fig4
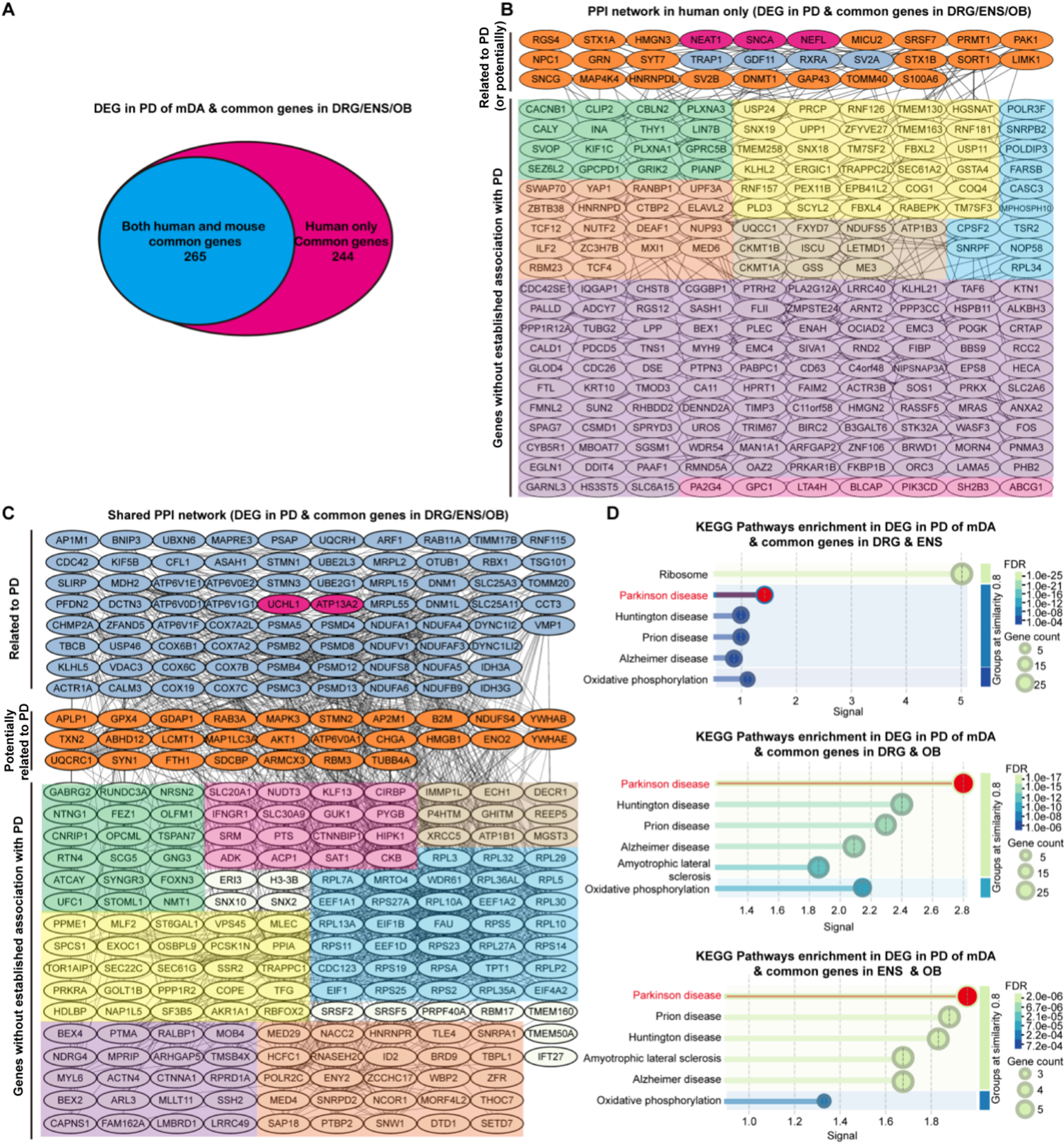
Figure 4. Cross-species comorbidity gene overlap and functional enrichment analysis. (A) Venn diagram showing the overlap between comorbidity-associated genes identified in humans only (244 genes) and those conserved across both human and mouse datasets (265 genes); (B) PPI network of comorbidity genes identified only in humans; (C) PPI network of comorbidity genes shared between human and mouse. Divided into three clusters: Related to PD, Potentially related to PD and Genes without established association with PD. Note: This classification is based on the literature and serves as a hypothesis-generating framework rather than a definitive annotation. “Genes without established association with PD” cluster are further grouped into seven functional submodules based on biological relevance: neural development and synapses (green), autophagy and protein clearance transport (yellow), mitochondrial function and oxidative stress (brown), inflammation and immune metabolism (pink), translation and ribosome/protein synthesis (blue), basic cellular functions and cytoskeleton (purple), and RNA processing/splicing/nuclear export and transcriptional regulation(orange); (D) KEGG pathway enrichment analysis of the conserved comorbidity genes expressed in both species. Related to PD: established PD relevance; Potentially related to PD: potential PD relevance; Genes without established association with PD. PPI: Protein–protein interaction; PD: Parkinson’s disease; KEGG: Kyoto Encyclopedia of Genes and Genomes; DEG: differentially expressed gene; mDA: midbrain dopaminergic; DRG: dorsal root ganglia; ENS: enteric nervous system; OB: olfactory bulb; FDR: false discovery rate.