Advancing computer vision in spine surgery: a comprehensive review of current applications and future directions
Abstract
Artificial intelligence is rapidly reshaping healthcare, with computer vision enabling automated interpretation of imaging across a wide range of clinical applications. Given its heavy reliance on imaging across perioperative settings, spine surgery is particularly well-suited for the integration of computer vision technologies. At the same time, spine surgery is among the fastest-growing and most resource-intensive specialties, with rising demands that underscore the need for technologies capable of enhancing precision, efficiency, and safety. In this narrative review, we will synthesize current applications of computer vision across the spine surgery workflow, outline key barriers for implementation, and discuss future directions for translating these tools into widespread clinical practice. Computer vision tools span a spectrum of maturity, with some already achieving clinical deployment for spinopelvic parameter measurement, pathology detection, and surgical planning, while intraoperative applications represent the most actively developing frontier. These innovations may redefine the standard of care in spine surgery, enabling a new era of surgical performance and data-informed decision-making.
Keywords
INTRODUCTION
Nearly one million adults undergo instrumented spinal surgery procedures in the United States every year[1]. Given our aging population, case volume is still expected to rise significantly, corresponding with an increase in degenerative disease. From 2004 to 2015, elective lumbar fusion volume in the National Inpatient Sample (NIS) increased by 62.3%, with an increase of 138.7% in those 65 and over[2]. Altogether, this is indicative of the continued shift toward more expensive and complex procedures, at a time when spinal surgery already poses some of the highest healthcare utilization costs[3]. These growing challenges highlight an urgent demand for innovative solutions to increase surgical efficiency, minimize complications, and optimize resource allocation in spine care.
The rapid expansion of artificial intelligence (AI) offers a promising path forward. Throughout this review, we will categorize these technologies hierarchically. AI represents the overarching domain of automated systems designed to perform tasks typically requiring human cognition. Machine learning (ML) is a subset of AI focused on algorithms that learn from data exposure, whereas deep learning (DL) is a specific form of ML leveraging multi-layered neural networks. In this context, computer vision refers to the broad application of AI techniques to interpret visual data, such as medical imaging and video. Computer vision has already been widely deployed for tasks such as object detection, image classification, and extraction of higher-order features[4].
The plethora of ordered patient imaging and utilization of “cameras” in the form of endoscopes and surgical microscopes have created vast amounts of visual neurosurgical data. Numerous surgical specialties have already begun leveraging visual data streams of their own, propelled by the development of large surgical video datasets, including the Heidelberg Colorectal Dataset[5] and SurgToolLoc[6]. Computer vision can leverage this data to enhance care throughout the surgical workflow, from preoperative planning to intraoperative guidance to postoperative monitoring. In this review, we will discuss current utilizations of computer vision in spine surgery specifically, and then expand into some of the challenges and future directions of these emerging technologies.
ONE PATIENT, MANY IMAGES
Patients routinely undergo spinal imaging studies across the perioperative continuum. Imaging is not confined to a single phase of care, but repeatedly obtained before, during, and after surgery to generate a continuous stream of visual data throughout a patient’s clinical course. This ubiquity creates a natural opportunity for computer vision tools to support scalable, automated analysis at every stage of spine care.
Radiographs remain the most commonly obtained first-line imaging, typically acquired from multiple views with standard anteroposterior (AP) and lateral projections. Specialized dynamic views such as flexion-extension or standing-sitting are often used to evaluate the spine during dynamic motion. In addition, newer full-length, low-dose X-ray imaging can capture a view of the patient from the vertex to the feet, aiding in an understanding of posture and lower extremity compensation[7]. Despite their widespread use, plain radiographs are limited by poor spatial resolution, reduced soft tissue contrast, and constraints inherent to single-plane images.
When higher-resolution imaging is necessary, computed tomography (CT) or magnetic resonance imaging (MRI) scans are employed depending on the clinical context and the granularity of information needed for operative planning[8]. These modalities provide detailed visualization of osseous and soft tissue structures, including the spinal cord, intervertebral discs, nerve roots, and surrounding vasculature. However, the volume and complexity of these studies often require time-consuming interpretation, which may be further prone to variability between readers.
There is a significant opportunity for computer vision to be leveraged across the vast spectrum of spinal imaging encountered throughout the surgical workflow. Modern systems draw on a diverse array of DL architectures, including convolutional neural networks (CNNs), vision transformers (ViTs), and hybrid approaches[9-11]. These models are often designed to progressively extract visual features in a “hierarchical” fashion, starting with low-level patterns such as edges, then eventually advancing to more complete objects such as complex anatomical structures. To perform specific tasks, models are typically trained in a supervised fashion using large, well-annotated datasets, where ground-truth labels guide the learning process[9].
