 may lead to the overdegradation of tumor suppressors or other proteins that defend the mRNA, which would equally contribute to oncogenesis[78]. The axis thus acts as a vital regulatory node with a status that is critically dependent on precise regulation and deregulation, which is a general route to pathology in various disease states.
Beyond its foundational roles in organ development and function, DHX36 also possesses a pivotal role in the pathology of various diseases, including multiple types of cancers[79,80], viral infections[81-84], tuberculosis (TB)[85], lung edema[86], obesity[87], male infertility, and embryonic lethality[88-90], as well as neurological diseases[91]. In the subsequent sections, we will discuss DHX36’s implications in disease treatment, exploring how its regulatory roles could transform therapies across a spectrum of severe human conditions.
Cancers
Using the GEPIA2 database (http://gepia2.cancer-pku.cn/#index)[92], mRNA expression of DHX36 (ENSG00000174953) across multiple cancer types revealed significantly elevated levels in diffuse large B-cell lymphoma (DLBC), pancreatic adenocarcinoma (PAAD), and thymoma (THYM), with no notable decrease in other cancer types [Figure 5]. Experimental data corroborate this, with multiple studies indicating heightened levels of helicase enzymes in various cancers [Table 1].
Figure 5. The mRNA expression of DHX36 (ENSG00000174953) in 33 different cancer types and the matched normal samples (http://gepia2.cancer-pku.cn/#analysis). T: Tumors tissues; N: normal tissues; BRCA: breast invasive carcinoma; CESC: cervical squamous cell carcinoma; COAD: colon adenocarcinoma; DLBC: diffuse large B-cell lymphoma; GBM: glioblastoma multiforme; KIRC: kidney renal clear cell carcinoma; KIRP: kidney renal papillary cell carcinoma; LIHC: liver hepatocellular carcinoma; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; OV: ovarian serous cystadenocarcinoma; PAAD: pancreatic adenocarcinoma; READ: rectum adenocarcinoma; SKCM: skin cutaneous melanoma; STAD: stomach adenocarcinoma; TGCT: testicular germ cell tumor; THCA: thyroid carcinoma; THYM: thymoma; UCEC: uterine corpus endometrial carcinoma; UCS: uterine carcinosarcoma; DHX36: DEAH-box helicase 36; ACC: adenoid cystic carcinoma; BLCA: bladder cancer; CHOL: cholangiocarcinoma; ESCA: esophageal carcinoma; HNSC: head and neck squamous cell carcinoma; LAML: acute myeloid leukemia; LGG: low-grade gliomas; MESO: mesothelioma; PCPG: pheochromocytoma and paraganglioma; PRAD: prostate adenocarcinoma; SARC: sarcoma; UVM: uveal Melanoma.
Roles of DHX36 in cancer, viral, tuberculosis, male infertility, neurological, and obesity diseases
| Diseases | Types | Results | References |
| Cancer | Testicular | 2; 2’-5’-oligoadenylate synthetase-2 upregulation and testicular cancer repression | [79] |
| Cervical | Cervical cancer progression without mechanistic investigation | [12] | |
| Colon | G-quadruplex-forming sequences containing lncRNA induce colon cancer cell migration by inhibiting DHX36 function | [96] | |
| Lung | DHX36 knockdown promoted tumor growth and colony formation | [97] | |
| Breast | DHX36 knockdown enhanced invasion but decreased migration of MDA-MB-231 and BT549 cells DHX36 knockdown increased tumor growth and cancer cell invasion in the xenograft mouse model | [112] | |
| DHX36 knockdown increased paired-like homeodomain 1, leading to cancer progression | [98] | ||
| DHX36 has an oncogenic effect, enhancing BC progression and invasiveness | [78] | ||
| Head and neck | Overexpression of the DHX36, OPA1 mitochondrial dynamin-like GTPase, and SUMO-specific peptidase-2 genes in HNSC tumors may be involved in the progression of HNSC | [115] | |
| Leukemia | DHX36 is associated with the Aven complex, promoting the survival of leukemic cells | [99] | |
| Expression of numerous transcription factors and oncogenes in response to DHX36 knockout is linked to cancer progression | [80] | ||
| Viruses | Oncolytic viruses | Knocking down MyD88/DHX36 enhanced the production of oncolytic herpes simplex virus in A172 cells, suggesting that DHX36 may improve oncolytic virus efficiency by increasing its yield within tumor cells | [84] |
| Hepatitis C virus | Manoalide inhibits the human DHX36 ATPase by targeting its helicase core, which is shared with HCV NS3, thereby reducing viral replication | [82] | |
| Hepatitis B virus | DHX36 binds strongly to this G4 structure and mutants in the HBV genome, indicating it may influence HBV replication and disease development | [81] | |
| Influenza A virus | DHX36 and eukaryotic translation initiation factor 2 alpha kinase 2 (PKR) interact to form a complex when dsRNA is present, leading to the formation of an antiviral stress granule (avSG) in cells infected with viruses | [83] | |
| The DDX1-DDX21-DHX36 complex functions as a dsRNA sensor for influenza A virus, triggering the TRIF pathway to activate type I interferon responses in the cytoplasm of myeloid dendritic cells | [130] | ||
| Tuberculosis | Bioinformatic analysis of genes revealed high expression of DHX36, BAX, and ARPC1B in TB patients | [85] | |
| Male infertility and embryonic lethality | DHX36 knockdown disrupts normal meiosis progression and hinders spermatogenesis by inducing dysregulated transcription of essential genes, such as SPO11 and SYCP1, which are crucial for chromosome pairing and recombination during meiosis | [56] | |
| DHX36-deficient mice showed significantly smaller testes and a complete absence of mature sperm, leading to a severe impairment in spermatogonial differentiation | [47] | ||
| Neurological diseases | DHX36’s role in regulating the translation of the G4C2 repeat sequence | [91] | |
| DHX36 knockdown in cultured neurons increased the number of neurons with mushroom-shaped spines | [133] | ||
| DHX36 knockdown inhibited brain-derived neurotrophic factor (BDNF), a vital protein for dendritic morphogenesis | [134] | ||
| Obesity | DHX36 unwinds the RNA-G4 structure of miR-26a, thereby enhancing its maturation and biological activity | [87] | |
Cancer involves epigenetic dysregulation beyond genetic mutations. The DHX36-m6A-G4 axis is crucial in post-transcriptional control, affecting cancer gene expression by resolving G4s in promoters, impacting chromatin and transcription[93]. DHX36 unwinds G4s, influencing oncogene silencing or tumor suppressor activation, with m6A modifications modulating this. mRNAs for chromatin modifiers contain G4s and m6A marks; DHX36 exposes m6A sites, guiding reader proteins such as YTHDF1 or YTHDF2, affecting histone and DNA modifier proteins and the epigenetic landscape[94]. G4 structures also regulate lncRNAs, with DHX36 activity influencing their role in recruiting repressive complexes, adding an RNA-driven epigenetic layer[95]. Overall, the DHX36-m6A-G4 axis integrates nucleic acid structures into gene expression, and its dysregulation promotes abnormal transcription in cancer. Targeting it offers a potential epigenetic therapy.
DHX36 generally induced tumor-repressive effects in specific cancers, such as in testicular cancer[79], colon[96], and lung[97] cancers. In other contexts, DHX36 has been demonstrated to facilitate tumor growth, specifically in the proliferation of breast cancer (BC)[98] and leukemia cells[80,99]. Also, cervical cancer recognized DHX36 in HeLa cell lysate without a mechanistic investigation[12]. The following subtitles will examine the roles of DHX36 in different cancer types, elucidating how it influences cancer biology in suppressive and proliferative capacities.
Testicular cancer
Although testicular cancer is relatively rare in the general population, it is one of the most common malignancies among men[100]. The majority of these cases originate from testicular germ cell tumors (TGCTs)[101], which have a strong potential to metastasize to organs such as the retroperitoneal lymph nodes[102], brain[103], and heart[104]. Emerging evidence suggests that some non-coding RNAs, including Piwi-like RNA-mediated gene silencing 1 (PIWI)-interacting RNAs (piRNAs), play key regulatory roles in TGCTs, acting as either tumor suppressors or oncogenes; thus, they can act as potential diagnostic biomarkers and for the development of novel therapies[105]. piRNAs are a class of testis-specific non-coding RNAs that have emerged as possible mediators in the oncogenesis and progression of testicular cancer. This regulatory function opens pathways for their use as diagnostic biomarkers and targets for novel therapies.
Recent studies show piR-36249 interacts with DHX36, promoting its translation and increasing 2’-5’-oligoadenylate synthetase 2 (OAS2) expression. This inhibits testicular cancer cell growth and migration, while promoting apoptosis[79]. In this way, DHX36 can potentially be implicated as a tumor suppressor in testicular cancer, mediating cellular pathways that curb cancer progression.
TGCTs pose a major threat to young men, with studies highlighting piRNAs in tumor development. The interaction between piR-36249 and DHX36 increases OAS2 expression, reducing tumor growth and spread. A key research gap is understanding how DHX36 exerts its tumor-suppressive effects, including other interactions. Future research should focus on in vivo validation, DHX36’s role in different TGCT subtypes, and whether targeting it alone or with treatments could improve outcomes. Developing piRNA-based diagnostics and therapies targeting DHX36 could transform TGCT management, pending further research.
Cervical cancer
Cervical cancer ranks among the top five most common cancers and cancer-related deaths in women worldwide. Approximately 90% of cervical cancers are squamous cell carcinomas, while the remaining 10% are adenocarcinomas[106]. A primary factor in the onset of cervical cancer is human papillomavirus (HPV) infection, a sexually transmitted virus that affects 160 to 289 out of 10,000 persons each year[107]. Regarding the role of DHX36 in cervical cancer, Vaughn et al.[12] observed an elevated level of DHX36 expression in HeLa cells, a commonly used cervical cancer cell line. However, the mechanistic role of DHX36 in cervical cancer progression remains unexplored.
Extensive research has identified pathways such as phosphoinositide 3-kinase (PI3K)/Protein Kinase B (AKT) and Proto-oncogene Int-1 homolog (Wnt)/β-catenin in cervical cancer; however, the role of DHX36 - a helicase linked to G4 resolution and oncogenic regulation - remains unclear. Vaughn et al.[12] noted higher DHX36 expression in HeLa cells, suggesting a link to cervical cancer; however, its specific roles in disease progression have yet to be examined. This indicates a significant research gap, as DHX36’s interactions with G4 structures and its influence in other cancers suggest it may impact oncogenic proteins or signaling pathways in cervical cancer. Future studies should investigate DHX36’s role, including its effects on targets such as transcription factors or cell cycle regulators and interactions with HPV oncoproteins. Exploring DHX36 as a therapeutic target or biomarker may offer new strategies for cervical cancer. Addressing these could improve understanding of cervical cancer biology and aid targeted therapy development.
Colon cancer
Colorectal cancer (CRC) ranks as the second most lethal cancer globally, with projected rises in incidence and mortality in the coming decades[108]. Notably, research has revealed that DHX36 plays a role in CRC progression through its interactions with specific lncRNAs. In a study about this interaction between lncRNAs and DHX36, Matsumura et al.[96] demonstrated that the G-quadruplex-forming sequence-containing lncRNA (GSEC) interacts with DHX36 to inhibit its activity, thereby enhancing colon cancer cell migration. When DHX36 expression was knocked down, the effects of GSEC knockdown on reduced cell migration were reversed, highlighting GSEC’s crucial role in promoting colon cancer cell migration by inhibiting DHX36 function. This study demonstrated that the lncRNA GSEC promotes CRC progression by inhibiting DHX36, thereby facilitating cancer cell migration. This underscores the role of RNA-protein interactions, especially G4 structures, in CRC development. While DHX36’s involvement in migration through GSEC is known, its broader regulatory roles in other oncogenic pathways are unclear. CRC therapies target pathways such as VEGF/vascular endothelial growth factor receptor (VEGFR), epidermal growth factor receptor (EGFR), and Wnt/β-catenin, but how DHX36 interacts with G4 in these pathways remains understudied. Since DHX36 can unwind G4 motifs, it may affect oncogene or tumor suppressor expression. Future research should explore DHX36’s influence on G4 elements in CRC signaling, potentially revealing new therapeutic targets. Combining DHX36 modulation with current treatments could improve outcomes, warranting further studies.