IMAGE SEGMENTATION
While images are initially composed of raw pixel data, both the human brain and computer vision systems must prioritize the identification of meaningful structures[12]. One of the core tasks in computer vision is image segmentation, which involves labeling specific parts or objects within an image. For spinal imaging, segmentation tasks typically involve the identification and delineation of regions of interest (ROIs), including key anatomical structures (e.g., vertebrae or intervertebral discs) and sites of pathology[13]. Proper segmentation is a foundational first step for almost all downstream tasks that we will discuss in this review, and images taken across the perioperative continuum rely on automated segmentation to establish accurate anatomical baselines.
AI algorithms have proven immensely helpful in automating segmentation tasks, where the traditional gold standard has relied on manual annotation by expert clinicians. This specifically involves surgeons or radiologists painstakingly labeling thousands of cross-sectional CT or MRI slices to identify anatomical structures and pathological regions. While highly accurate, this process is inefficient and not scalable. Earlier attempts to automate segmentation relied on classical mathematical techniques such as edge detection, thresholding, and region growing[14]. Although applicable in some contexts, these methods often fall short when applied to complex and heterogeneous medical images, which contain noise, artifacts, and overlapping tissue boundaries. DL-based segmentation methods can overcome these limitations, dramatically improving both the efficiency and accuracy of image segmentation across the medical domain[12].
Vertebrae across imaging modalities
As outlined in Table 1, significant work has focused on the segmentation of radiographs, given that they remain one of the most commonly used and accessible imaging modalities in spine surgery.
Selected studies of vertebral segmentation models from radiographs
| Study (Year) | Vertebrae | Dataset | Algorithm(s) | Metric | Performance |
| Zamora et al.[15] (2003) | Cervical and lumbar | NHANES II database 100 cervical, 100 lumbar radiographs | Active shape model Generalized Hough transform Deformable model | “Success rate,” cervical | 75% |
| “Success rate,” lumbar | 49% | ||||
| Koompairojn et al.[16] (2006) | Lumbar | NHANES II database 86 lumbar radiographs | Scale-invariant feature transform, Speeded-up robust features Active appearance model | Point-to-point error | 9.74 ± 2.54 pixels |
| Lecron et al.[17] (2012) | Cervical | Internal single-center 49 cervical radiographs | Active shape model | Point-to-line error | 4.43 pixels |
| Orientation error | 2.29 degrees | ||||
| Li et al.[18] (2016) | Lumbar | Microsoft dataset 50 C-arm lumbar radiographs | Feature-fusion deep learning model | Localization error, non-pathological cases | 1.23 ± 4.01 mm |
| Identification rate, non-pathological cases | 89.02% | ||||
| Localization error, pathological cases | 2.23 ± 8.76 mm | ||||
| Identification rate, pathological cases | 80.37% | ||||
| Al-Arif et al.[19] (2018) | Cervical | Internal single-center 296 lateral cervical radiographs | Fully convolutional network U-Net CNN U-Net-S CNN | Pixel-wise accuracy | 97.01% |
| Dice score | 0.84 | ||||
| Shape error | 1.69 mm | ||||
| Cho et al.[20] (2020) | Lumbar | Internal single-center 780 lateral lumbar radiographs | U-Net | Dice score | 0.821 |
| AUC | 0.914 | ||||
| Pixel-wise accuracy | 0.862 | ||||
| Shin et al.[21] (2020) | Cervical | Internal single-center 975 cervical lateral radiographs | U-Net | Pixel-wise accuracy | 96.67% |
| Kim et al.[22] (2021) | Lumbar | Internal single-center 797 lumbar radiographs | Pose-Net M-Net LevelSet | Dice score | 0.916 ± 0.022 |
| Precision | 0.846 ± 0.036 | ||||
| Sensitivity | 0.901 ± 0.029 | ||||
| Specificity | 0.996 ± 0.017 | ||||
| Hausdorff distance | 8.90 ± 3.23 mm | ||||
| Chen et al.[23] (2024) | Cervical and Lumbar | MEASURE1 trial: 293 cervical, 219 lumbar radiographs PREVENT trial: 132 cervical, 94 lumbar radiographs NHANES II: 445 radiographs | VertXNet (U-Net + Mask R-CNN via ensemble rule) | Dice score | 0.90 |
| Dice score, external PREVENT | 0.89 | ||||
| Dice score, external NHANES II | 0.88 |