Lung cancer
Lung cancer is a common and deadly cancer worldwide, mainly caused by tobacco smoking. It is frequently diagnosed and is the leading cause of cancer deaths[109]. A bioinformatics study was done by Cui et al.[97], who identified DHX36 as an independent prognostic indicator for non-small cell lung cancer (NSCLC), revealed through the cancer genome atlas (TCGA) and gene expression omnibus (GEO) bioinformatic analysis. They found that knocking down DHX36 enhances tumor growth and colony formation. These findings suggest that DHX36 acts as a tumor suppressor in lung cancer, opening potential avenues for therapeutic targeting. Although bioinformatic analyses highlight DHX36’s promise, experimental validation is essential. Therefore, we recommend further research to evaluate the impact of DHX36 knockdown on these key target pathways, which may help refine strategies in lung cancer management.
The study of Cui et al.[97] highlighted DHX36 as a new prognostic marker and potential tumor suppressor, offering insights into NSCLC. While bioinformatics link DHX36 to tumor suppression, there’s a gap in experimental validation, especially its interactions with VEGF, KRAS proto-oncogene (KRAS), and PI3K/AKT/mechanistic target of rapamycin kinase (mTOR). Future studies should validate DHX36’s mechanisms in vitro and in vivo, and evaluate its potential with current therapies. Exploring DHX36’s role in the tumor microenvironment and immune regulation could improve NSCLC treatment.
Breast cancer
The global burden of BC incidence and mortality continues to increase worldwide because of the growth and aging of the population[110]. By 2040, the burden of BC is expected to rise to over three million new cases and one million deaths every year[111]. Regarding the role of DHX36 in BC, Zeng et al.[112] demonstrated that DHX36 lentiviral knockdown in BC cell lines (MDA-MB-231 and BT549) enhanced their invasion capabilities and disrupted the cell cycle of cells. Besides, DHX36 knockdown synchronized the S-phase cell population and desensitized cells to the cytotoxic effect of cisplatin. In a mouse xenograft tumor model, DHX36 lentiviral knockdown increased tumor growth and invasiveness and decreased cellular apoptosis. These findings highlight DHX36’s role in moderating BC progression.
In a recent study, Lu et al.[62] investigated the link between DHX36 activity and its roles in transcriptional regulation in Michigan cancer foundation-7 (MCF-7) through inducing cleavage under targets and tagmentation (CUT&Tag) for protein-DNA interactions mapping. Subsequently, the genes directly regulated by DHX36 were identified using the auxin-inducible degron protein degradation system and an improved method for sequencing nascent RNA, acrylonitrile-mediated uridine-to-cytidine conversion sequencing (AMUC-seq). The results revealed a significant enrichment of G4 structures at DHX36 target sites, predominantly located in active genomic regions. These findings underscore the potential role of DHX36 in modulating gene transcription through G4 structures. Further insights are provided by Booy et al.[98], who identified a regulatory relationship between DHX36 and the tumor suppressor gene paired-like homeodomain 1 (PITX1) in BC cells. This study revealed that the endogenously expressed DHX36 protein interacts with PITX1 mRNA, modulating its protein expression in BC cells (MCF-7). Decreased DHX36 expression significantly upregulated PITX1, while only marginally altering mRNA levels, suggesting that DHX36 plays a role in controlling BC development. However, given the complexity of DHX36’s regulatory roles, more detailed investigations are crucial to fully elucidate its potential as a therapeutic target in BC.
Recently, our laboratory studied the oncogenic role of DHX36 in BC progression and invasiveness[78]. The research demonstrates that elevated expression of DHX36 is associated with enhanced malignant behaviors in BC cells, including increased proliferation, migration, and invasion. Mechanistically, DHX36 likely exerts its oncogenic effects by regulating the expression of key genes involved in cancer progression, potentially through its known function as a G4 helicase that modulates gene expression at both transcriptional and post-transcriptional levels. These findings suggest that DHX36 could serve as a potential prognostic biomarker and a novel therapeutic target for BC treatment, particularly for aggressive or metastatic forms of the disease.
A significant gap exists in understanding DHX36’s pathways, interactions in oncogenic networks, and its potential as a biomarker or therapeutic target across BC subtypes. Future research should explore its tissue-specific functions, interactions with pathways such as PI3K/AKT/mTOR, and clinical significance in vivo. Examining DHX36’s role in chemotherapy resistance, especially in triple-negative breast cancer (TNBC), could inform new combination therapies. Addressing these gaps is vital to harnessing DHX36’s potential to improve BC treatment and outcomes.
Head and neck cancers
Head and neck cancers (HNC) primarily originate in the squamous cells lining mucosal surfaces of the head and neck, affecting regions such as the mouth, throat, and voice box[113]. Less commonly, HNC may arise in the salivary glands, sinuses, or muscles or nerves in the head and neck[114]. To elucidate the role of DHX36 in HNC, a comprehensive in silico analysis was conducted to identify the gene signature on chromosome 3q. The signature encompassed DHX36, OPA1 mitochondrial dynamin-like GTPase (OPA1), and SUMO-specific peptidase 2 (SENP2), which showed significant correlation in head and neck squamous cell carcinomas (HNSC)[115]. This study offers a promising pathway for understanding HNC pathogenesis, especially in relation to angiogenesis. However, the lack of experimental validation highlights the need for functional studies to clarify the mechanistic roles of EGFR, PI3K/AKT/mTOR, Janus Kinase (JAK)/signal transducer and activator of transcription (STAT), Nuclear Factor Kappa B (NF-κB), and hepatocyte growth factor (HGF)/MET proto-oncogene (MET) in HNC development. Future research should focus on in vitro and in vivo experiments to verify their functions, assess their potential as therapeutic targets, and examine their interactions with established oncogenic pathways.
Leukemia
Leukemia is the most prevalent form of childhood cancer, accounting for 28% of all cases, followed closely by brain and other nervous system tumors[116]. The pathogenesis of leukemia is multifactorial, involving the dysregulation of several essential genes that present viable targets for therapeutic intervention. Recent findings from Thandapani et al.[99] determined the role of Aven in stimulating mRNA translation of the mixed lineage leukemia proto-oncogene, facilitating the transcription of leukemia-associated genes. Crucially, DHX36 has been identified as a component of the Aven complex, which is essential for the efficient translation of G4 mRNAs and promotes the survival of leukemic cells. Studies with DHX36 knockout in Jurkat cells show that losing DHX36 unwinds G4 structures in promoters and genes, increasing transcription factors and oncogenes[80]. Continued research into DHX36’s interactions and regulatory functions in leukemia may yield novel strategies for managing this prevalent childhood cancer.
The discussion of Aven and DHX36 emphasizes their role in supporting leukemic cell survival via mRNA translation and G4 regulation, making them promising targets. However, research gaps exist, particularly regarding their interactions with other oncogenic pathways in vivo and across leukemia subtypes. Future studies should focus on inhibiting DHX36’s G4 unwinding in preclinical models and combining this with current treatments to combat resistance. More research into DHX36’s regulation in leukemogenesis may reveal new biomarkers or drug targets, advancing personalized medicine for pediatric leukemia.
The context-dependent dual role of DHX36 in cancer
DHX36 has a dual role in cancer, which is due to several reasons. First, while DHX36’s core mechanism is G4 unwinding, its ultimate effect is dictated by cellular context and target gene specificity: in scenarios where it primarily resolves G4s in tumor suppressor genes such as PITX1 in BC[98]. Its activity is tumor-suppressive, whereas targeting G4s in oncogenes, as identified in leukemia[80], shifts its role to pro-tumorigenic. Second, the outcome is strongly influenced by the cell’s m6A landscape. Unwinding a G4 to expose an m6A site can have opposing effects - for instance, YTHDF1 binding could enhance translation of an oncogene, while YTHDF2 binding could promote decay of a tumor suppressor mRNA - making the epitranscriptomic state a critical determinant. Finally, DHX36’s interactions with specific cofactors, such as the lncRNA GSEC in colon cancer[96] or the Aven complex in leukemia[99], can modulate its activity and guide target selection, thereby shifting its overall influence toward oncogenic or tumor-suppressive pathways.
Viral infection
The DHX36-m6A-G4 axis adds a layer of epigenetic and post-transcriptional regulation at the host-virus interface. It shows how m6A, the most common mRNA modification, prevents G4 RNA structure formation, affecting their access to the resolvase DHX36[117]. During viral infections, both host and viruses utilize this mechanism: the host regulates G4-containing immune genes, while viruses encode G4 structures in their genomes, where DHX36 influences replication or translation in an m6A-dependent way. For example, a key G4 in the hepatitis B virus (HBV) pre-core promoter, which DHX36 binds strongly[81], suggests that its stability and function are affected by the epitranscriptomic state of viral RNA. Consequently, targeting this axis offers a novel therapeutic approach, with small molecules potentially modulating DHX36 activity or m6A levels to destabilize essential viral G4s or enhance G4-driven antiviral host responses.
In the following paragraphs, we will discuss the role of DHX36 for improving the yield of oncolytic viruses for cancer treatment and its induced antiviral effects against the hepatitis C virus (HCV), the HBV, and the influenza A virus [Table 1 and Figure 6].
Figure 6. Schematic representation of DHX36 involvement in viral infections. DHX36 interacts with multiple viruses and modulates their replication or host responses. In oncolytic viruses, DHX36 downregulates MX2, enhancing herpes simplex virus-1 (HSV-1) yield and contributing to glioblastoma cell killing. In Hepatitis C virus (HCV) infection, DHX36 and viral NS3 promote viral replication. In Hepatitis B virus (HBV), DHX36 shows strong binding affinity to G-quadruplex (G4) structures, though the biological consequences remain to be clarified (needs future investigations). In Influenza A virus (IAV) infection, DHX36 forms a complex with PKR, promoting antiviral stress granule (avSG) formation and enhancing type I interferon (IFN) responses. The red arrow means decrease or downregulation. The green arrow means increase or upregulation. DHX36: DEAH-box helicase 36.
Oncolytic Viruses: Oncolytic viruses have emerged as a promising therapeutic approach for various types of cancer, leveraging their ability to replicate selectively in tumor tissues[118]. The effectiveness of these viruses is attributed to several mechanisms, including the direct lysis of cells, induction of immunological cell death, and activation of both host innate and adaptive immune responses[119]. Various viruses, including herpesviruses, retroviruses, adenoviruses, vaccinia, and poliovirus, have undergone clinical trials to evaluate their safety and efficacy in cancer treatments[120]. Research by Ren et al.[84] highlighted the association between MX dynamin-like GTPase 2 (MX2) protein accumulation and decreased herpes simplex virus-1 (oHSV-1) yield in glioblastoma (A172) cell lines. Interestingly, MX2 depletion resulted in a decrease in mRNA levels of MYD88 (an innate immune signal transduction adaptor), and DHX36 in A172 cells. In addition, MYD88/DHX36 knockdown increased virus yield in A172 cells, suggesting that DHX36 may enhance the effectiveness of oncolytic viruses by increasing their yield within tumor cells.
m6A is an RNA modification affecting herpes simplex virus (oHSV) activity, with HSV-1 infection lowering m6A by degrading methyltransferase 14 (METTL14) via infected cell protein 0 (ICP0). This reduction enhances the anti-tumor effects of oHSV in glioma models[121]. Targeting m6A components such as METTL14 or insulin-like growth factor 2 MRNA binding protein 3 (IGF2BP3) may enhance oHSV cancer therapy. More research on DHX36 and m6A in G4 regulation in oncolytic viruses is suggested for cancer treatment.
Hepatitis C virus: HCV is a hepatotropic RNA virus that can induce acute and chronic hepatitis, often resulting in cirrhosis and hepatocellular carcinoma[122]. Given its high prevalence and chronicity, HCV presents a globally significant public health challenge[123]. HCV’s non-structural protein 3 (NS3 protein) is known to possess nucleoside triphosphatase (NTPase) and RNA helicase activities, which are essential for viral replication. Consequently, NS3 is proposed as an ideal target for the development of antiviral drugs[124]. Research by Salam et al.[82] demonstrated that manoalide could inhibit the ATPase activity of human DHX36, as well as RNA binding and helicase activities of NS3, by targeting the helicase core domain conserved in both HCV NS3 and DHX36, thereby effectively inhibiting viral replication.
Hepatitis B virus: With approximately 250 million people worldwide chronically infected with HBV, this virus is a significant cause of cirrhosis and hepatocellular carcinoma[125]. Although nucleotide analogs and interferons (IFNs) can inhibit HBV replication to alleviate the disease, this does not fully eradicate the virus. Therefore, developing new anti-HBV medicines is essential to control HBV worldwide[126]. Helicases are vital in regulating viral replication and modulating innate immune signaling pathways[127]. Meier-Stephenson et al.[81] reported a unique G4 structure in the pre-core promoter region of the HBV genome, which is crucial for the viral life cycle.
Influenza A virus: Retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs) function as cytoplasmic sensors for viral RNA to initiate antiviral responses, including the production of type I IFN[128]. Influenza A viruses lacking the non-structural protein 1 (NS1) efficiently induce antiviral stress granules (avSG) and increase the production of IFN[129]. Yoo et al.[83] demonstrated that DHX36 is a crucial molecule for RIG-I signaling by regulating the activation of eukaryotic translation initiation factor 2 alpha kinase 2 (PKR) in host cells infected with the influenza A virus ΔNS1 strain. In this context, DHX36 and PKR form a complex in the presence of double-stranded RNA (dsRNA), triggering the formation of an avSG in virus-infected cells. Consequently, DHX36 facilitates the binding of dsRNA and the phosphorylation of PKR through its ATPase/helicase activity. Additionally, Zhang et al.[130] identified the DEAD-box helicase 1 (DDX1)-DEAD-box helicase 21 (DDX21)-DHX36 complex as a dsRNA sensor for influenza A virus, utilizing the TIR domain-containing adaptor-inducing interferon (IFN)-beta (TRIF) pathway to activate type I IFN responses in the cytosol of myeloid dendritic cells.
Tuberculosis
TB is caused by Mycobacterium tuberculosis (M. tuberculosis), which infects approximately one-quarter of the world’s population, of whom 5%-10% develop active disease[131]. The potential epigenetic role of the DHX36-m6A-G4 axis in TB presents a compelling, though still emerging, mechanism by which the host orchestrates its immune response to M. tuberculosis. The observed upregulation of DHX36 in the peripheral blood mononuclear cells of TB patients[85] suggests that its involvement is more than incidental. We can hypothesize that this helicase may function at the nexus of RNA epigenetics and immune signaling, where it could resolve G4 structures in the mRNAs of critical immune genes, such as those within the tumor necrosis factor (TNF) signaling pathway, in a manner regulated by m6A modifications [Table 1].
Male infertility and embryonic lethality
The role of DHX36 in male fertility and embryonic development centers on the DHX36-m6A-G4 epigenetic axis, where it regulates genomic stability and gene expression. DHX36 deficiency leads to infertility marked by meiotic arrest and dysregulated genes such as SPO11 and SYCP1[56]. Further research by Gao et al.[47] revealed the role of DHX36 in spermatogenesis of DHX36-deleted mice. Due to the absence of DHX36 unwinding G4 structures, affected mice exhibited markedly small testes and a lack of mature sperm, resulting in a sharp decrease in spermatogonial differentiation. These findings underscore the pivotal role of DHX36 in regulating the intricate processes of spermatogenesis and its importance in male fertility [Table 1].
Beyond its role in male fertility, DHX36 also plays a critical role in the development of embryonic tissue. Its absence in murine models leads to severe heart abnormalities, primarily due to the insufficient growth and differentiation of heart muscle cells[89]. Jiang et al.[66] reported that DHX36 deficiency is associated with heart failure and mortality rates, coupled with impaired neonatal heart regeneration. Also, comprehensive multi-omics data analyses of cardiovascular disease have revealed that DHX36 expression correlates with reduced incidence of cardiovascular disease[90]. Additionally, Lai et al.[88] identified that DHX36 knockout in the germ line resulted in embryonic lethality, primarily due to the onset of hemolytic anemia and a differentiation defect at the proerythroblast stage. Thus, DHX36 is illustrated to be necessary for hematopoiesis, as evidenced by significant decreases in mice’s red blood cells (RBCs), platelet, and white blood cells (WBCs) counts.
Neurodegenerative diseases
DHX36 is crucial for the proper development and function of the nervous system. This section examines its role in neurological diseases, synaptic plasticity, and potential implications for brain aging [Table 1], revealing that the DHX36-m6A-G4 axis emerges as a critical, yet vulnerable, regulatory mechanism in neurological health, with its dysfunction directly implicated in disease and aging. G4s are implicated in fragile X syndrome, a condition caused by trinucleotide cytosine-guanine-guanine (CGG) expansion that silences the fragile X mental retardation gene[132]. Research by Tseng et al.[91] uncovered DHX36’s role in regulating the translation of the 4 guanine - 2 cytosine (G4C2) repeat sequence. This function opens new avenues for therapeutic use in nucleotide repeat expansion disorders, emphasizing DHX36’s potential in addressing these complex neurological challenges.
DHX36 is also involved in the synaptic plasticity of cultured neurons, an essential process in learning and memory. It regulates the dendritic localization of precursor miRNA-134, which is vital for synaptic functions[133]. DHX36 knockdown in cultured neurons increased the number of neurons with mushroom-shaped spines and an increase in spine volume, indicative of enhanced synaptic connections. This suggests that DHX36 is critical for maintaining the structural integrity of synapses and facilitating neuronal synaptic plasticity. Additionally, DHX36 knockdown inhibited brain-derived neurotrophic factor (BDNF), a vital protein for dendritic morphogenesis[134]. The modulation of BDNF expression further demonstrates the importance of DHX36 in neuronal health and development.
DHX36 may influence brain aging by regulating autophagy genes and DNA repair, potentially contributing to cognitive decline[14]. In addition, Frobel and Hänsel‐Hertsch[135] hypothesized that the acetylation of helicases, including DHX36, increases with age in response to the decline in sirtuin activity. Acetylated helicases bind G4 structures but are less efficient at resolving them, possibly leading to genome instability and brain aging. More research is needed to understand how DHX36 and related helicases can elucidate brain aging mechanisms.
Obesity
Obesity is a complex condition marked by excess body fat, increasing the risk of various diseases and health issues[136]. The rising prevalence of obesity poses a significant public health challenge and has been linked to detrimental effects on conditions such as asthma, affecting its morbidity and therapeutic responses[137]. The role of the DHX36-m6A-G4 axis in obesity highlights a crucial epigenetic mechanism linking nucleic acid structure to metabolic issues. Central to this is the finding that DHX36 is essential for unwinding the G4 structure in the primary transcript of miR-26a, a microRNA whose maturation is significantly impaired in obesity. miR-26a plays a crucial role in various physiological processes and has gained interest as a potential therapeutic target for several human diseases. Notably, miR-26a expression is reduced in obesity, contributing to insulin resistance and metabolic disorders[138]. This reduction plays a pivotal role in the dysregulation of metabolic pathways associated with obesity. Liu et al.[87] demonstrated the importance of RNA-G4s in controlling the biogenesis and function of miR-26a. DHX36 unwinds the RNA-G4 structure of miR-26a, enhancing its maturation and biological activity. This unwinding is crucial for miR-26a’s regulatory functions in metabolism. The interaction between DHX36, RNA G4 structures, and miR-26a forms a key regulatory axis in obesity development. Obese mouse models showed reduced miR-26a maturation and DHX36 expression in the liver [Table 1], highlighting DHX36’s role in regulating miR-26a levels in obesity. Targeting the DHX36/RNA-G4/miR-26a axis could provide new treatments for obesity and metabolic disorders. Modulating DHX36 to boost miR-26a maturation may restore its regulatory role and reduce obesity’s health impacts.
THERAPEUTIC TARGETING OF THE DHX36-M6A-G4 AXIS: OPPORTUNITIES AND CHALLENGES
The most studied group of compounds that act on this axis is G4-stabilizing ligands, which act by binding and stabilizing G4 structures to either inhibit or activate their recognition and resolution by DHX36. Some potential therapies have made their way up to clinical development, including telomestatin, pyridostatin, and BRACO-19, which have strong anticancer properties by stabilizing the G4s in oncogene promoters, including c-MYC and KRAS, resulting in transcriptional repression and cancer cell growth repression[3,39,46]. Markedly, CX-5461 has advanced to phase I/II clinical trials in hematologic malignancies, and quarfloxin (CX-3543) was the first G4-interactive compound to enter clinical trials, but its development was eventually halted because of formulation issues and not due to an efficacy concern[46]. Nonetheless, there are still significant challenges to overcome, especially in selectivity; the human genome has hundreds of thousands of possible G4-forming sequences[31], and it is incredibly hard to selectively stabilize disease-relevant G4s as opposed to other physiologically significant sequences. Genome-wide mapping, G4 enrichment of the regulatory regions of oncogenes and tumor suppressor genes have been reported[32,33], and it is feared that G4 ligands may inadvertently silence protective genes. Also, other compounds, such as TMPyP4, have off-target effects with intercalation of duplex DNA, which may lead to DNA damage[3,46]. The next-generation ligands, such as C066-3108 and {Benzo[lmn][3,8]phenanthroline-1,3,6,8(2H,7H)-tetrone, 2,7-bis[3-(4-morpholinyl)propyl]-4-[[2-(1-pyrrolidinyl)ethyl]amino]} (CM03), which display improved specificity for G4s in oncogene promoters, represent a step toward overcoming these shortcomings[46].
Oncolytic virotherapy takes advantage of the dual capacity of DHX36 in antiviral immunity to increase therapeutic effect on cancers. Transient DHX36 inhibition may enhance oHSV-1 replication in glioblastoma cells because it was found that DHX36 knockdown increases oHSV-1 production[84]. The DDX1-DDX21-DHX36 complex is a sensor of type I IFN stimulation through the TRIF pathway[130]. In oncolytic virotherapy, the antiviral response of this antiviral may restrict viral replication, and a transient DHX36 inhibition might therefore protect the virus against antiviral responses. On the other hand, DHX36 also stimulates the formation of avSGs by activating PKR[83], which can compromise immunogenic cell death and antitumor immunity, resulting in a therapeutic paradox with the net effect determined by the ratio between viral replication and immune activation. The latest finding of degradation of METTL14 by ICP0 through HSV-1 infection, decreasing m6A levels to increase oHSV antitumor effects in glioma[121], indicates that there are complicated interactions between viral infection, m6A modification, and DHX36 activity that need to be comprehended to achieve the best therapeutic combinations. Translation clinical tumor heterogeneity and variable baseline IFN signaling are a challenge to clinical translation and require strategies to stratify patients[139]. Although it is shown that DHX36 modulation combined with oncolytic virotherapy is potentially effective, clinical validation remains unconfirmed. Additionally, timing and levels of DHX36 inhibition will be critical to achieve improved viral replication and sustained antitumor immunity.
The use of proteolysis-targeting chimeras (PROTACs) represents a novel approach for the targeted degradation of DHX36. RNA-G4-based PROTACs, which exploit DHX36’s high-affinity interaction with RNA G-quadruplex structures as the recognition motif, have been developed[67,140]. These bifunctional molecules bind both DHX36 and E3 ubiquitin ligases, leading to ubiquitination and proteasomal degradation, resulting in more complete and sustained effects compared with traditional enzymatic inhibitors. RNA-G4-based PROTACs were effective in C2C12 myoblasts, blocking DHX36-mediated APP expression and inhibiting skeletal muscle stem cell proliferation by downregulating GNAI2. This recapitulated the effects of DHX36 knockout and demonstrated the potential of targeted degradation[67]. Nevertheless, PROTAC technology has several limitations, including high molecular weight and poor pharmacokinetics, which result in low oral bioavailability and limited tissue distribution. This is particularly relevant for DHX36 in muscle or brain, where it performs essential developmental functions[51,52]. There are still issues of selectivity, with other G4-binding proteins (other helicases and hnRNPs) potentially being unintentionally targeted and degraded. The critical role of DHX36 in spermatogenesis[17] and cardiac differentiation[53] raises significant concerns about on-target toxicity from complete DHX36 loss, whereas partial or temporary loss would be preferable but difficult to achieve with current PROTAC designs. Despite these challenges, RNA-G4-based PROTACs provide a powerful tool for studying DHX36 function and represent an innovative, mechanism-based approach that, with further improvements in pharmacokinetics and selectivity, may ultimately yield therapeutic leads.
FUTURE PERSPECTIVES
Although the critical role of the DHX36-m6A-G4 axis is proven, many research opportunities in the future are vital in applying this information to treatment interventions[7]. The future studies ought to focus on the role of m6A modifications around or within G4 structures in regulating the binding and unwinding properties of DHX36. The understanding of this interaction on a molecular basis would result in novel treatment regimens. Creating innovative tools, e.g., RNA-G4-based PROTACs to control DHX36 activity, promises the opportunity to regulate the G4 structures and gene expression precisely in disease models[140]. At the same time, it is essential to enhance computational algorithms, predicting G4 formation, DHX36 binding motifs, and m6A sites, to identify the target correctly and comprehend the functions of the axis in various tissues and diseases. The computational forecasting of interaction between DHX36 and G4 structures is essential to comprehending the strength of interaction, experiment design, and interactions to prioritize further research.
DHX36 is extremely active in the human testis and is key to spermatogenesis. It activates important meiotic genes, such as SPO11 and SYCP1, and binds the c-KIT promoter to activate spermatogonial differentiation[47,56]. Nevertheless, the complete spectrum of genes regulated by DHX36 in spermatogenesis remains to be determined, and additional efforts to determine the mechanistic action of DHX36, particularly in testis-specific developmental genes, are needed. Moreover, the role of DHX36 in female fertility (especially in oogenesis and other pathologies such as premature ovarian failure) is something that needs to be researched urgently because the rate of female infertility is extremely high. Researchers ought to understand the role of DHX36 in regulating the genes of premature ovarian failure, e.g., forkhead box O3 (FOXO3), forkhead box L2 (FOXL2), and bone morphogenetic protein 15 (BMP15).
DHX36 is a key player in postnatal cardiovascular development and functions as it affects YAP1, HEXIM1, and NKX2-5 expression and thus increases ventricular chamber development[66]. It should be further researched to clarify its regulatory impact on genes involved in cardiac development to elucidate the relevant signal pathways, including Wnt/β-catenin signaling. We also promote studies of the regulatory consequences of DHX36 on genes of heart development and electric conduction of impulses, such as atrial natriuretic factor, the gap‐junction protein Connexin 40[141], T‐box transcription factors-5, T‐box transcription factors-20[142], notch receptor 1 (NOTCH1)[143], cytokine bone morphogenetic protein-10[144], and gap junction protein alpha-5[145].
The action of DHX36 in muscle regeneration under aging and muscle illness is a subject that needs to be researched widely to establish new approaches towards the treatment of muscle-wasting illnesses[49]. Moreover, there is only a single study that identified the role of DHX36 when it comes to neural and memory development[66]. The way it contributes to the development of neurons in infants and how it contributes to neurological diseases in adults is unclear. The investigations of DHX36 use in the prevention and treatment of the development of neurological diseases such as Alzheimer’s and Parkinson’s diseases are ongoing.
Cancer is one of the main causes of death, which underscores the necessity to get into the contextual aspects of DHX36[80,96,98,99]. Its tumor-suppressive activity has been observed in certain types of cancers, such as testicular and lung cancer. However, in other cases, such as BC and leukemia, its activity needs to be clarified. Its implication in cancer, such as in hepatocellular carcinoma, cervical carcinoma, osteosarcoma, and colon cancer, remains unknown and thus needs further investigation. Future research ought to investigate the resolvase activity of DHX36 and the most important genes regulated by DHX36, in particular in pathways such as the NOTCH, Hedgehog, PI3K-AKT-mTOR, transforming growth factor beta 1 (TGF-β)-SMAD family member 4 (SMAD4), and insulin-like growth factor 2 (IGF2)-insulin-like growth factor 1 receptor (IGF1R). Also, DHX36 has been demonstrated to upregulate oncolytic production of viruses in tumours, implying that its activity can be controlled to improve virotherapy.
DHX36 is a viral sensor used in the control of viral diseases that identifies nucleic acids. Since it causes serious viral illnesses, there is an urgent need to carry out comprehensive research to come up with specific antiviral treatment[82,83]. The relationship between the action of available antiviral medications on DHX36 and studies on natural compounds with high potency and low acuteness should be investigated to determine whether they can target G4s or regulate the action of DHX36. It is also important to develop sophisticated software to forecast small-molecule affinity to G4s to move forward with drug discovery.
To conclude, the multiple functions of DHX36 in dealing with G4s offer a good opportunity to learn about development and fight illnesses. DHX36 modulators would be a great addition to current treatments, including immune checkpoint inhibitors or antivirals, and yield strong new combination therapies. Multi-disciplinary research is crucial to fully understand the mechanistic roles of DHX36, which could one day be used to produce new treatments for developmental disorders, regenerative medicine, and disease prevention and management.
CONCLUSION
The DHX36-m6A-G4 axis is one of the most significant epigenetic networks when m6A modifications and the G4 structures interact, and DHX36 is the main processor that converts these interactions into particular outcomes in terms of gene expression. It is essential to note that the interaction of DHX36 and m6A plays a vital role in the regulation of G4 structures, which subsequently impacts the formation of tissues and the progression of the disease. In this direction, m6A is labeled on RNA to be processed and has the capacity to affect G4. DHX36 identifies and opens these G4 structures to reveal m6A marks so that these can be used to control the expression of genes in important developmental events.
Genes are essential in cell differentiation during embryonic development, and the DHX36-m6A-G4 axis can do this by facilitating the activation or silencing of genes at the appropriate moments. As an example, such an axis is essential to development, where m6A-directed G4 resolution of DHX36 triggers gene expression required in spermatogenesis (e.g., c-KIT and SPO11), cardiogenesis (e.g., YAP1 and NKX2-5), and neurogenesis.
This axis may be detrimental to the loss or dysfunction in conditions such as cancer. The presence of m6A in tumor suppressor genes and a decrease in DHX36 activity that hinders G4 unwinding are the dysregulation features. It results in repressive G4 structures, which remain and inhibit vital pathways that promote tumor proliferation. The knowledge of this axis provides invaluable clues on the manner in which cells lose control in cancer and other illnesses, and as such, it is an important junction in the developmental as well as the pathological processes involved.
DECLARATIONS
Acknowledgment
The elements in Figure 1, Figure 3, Figure 4 and Graphical Abstract were created with BioRender.com (Created in BioRender. ElFar A (2026) https://app.biorender.com/illustrations/696b076832c5b92b1b269503?slideId=db04ab6b-1085-49bd-ae05-5abd79ea6954).
Authors’ contributions
Conceptualized the review, performed the literature review and data curation, contributed to table creation: El-Far A, Cheng J, Fu L, Li M, Jiang X, Elmetwalli A, Linehan AG, Li HJ, Fu X, Fu J
Wrote the original draft and prepared the figures: El-Far A
All authors contributed to manuscript revision, editing, and have read and approved the final version.
Availability of data and materials
Not applicable.
AI and AI-assisted tools statement
Not applicable.
Financial support and sponsorship
This work was partially supported by the National Natural Science Foundation of China (Grant No. 82073263) and the Joint Innovation Special Project of the Science and Technology Plan of Sichuan Province (Grant No. 2022YFS0623). The New York University Research Catalyst Grant also supported this project.
Conflicts of interest
Fu J is the Guest Editor of the Special Issue “Power of Epigenetics in Tumor Detection, Treatment, and Prognosis” of Journal of Translational Genetics and Genomics and the Section Editor of Journal of Translational Genetics and Genomics. He was not involved in any steps of the editorial process, notably including reviewer selection, manuscript handling, or decision-making. The other authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
Supplementary Materials
REFERENCES
0.
2. Teng X, Hu D, Dai Y, et al. Discovery of a G-quadruplex unwinder that unleashes the translation of G-quadruplex-containing mRNA without inducing DNA damage. Angew Chem Int Ed. 2024;63:e202407353.
3. Siddiqui-Jain A, Grand CL, Bearss DJ, Hurley LH. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc Natl Acad Sci U S A. 2002;99:11593-8.
4. De Armond R, Wood S, Sun D, Hurley LH, Ebbinghaus SW. Evidence for the presence of a guanine quadruplex forming region within a polypurine tract of the hypoxia inducible factor 1alpha promoter. Biochemistry. 2005;44:16341-50.
5. Sahayasheela VJ, Yu Z, Hidaka T, Pandian GN, Sugiyama H. Mitochondria and G-quadruplex evolution: an intertwined relationship. Trends Genet. 2023;39:15-30.
6. Antonicka H, Sasarman F, Nishimura T, Paupe V, Shoubridge EA. The mitochondrial RNA-binding protein GRSF1 localizes to RNA granules and is required for posttranscriptional mitochondrial gene expression. Cell Metab. 2013;17:386-98.
7. Zhang Y, Zhao J, Chen X, et al. DHX36 binding induces RNA structurome remodeling and regulates RNA abundance via m6A reader YTHDF1. Nat Commun. 2024;15:9890.
8. Hong J, Xu K, Lee JH. Biological roles of the RNA m6A modification and its implications in cancer. Exp Mol Med. 2022;54:1822-32.
9. Lu N, Rong G, Han W. Epi-immunotherapy in cancer treatment: mechanisms, clinical progress, and future directions. Cancer Innov. 2025;4:e70023.
10. Chen Y, Zhou Z, Chen Y, Chen D. Reading the m6A-encoded epitranscriptomic information in development and diseases. Cell Biosci. 2024;14:124.
12. Vaughn JP, Creacy SD, Routh ED, et al. The DEXH protein product of the DHX36 gene is the major source of tetramolecular quadruplex G4-DNA resolving activity in HeLa cell lysates. J Biol Chem. 2005;280:38117-20.
13. Dell'Oca MC, Quadri R, Bernini GM, et al. Spotlight on G-quadruplexes: from structure and modulation to physiological and pathological roles. Int J Mol Sci. 2024;25.
14. Antcliff A, McCullough LD, Tsvetkov AS. G-quadruplexes and the DNA/RNA helicase DHX36 in health, disease, and aging. Aging. 2021;13:25578-87.
15. Fu JJ, Li LY, Lu GX. Molecular cloning and characterization of human DDX36 and mouse Ddx36 genes, new members of the DEAD/H box superfamily. Acta Biochim. Biophys Sin. 2002;34:655-61.
16. Fu JJ, Li LY, Liu SF, Xing XW, Liu G, Lu GX. Expression research for human DDX36 and mouse Ddx36 gene in the adult testis. Acta Genet Sin. 2003;30:201-8.
17. Fu L, Wu Q, Fu J. Exploring the biological roles of DHX36, a DNA/RNA G-quadruplex helicase, highlights functions in male infertility: a comprehensive review. Int J Biol Macromol. 2024;268:131811.
18. Creacy SD, Routh ED, Iwamoto F, Nagamine Y, Akman SA, Vaughn JP. G4 resolvase 1 binds both DNA and RNA tetramolecular quadruplex with high affinity and is the major source of tetramolecular quadruplex G4-DNA and G4-RNA resolving activity in HeLa cell lysates. J Biol Chem. 2008;283:34626-34.
19. Tran H, Schilling M, Wirbelauer C, Hess D, Nagamine Y. Facilitation of mRNA deadenylation and decay by the exosome-bound, DExH protein RHAU. Mol Cell. 2004;13:101-11.
20. Newman M, Sfaxi R, Saha A, Monchaud D, Teulade-Fichou MP, Vagner S. The G-quadruplex-specific RNA helicase DHX36 regulates p53 Pre-mRNA 3’-end processing following UV-induced DNA damage. J Mol Biol. 2017;429:3121-31.
21. Chen MC, Tippana R, Demeshkina NA, et al. Structural basis of G-quadruplex unfolding by the DEAH/RHA helicase DHX36. Nature. 2018;558:465-9.
22. Chalupníková K, Lattmann S, Selak N, Iwamoto F, Fujiki Y, Nagamine Y. Recruitment of the RNA helicase RHAU to stress granules via a unique RNA-binding domain. J Biol Chem. 2008;283:35186-98.
23. Heddi B, Cheong VV, Martadinata H, Phan AT. Insights into G-quadruplex specific recognition by the DEAH-box helicase RHAU: solution structure of a peptide-quadruplex complex. Proc Natl Acad Sci U S A. 2015;112:9608-13.
24. Kwok CK, Merrick CJ. G-quadruplexes: prediction, characterization, and biological application. Trends Biotechnol. 2017;35:997-1013.
25. Nakata M, Kosaka N, Kawauchi K, Miyoshi D. Quantitative effects of the loop region on topology, thermodynamics, and cation binding of DNA G-quadruplexes. ACS Omega. 2024;9:35028-36.
26. GELLERT M, LIPSETT MN, DAVIES DR. Helix formation by guanylic acid. Proc Natl Acad Sci U S A. 1962;48:2013-8.
28. Fracchioni G, Vailati S, Grazioli M, Pirota V. Structural unfolding of G-quadruplexes: from small molecules to antisense strategies. Molecules. 2024;29.
29. Shiekh S, Kodikara SG, Balci H. Structure, topology, and stability of multiple G-quadruplexes in long telomeric overhangs. J Mol Biol. 2024;436:168205.
30. Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S. Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res. 2006;34:5402-15.
31. Chambers VS, Marsico G, Boutell JM, Di Antonio M, Smith GP, Balasubramanian S. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat Biotechnol. 2015;33:877-81.
32. Hänsel-Hertsch R, Spiegel J, Marsico G, Tannahill D, Balasubramanian S. Genome-wide mapping of endogenous G-quadruplex DNA structures by chromatin immunoprecipitation and high-throughput sequencing. Nat Protoc. 2018;13:551-64.
33. Marsico G, Chambers VS, Sahakyan AB, et al. Whole genome experimental maps of DNA G-quadruplexes in multiple species. Nucleic Acids Res. 2019;47:3862-74.
34. Kwok CK, Marsico G, Sahakyan AB, Chambers VS, Balasubramanian S. rG4-seq reveals widespread formation of G-quadruplex structures in the human transcriptome. Nat Methods. 2016;13:841-4.
35. Guo JU, Bartel DP. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science. 2016;353:aaf5371.
36. Bugaut A, Balasubramanian S. 5'-UTR RNA G-quadruplexes: translation regulation and targeting. Nucleic Acids Res. 2012;40:4727-41.
37. Millevoi S, Moine H, Vagner S. G-quadruplexes in RNA biology. Wiley Interdiscip Rev RNA. 2012;3:495-507.
38. Decorsière A, Cayrel A, Vagner S, Millevoi S. Essential role for the interaction between hnRNP H/F and a G quadruplex in maintaining p53 pre-mRNA 3’-end processing and function during DNA damage. Genes Dev. 2011;25:220-5.
39. Kumari S, Bugaut A, Huppert JL, Balasubramanian S. An RNA G-quadruplex in the 5’ UTR of the NRAS proto-oncogene modulates translation. Nat Chem Biol. 2007;3:218-21.
40. Shahid R, Bugaut A, Balasubramanian S. The BCL-2 5’ untranslated region contains an RNA G-quadruplex-forming motif that modulates protein expression. Biochemistry. 2010;49:8300-6.
41. Balkwill GD, Derecka K, Garner TP, Hodgman C, Flint AP, Searle MS. Repression of translation of human estrogen receptor alpha by G-quadruplex formation. Biochemistry. 2009;48:11487-95.
42. Morris MJ, Basu S. An unusually stable G-quadruplex within the 5’-UTR of the MT3 matrix metalloproteinase mRNA represses translation in eukaryotic cells. Biochemistry. 2009;48:5313-9.
43. Puig Lombardi E, Londoño-Vallejo A. A guide to computational methods for G-quadruplex prediction. Nucleic Acids Res. 2020;48:1-15.
44. Kikin O, D’Antonio L, Bagga PS. QGRS Mapper: a web-based server for predicting G-quadruplexes in nucleotide sequences. Nucleic Acids Res. 2006;34 Suppl:W676-82.
45. Bedrat A, Lacroix L, Mergny JL. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res. 2016;44:1746-59.
46. Iachettini S, Biroccio A, Zizza P. Therapeutic use of G4-ligands in cancer: state-of-the-art and future perspectives. Pharmaceuticals. 2024;17.
47. Gao X, Ma W, Nie J, et al. A G-quadruplex DNA structure resolvase, RHAU, is essential for spermatogonia differentiation. Cell Death Dis. 2015;6:e1610.
48. Chen X, Yuan J, Xue G, et al. Translational control by DHX36 binding to 5’UTR G-quadruplex is essential for muscle stem-cell regenerative functions. Nat Commun. 2021;12:5043.
49. Chen X, Xue G, Zhao J, et al. Lockd promotes myoblast proliferation and muscle regeneration via binding with DHX36 to facilitate 5’ UTR rG4 unwinding and Anp32e translation. Cell Rep. 2022;39:110927.
50. Huang X, Zhao K, Jiang M, Qiu D, Zhou J, Yang Z. The G4 resolvase RHAU regulates ventricular trabeculation and compaction through transcriptional and post-transcriptional mechanisms. J Biol Chem. 2022;298:101449.
51. Marshall PR, Davies J, Zhao Q, et al. DNA G-quadruplex is a transcriptional control device that regulates memory. J Neurosci. 2024;44:e0093232024.
52. Schultz RJ, Amaral JZ, Bridges CS, et al. Gartland classification concordance of supracondylar fractures among pediatric emergency medicine physicians, radiologists, and orthopedic surgeons. Pediatr Radiol. 2024;54:1180-6.
53. Fu X, Zhang F, Dong X, et al. Adapting cytoskeleton-mitochondria patterning with myocyte differentiation by promyogenic PRR33. Cell Death Differ. 2025;32:177-93.
54. Guo Q, Luo Q, Song G. Control of muscle satellite cell function by specific exercise-induced cytokines and their applications in muscle maintenance. J Cachexia Sarcopenia Muscle. 2024;15:466-76.
56. Zhang K, Zhang T, Zhang Y, et al. DNA/RNA helicase DHX36 is required for late stages of spermatogenesis. J Mol Cell Biol. 2023;14:mjac069.
57. Siebert-Kuss LM, Krenz H, Tekath T, et al. Transcriptome analyses in infertile men reveal germ cell-specific expression and splicing patterns. Life Sci Alliance. 2023;6:e202201633.
58. Johnson ML, Wang R, Sperry AO. Novel localization of aurora a kinase in mouse testis suggests multiple roles in spermatogenesis. Biochem Biophys Res Commun. 2018;503:51-5.
59. Gui Y, Yuan S. Epigenetic regulations in mammalian spermatogenesis: RNA-m6A modification and beyond. Cell Mol Life Sci. 2021;78:4893-905.
60. Fulka H, Langerova A. Nucleoli in embryos: a central structural platform for embryonic chromatin remodeling? Chromosome Res. 2019;27:129-40.
61. Jiao YX, Bu GW, Wu YW, et al. DHX36-mediated G-quadruplexes unwinding is essential for oocyte and early embryo development in mice. Sci Bull. 2025;70:1295-310.
62. Lu QW, Liu SY, Liao XQ, et al. RNA G-quadruplex removal promotes a translational switch after meiosis resumption. Nucleic Acids Res. 2025;53:gkaf067.
63. Wu D, Liu C, Ding L. Follicular metabolic dysfunction, oocyte aneuploidy and ovarian aging: a review. J Ovarian Res. 2025;18:53.
64. Völkers M, Preiss T, Hentze MW. RNA-binding proteins in cardiovascular biology and disease: the beat goes on. Nat Rev Cardiol. 2024;21:361-78.
65. Yang C, Yao J, Yi H, Huang X, Zhao W, Yang Z. To unwind the biological knots: the DNA/RNA G-quadruplex resolvase RHAU (DHX36) in development and disease. Animal Model Exp Med. 2022;5:542-9.
66. Jiang M, Hu H, Zhao K, et al. The G4 resolvase RHAU modulates mRNA translation and stability to sustain postnatal heart function and regeneration. J Biol Chem. 2021;296:100080.
67. Zhang K, Nie Q, Li M, et al. RNA G-quadruplex structure-based PROTACs for targeted DHX36 protein degradation and gene activity modulation in mammalian cells. Nucleic Acids Res. 2025;53:gkaf039.
68. Scalzitti S, Mariani D, Setti A, et al. Lnc-SMaRT translational regulation of spire1, a new player in muscle differentiation. J Mol Biol. 2022;434:167384.
69. Marshall PR, Bredy TW. What does the future hold for the study of nucleic acid modifications in the brain? In: Bredy TW, Editor. DNA Modifications in the Brain. Academic Press; 2017. pp. 149-59.
70. Fan Y, Lv X, Chen Z, Peng Y, Zhang M. m6A methylation: critical roles in aging and neurological diseases. Front Mol Neurosci. 2023;16:1102147.
71. Chen J, Zhang YC, Huang C, et al. m6A regulates neurogenesis and neuronal development by modulating histone methyltransferase Ezh2. Genom Proteom Bioinform. 2019;17:154-68.
72. Fleming AM, Nguyen NLB, Burrows CJ. Colocalization of m6A and G-quadruplex-forming sequences in viral RNA (HIV, Zika, Hepatitis B, and SV40) suggests topological control of adenosine N6-methylation. ACS Cent Sci. 2019;5:218-28.
73. Zhen D, Wu Y, Zhang Y, et al. m6A reader: epitranscriptome target prediction and functional characterization of N6-methyladenosine (m6A) readers. Front Cell Dev Biol. 2020;8:741.
74. Li F, Li W. Readers of RNA modification in cancer and their anticancer inhibitors. Biomolecules. 2024;14:881.
75. Fitzsimmons CM, Batista PJ. It’s complicated… m6A-dependent regulation of gene expression in cancer. Biochim Biophys Acta Gene Regul Mech. 2019;1862:382-93.
76. Chen L, Gao Y, Xu S, et al. N6-methyladenosine reader YTHDF family in biological processes: structures, roles, and mechanisms. Front Immunol. 2023;14:1162607.
77. Mizumoto A, Yokoyama Y, Miyoshi T, Takikawa M, Ishikawa F, Sadaie M. DHX36 maintains genomic integrity by unwinding G-quadruplexes. Genes Cells. 2023;28:694-708.
78. Wei C, Xu D, Cheng J, et al. DHX36 plays an oncogenic role in breast cancer progression and invasiveness. Sci Rep. 2025;16:1156.
79. Wang Q, Chen P, Wang X, et al. piR-36249 and DHX36 together inhibit testicular cancer cells progression by upregulating OAS2. Noncoding RNA Res. 2023;8:174-86.
80. Gumina JM, Richardson AE, Shojiv MH, et al. Differential gene expression following DHX36/G4R1 knockout is associated with G-quadruplex content and cancer. Int J Mol Sci. 2024;25:1753.
81. Meier-Stephenson V, Badmalia MD, Mrozowich T, et al. Identification and characterization of a G-quadruplex structure in the pre-core promoter region of hepatitis B virus covalently closed circular DNA. J Biol Chem. 2021;296:100589.
82. Salam KA, Furuta A, Noda N, et al. Inhibition of hepatitis C virus NS3 helicase by manoalide. J Nat Prod. 2012;75:650-4.
83. Yoo JS, Takahasi K, Ng CS, et al. DHX36 enhances RIG-I signaling by facilitating PKR-mediated antiviral stress granule formation. PLoS Pathog. 2014;10:e1004012.
84. Ren Y, Chen M, Wu G, et al. High accumulation of Mx2 renders limited multiplication of oncolytic herpes simplex virus-1 in human tumor cells. Sci Rep. 2021;11:21227.
85. Zhang Y, Li Y, Li H, et al. DHX36, BAX, and ARPC1B may be critical for the diagnosis and treatment of tuberculosis. Can Respir J. 2020;2020:4348371.
86. Migneault F, Gagnon F, Pascariu M, et al. Post-transcriptional modulation of aENaC mRNA in alveolar epithelial cells: involvement of its 3’ untranslated region. Cell Physiol Biochem. 2019;52:984-1002.
87. Liu G, Du W, Xu H, et al. RNA G-quadruplex regulates microRNA-26a biogenesis and function. J Hepatol. 2020;73:371-82.
88. Lai JC, Ponti S, Pan D, et al. The DEAH-box helicase RHAU is an essential gene and critical for mouse hematopoiesis. Blood. 2012;119:4291-300.
89. Nie J, Jiang M, Zhang X, et al. Post-transcriptional regulation of Nkx2-5 by RHAU in heart development. Cell Rep. 2015;13:723-32.
90. Nikpay M, Soubeyrand S, Tahmasbi R, McPherson R. Multiomics Screening Identifies Molecular Biomarkers Causally Associated With the Risk of Coronary Artery Disease. Circ Genom Precis Med. 2020;13:e002876.
91. Tseng YJ, Sandwith SN, Green KM, et al. The RNA helicase DHX36-G4R1 modulates C9orf72 GGGGCC hexanucleotide repeat-associated translation. J Biol Chem. 2021;297:100914.
92. Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47:W556-60.
93. He PC, He C. m6A RNA methylation: from mechanisms to therapeutic potential. EMBO J. 2021;40:e105977.
94. Zaccara S, Jaffrey SR. A unified model for the function of YTHDF proteins in regulating m6A-modified mRNA. Cell. 2020;181:1582-1595.e18.
96. Matsumura K, Kawasaki Y, Miyamoto M, et al. The novel G-quadruplex-containing long non-coding RNA GSEC antagonizes DHX36 and modulates colon cancer cell migration. Oncogene. 2017;36:1191-9.
97. Cui Y, Li Z, Cao J, et al. The G4 Resolvase DHX36 possesses a prognosis significance and exerts tumour suppressing function through multiple causal regulations in non-small cell lung cancer. Front Oncol. 2021;11:655757.
98. Booy EP, Howard R, Marushchak O, et al. The RNA helicase RHAU (DHX36) suppresses expression of the transcription factor PITX1. Nucleic Acids Res. 2014;42:3346-61.
99. Thandapani P, Song J, Gandin V, et al. Aven recognition of RNA G-quadruplexes regulates translation of the mixed lineage leukemia protooncogenes. Elife. 2015;4:e06234.
100. Yuen KL, Pandit K, Puri D, Yodkhunnatham N, Bagrodia A. Testicular cancer with small metastatic burden: optimal approach in 2024. Curr Opin Urol. 2024;34:204-9.
101. Islam R, Heyer J, Figura M, et al. T cells in testicular germ cell tumors: new evidence of fundamental contributions by rare subsets. Br J Cancer. 2024;130:1893-903.
102. Tarrant WP, Czerniak BA, Guo CC. Relationship between primary and metastatic testicular germ cell tumors: a clinicopathologic analysis of 100 cases. Hum Pathol. 2013;44:2220-6.
103. Iida K, Naiki T, Kawai N, et al. Metastasectomy as optimal treatment for late relapsing solitary brain metastasis from testicular germ cell tumor: a case report. BMC Res Notes. 2014;7:865.
104. Rascol O, Ferreira J, Nègre-Pages L, et al. A proof-of-concept, randomized, placebo-controlled, multiple cross-overs (n-of-1) study of naftazone in Parkinson’s disease. Fundam Clin Pharmacol. 2012;26:557-64.
105. Calin GA, Croce CM. MicroRNA-cancer connection: the beginning of a new tale. Cancer Res. 2006;66:7390-4.
106. Aimagambetova G, Terzic M, Marat A, et al. Cervical cancer natural history, diagnosis, and treatment: from molecular events to clinical management. In: Rezaei N, Editors. gynecological cancers: an interdisciplinary approach. Springer, Cham; 2024. pp. 77-116.
108. Hossain MS, Karuniawati H, Jairoun AA, et al. Colorectal cancer: a review of carcinogenesis, global epidemiology, current challenges, risk factors, preventive and treatment strategies. Cancers. 2022;14:1732.
109. Marshall HM, Fong KM. Lung cancer screening - time for an update? Lung Cancer. 2024;196:107956.
110. Sedeta ET, Jobre B, Avezbakiyev B. Breast cancer: global patterns of incidence, mortality, and trends. J Clin Oncol. 2023;41:10528.
111. Arnold M, Morgan E, Rumgay H, et al. Current and future burden of breast cancer: global statistics for 2020 and 2040. Breast. 2022;66:15-23.
112. Zeng Y, Qin T, Flamini V, et al. Identification of DHX36 as a tumour suppressor through modulating the activities of the stress-associated proteins and cyclin-dependent kinases in breast cancer. Am J Cancer Res. 2020;10:4211-33.
113. Barsouk A, Aluru JS, Rawla P, Saginala K, Barsouk A. Epidemiology, risk factors, and prevention of head and neck squamous cell carcinoma. Med Sci. 2023;11:42.
114. Son E, Panwar A, Mosher CH, Lydiatt D. Cancers of the major salivary gland. J Oncol Pract. 2018;14:99-108.
115. Karatas OF, Capik O, Barlak N, Aydin Karatas E. Comprehensive in silico analysis for identification of novel candidate target genes, including DHX36, OPA1, and SENP2, located on chromosome 3q in head and neck cancers. Head Neck. 2021;43:288-302.
117. Chen T, Greene GH, Motley J, et al. m6A modification plays an integral role in mRNA stability and translation during pattern-triggered immunity. Proc Natl Acad Sci U S A. 2024;121:e2411100121.
118. Lin D, Shen Y, Liang T. Oncolytic virotherapy: basic principles, recent advances and future directions. Signal Transduct Target Ther. 2023;8:156.
120. Kaufman HL, Kohlhapp FJ, Zloza A. Oncolytic viruses: a new class of immunotherapy drugs. Nat Rev Drug Discov. 2015;14:642-62.
121. Dai W, Tian R, Yu L, et al. Overcoming therapeutic resistance in oncolytic herpes virotherapy by targeting IGF2BP3-induced NETosis in malignant glioma. Nat Commun. 2024;15:131.
123. Mundim AEFM, de Castro FOF, Albuquerque MBB, Vilanova-Costa CAST, Pfrimer IAH, Silva AMTC. Major mutations in the NS3 gene region of hepatitis C virus related to the resistance to direct acting antiviral drugs: a systematic review. Virusdisease. 2020;31:220-8.
124. Frick DN. The hepatitis C virus NS3 protein: a model RNA helicase and potential drug target. Curr Issues Mol Biol. 2007;9:1.
125. Kew MC. Epidemiology of chronic hepatitis B virus infection, hepatocellular carcinoma, and hepatitis B virus-induced hepatocellular carcinoma. Pathol Biol. 2010;58:273-7.
126. Yu X, Gao Y, Zhang X, et al. Hepatitis B: model systems and therapeutic approaches. J Immunol Res. 2024;2024:4722047.
127. You H, Ma L, Wang X, et al. The emerging role of DEAD/H-box helicases in hepatitis B virus infection. Front Cell Infect Microbiol. 2022;12:1062553.
128. Xu XX, Wan H, Nie L, Shao T, Xiang LX, Shao JZ. RIG-I: a multifunctional protein beyond a pattern recognition receptor. Protein Cell. 2018;9:246-53.
129. Onomoto K, Jogi M, Yoo JS, et al. Critical role of an antiviral stress granule containing RIG-I and PKR in viral detection and innate immunity. PLoS ONE. 2012;7:e43031.
130. Zhang Z, Kim T, Bao M, et al. DDX1, DDX21, and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity. 2011;34:866-78.
131. Vasiliu A, Martinez L, Gupta RK, et al. Tuberculosis prevention: current strategies and future directions. Clin Microbiol Infect. 2024;30:1123-30.
132. Stefanovic S, DeMarco BA, Underwood A, Williams KR, Bassell GJ, Mihailescu MR. Fragile X mental retardation protein interactions with a G quadruplex structure in the 3’-untranslated region of NR2B mRNA. Mol Biosyst. 2015;11:3222-30.
133. Bicker S, Khudayberdiev S, Weiß K, Zocher K, Baumeister S, Schratt G. The DEAH-box helicase DHX36 mediates dendritic localization of the neuronal precursor-microRNA-134. Genes Dev. 2013;27:991-6.
134. Zampa F, Bicker S, Schratt G. Activity-dependent Pre-miR-134 dendritic localization is required for hippocampal neuron dendritogenesis. Front Mol Neurosci. 2018;11:171.
135. Frobel J, Hänsel-Hertsch R. The age-related decline of helicase function-how G-quadruplex structures promote genome instability. FEBS Lett. 2025;599:267-74.
137. Bloodworth MH, Staso PJ, Huang S, et al. Impact of metabolic and weight components on incident asthma using a real-world cohort. Ann Allergy Asthma Immunol. 2024;133:660-666.e5.
138. Fu X, Dong B, Tian Y, et al. MicroRNA-26a regulates insulin sensitivity and metabolism of glucose and lipids. J Clin Invest. 2015;125:2497-509.
139. Appleton E, Chiocca EA, Ungerechts G, Melcher A, Vile R. Oncolytic viruses as anticancer agents: clinical progress and remaining challenges. Lancet. 2025;406:1295-312.
140. Patil KM, Chin D, Seah HL, Shi Q, Lim KW, Phan AT. G4-PROTAC: targeted degradation of a G-quadruplex binding protein. Chem Commun. 2021;57:12816-9.
141. Houweling AC, Somi S, Van Den Hoff MJ, Moorman AF, Christoffels VM. Developmental pattern of ANF gene expression reveals a strict localization of cardiac chamber formation in chicken. Anat Rec. 2002;266:93-102.
142. Greulich F, Rudat C, Kispert A. Mechanisms of T-box gene function in the developing heart. Cardiovasc Res. 2011;91:212-22.
143. Del Monte-Nieto G, Ramialison M, Adam AAS, et al. Control of cardiac jelly dynamics by NOTCH1 and NRG1 defines the building plan for trabeculation. Nature. 2018;557:439-45.
144. Chen H, Shi S, Acosta L, et al. BMP10 is essential for maintaining cardiac growth during murine cardiogenesis. Development. 2004;131:2219-31.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data






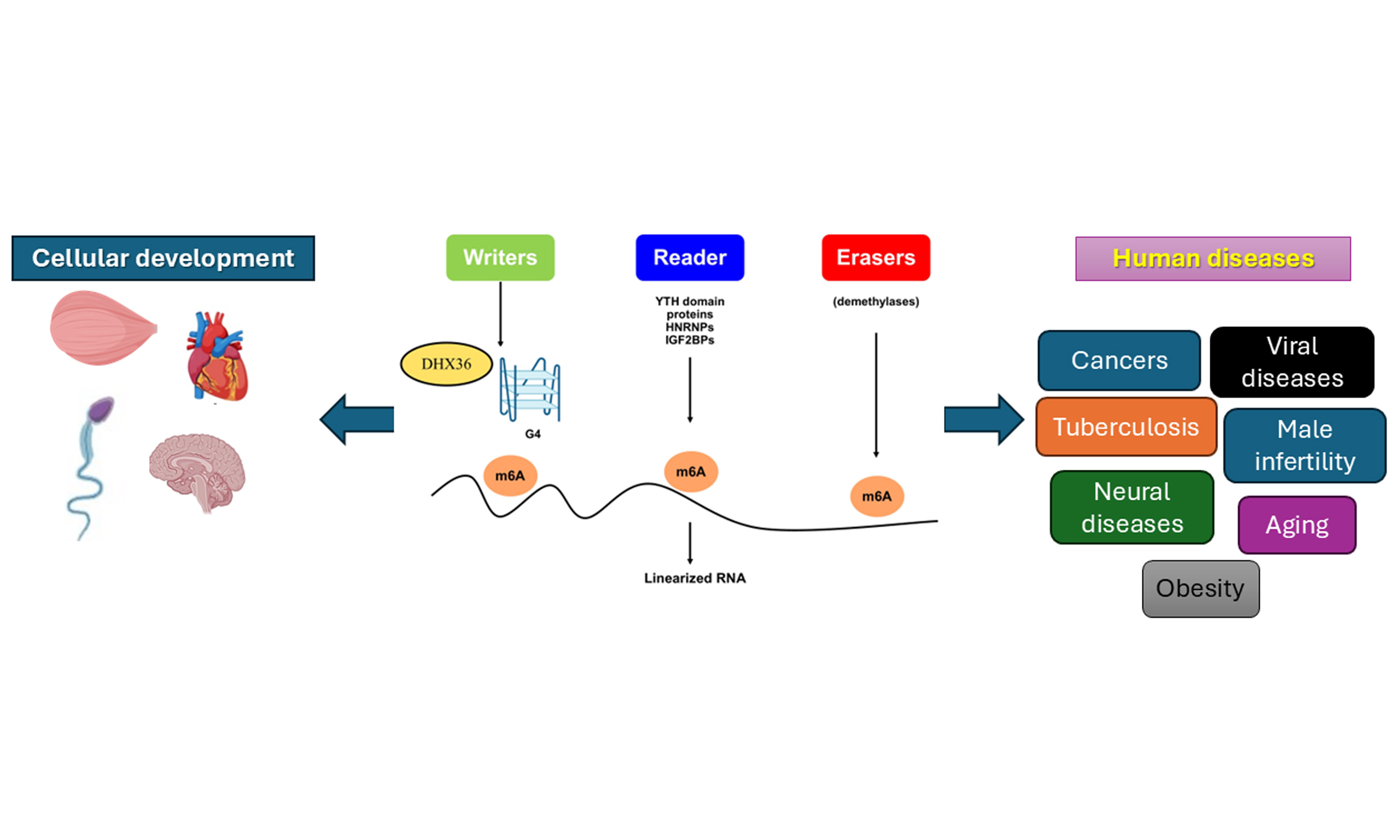
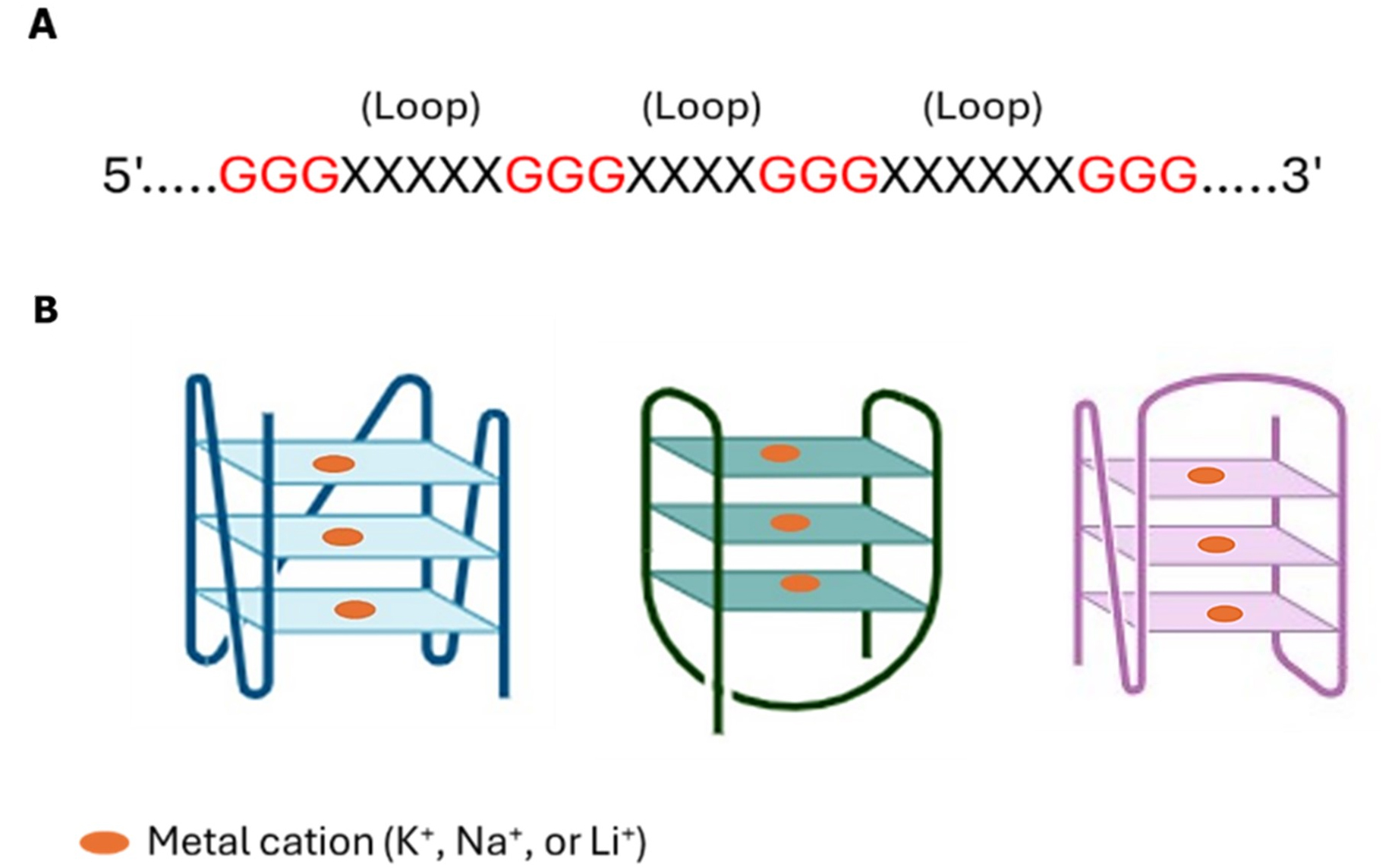
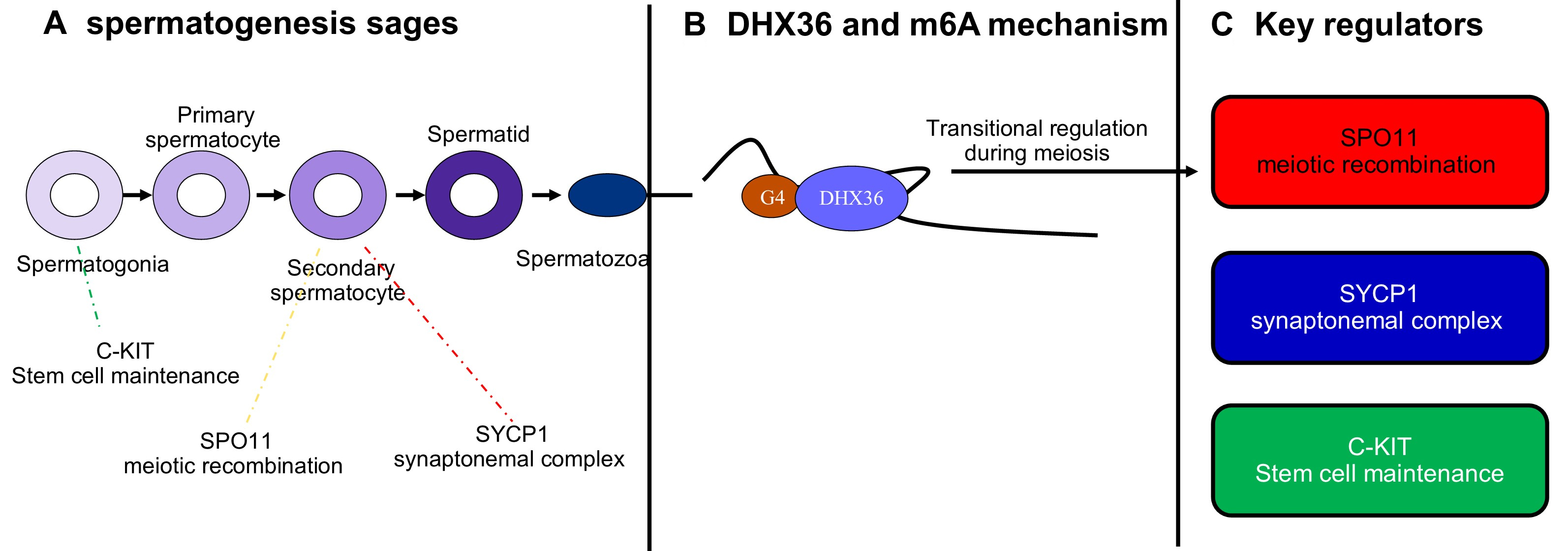
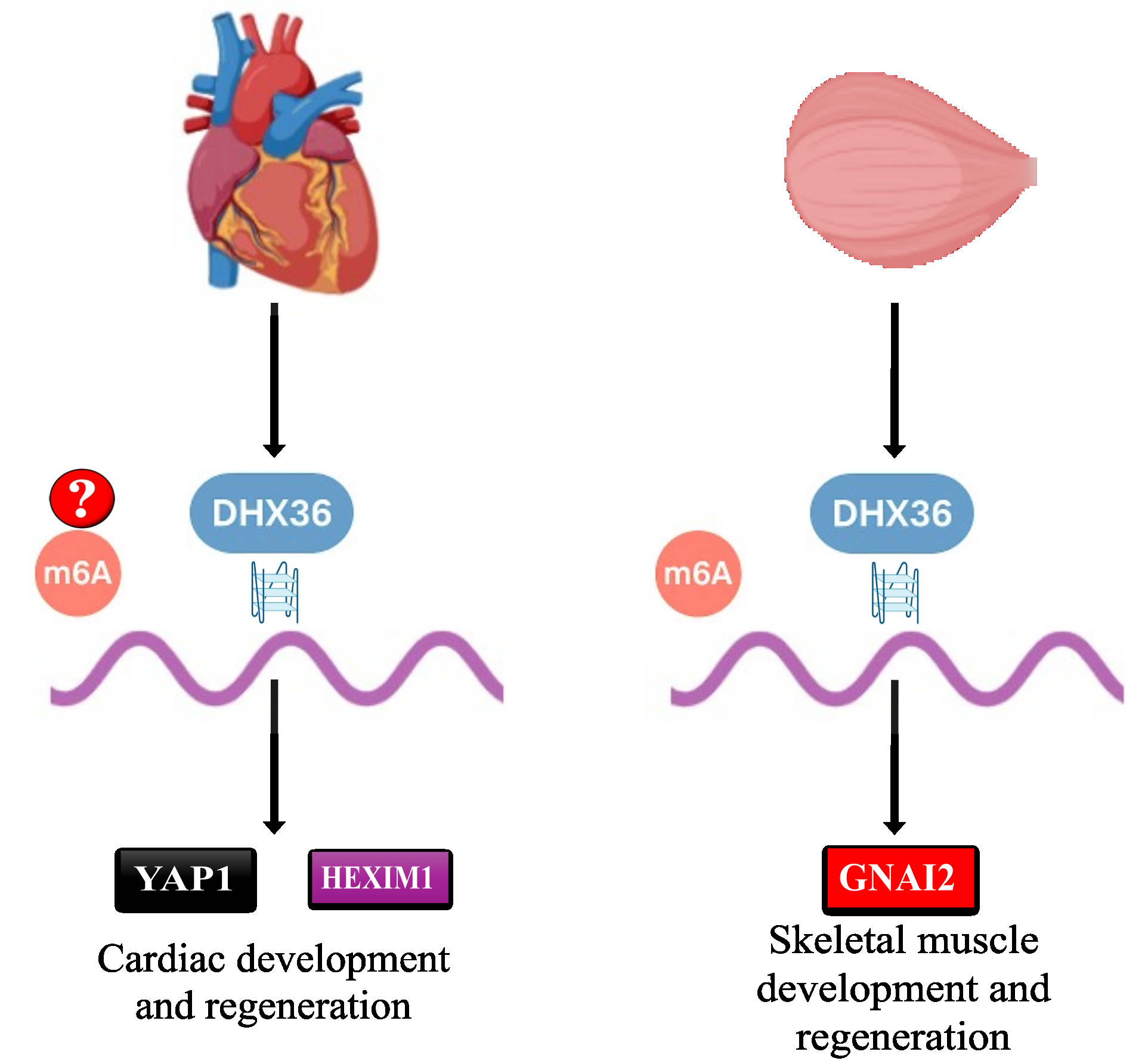
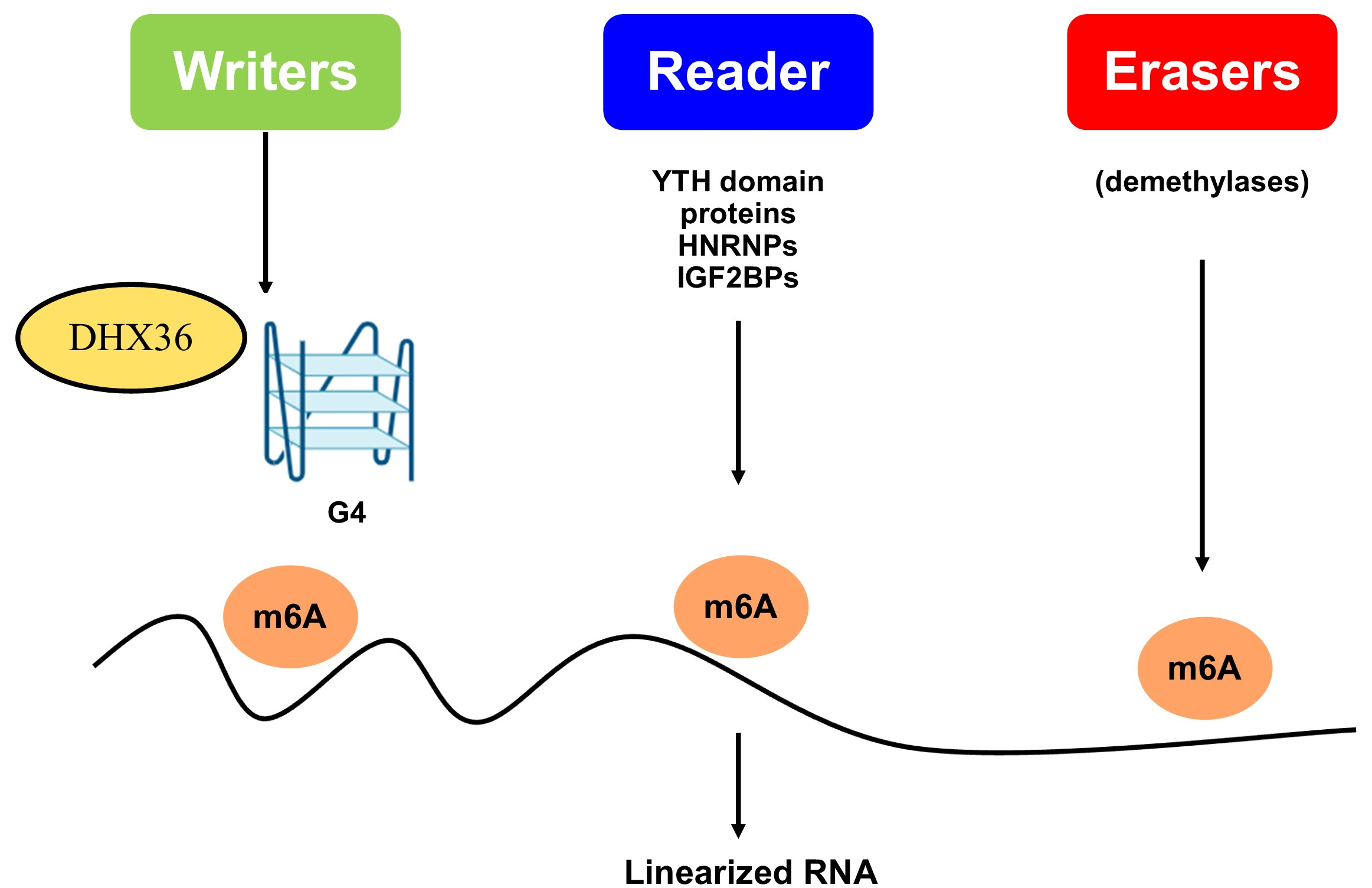
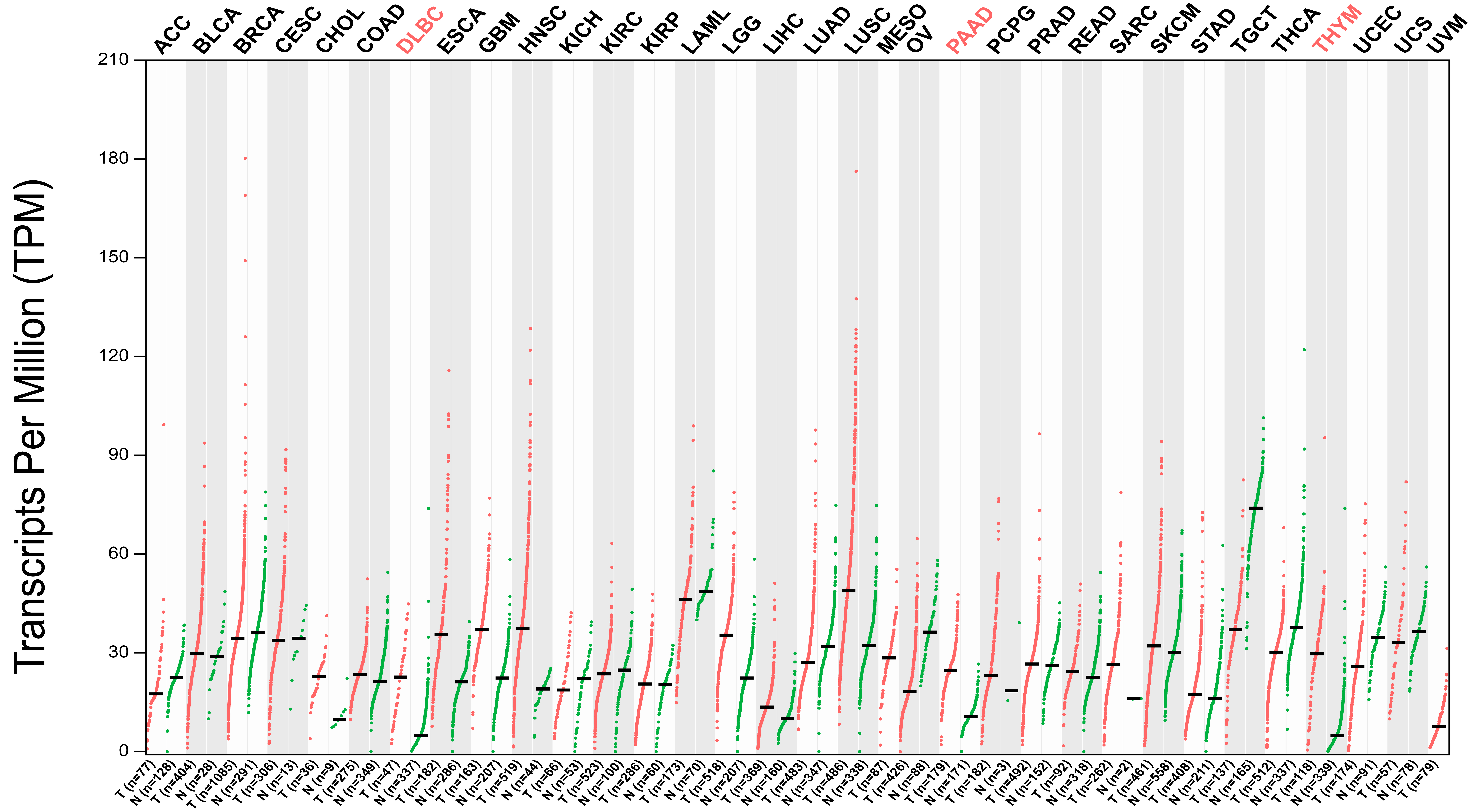
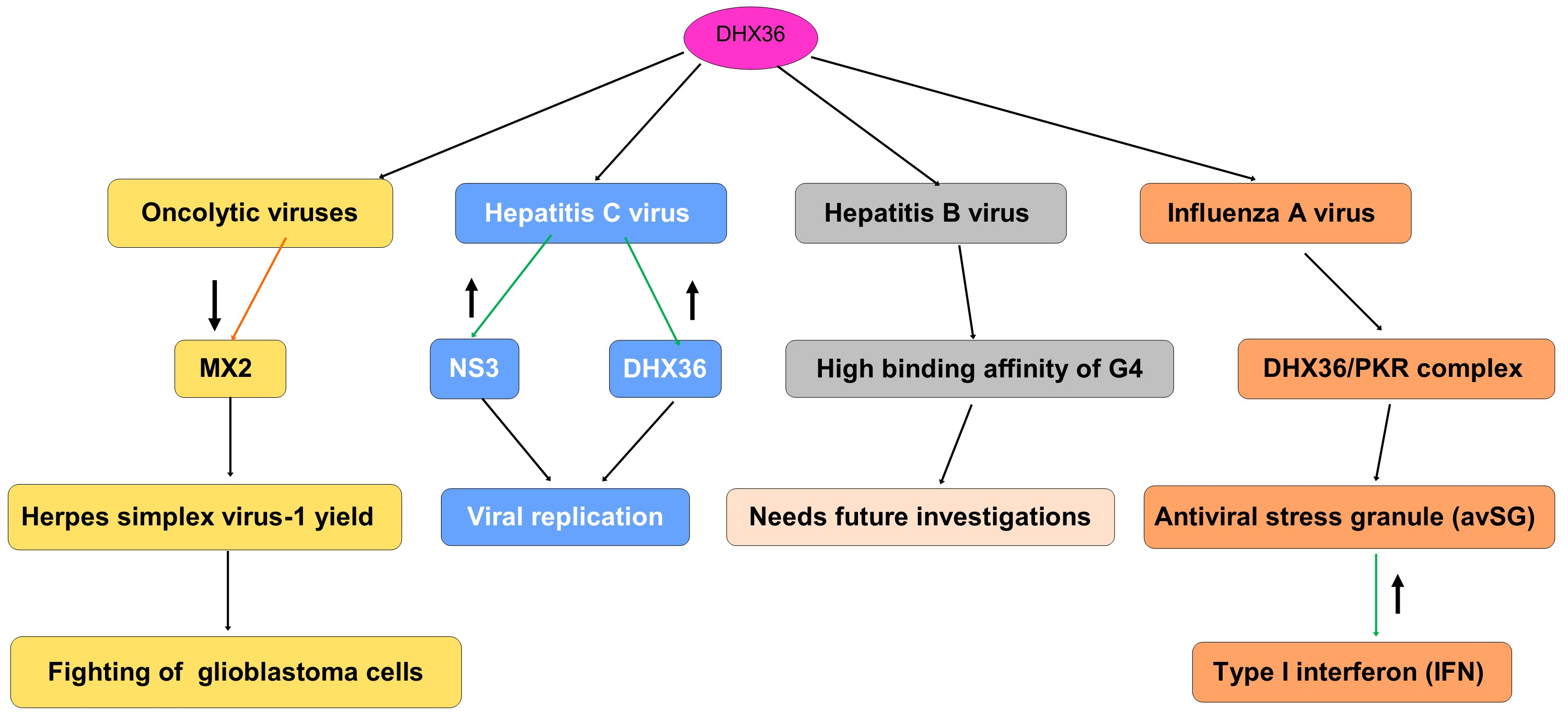












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].