The primary focus has been on vertebral body identification, with early work utilizing simpler algorithms including generalized Hough transforms, active appearance models, and active shape models[15-17]. These early methods achieved moderate accuracy, with segmentation errors on the order of a few millimeters, but typically required careful initialization or user input. During the 2010s, the first studies emerged leveraging AI for vertebral segmentation tasks. At this point, evaluation metrics began to include pixel-wise overlap scores (Dice, Jaccard) and boundary distances, replacing the crude millimeter error or detection rate metrics of prior decades[24]. One of the earliest publications from Li et al. utilized a basic CNN and feature fusion DL model to detect lumbar vertebrae[18]. Notably, this study used partially occluded C-arm imaging, better simulating the real-world quality of perioperative settings. The model had a vertebra identification rate of 89%, with a mean localization error of 1.2 mm for non-pathological cases. A subsequent study by Al-Arif et al. introduced a 3-step process to segment cervical vertebrae, consisting of global localization via a deep fully convolutional network, center localization via a deep probabilistic spatial regression network, and vertebrae segmentation via a shape-aware deep segmentation network[19]. This fully automatic framework achieved a Dice score of 0.84 with a mean shape error of 1.69 mm without requiring any human feedback. These initial DL applications marked a shift toward data-driven techniques, although they often still required multi-stage pipelines, such as detection followed by segmentation.
The advent of deep CNNs in medical imaging rapidly accelerated progress in spinal radiograph segmentation. The combination of fully convolutional architectures and larger training datasets yielded significant improvements in model performance. The popular U-Net architecture[25] was applied by Cho et al. and Shin et al. on lumbar and cervical vertebrae segmentation, respectively, resulting in strong segmentation performance[20,21]. A subsequent 2021 study by Kim et al., evaluating combinations of different CNNs, ultimately found that a Pose-Net + M-Net + LevelSet method yielded the highest performance on lumbar vertebrae segmentation, with a mean Dice score of 0.916 and a Hausdorff distance of 8.9 mm[22]. More recently, in 2024, Chen et al. presented a novel VertXNet model combining U-Net semantic segmentation and Mask Region-based Convolutional Neural Network (R-CNN) instance segmentation via the ensemble rule[23]. On both cervical and lumbar vertebrae segmentation tasks from lateral radiographs, VertXNet demonstrated superior performance to state-of-the-art (SOTA) models with a mean Dice score of 0.90, and scores of 0.89 and 0.88 on separate external datasets.
In parallel, AI segmentation efforts have expanded to CT and MRI scans, which pose distinct methodological challenges [Table 2]. While radiographic segmentation typically involves two-dimensional (2D) anatomical outlines with limited depth cues, CT and MRI provide volumetric data that require three-dimensional (3D) or multi-slice segmentation strategies, resulting in richer anatomical detail but higher computational demands. Early models, therefore, exhibited only modest performance, with limited generalizability and slow processing times.
Selected studies of vertebral segmentation models from CT and MRI imaging
| Study (Year) | Vertebrae | Dataset | Algorithm(s) | Metric | Performance |
| Janssens et al.[26] (2017) | Lumbar | MICCAI 2016 xVertSeg challenge Challenge: 15 spine CT scans | LocalizationNet (3D FCN) SegmentationNet (3D U-Net-like FCN) | Dice score | 0.958 ± 0.008 |
| Jaccard coefficient | 0.919 ± 0.015 | ||||
| Hausdorff distance | 4.32 ± 2.60 mm | ||||
| Symmetric surface distance | 0.37 ± 0.06 mm | ||||
| Lessmann et al.[27] (2019) | Cervical, thoracic, lumbar | MICCAI 2014: 15 thoracic and lumbar CT scans xVertSeg.v1: 15 lumbar CT scans National Lung Screening Trial: 55 low-dose chest CT scans 15 lumbar spine CT scans 23 T2-weighted sagittal MR scans | Iterative instance segmentation FCN inspired by U-Net | Dice score, CT average | 0.949 ± 0.021 |
| Dice score, MR average | 0.944 ± 0.033 | ||||
| Surface distance, CT average | 0.2 ± 0.1 mm | ||||
| Surface distance, MR average | 0.4 ± 0.3 mm | ||||
| Chen et al. as cited by Sekuboyina et al.[28] (2021) | Cervical, thoracic, lumbar | VerSe 2019 challenge | 3D U-Net for spine localization 3D U-Net for vertebral segmentation 3D ResNet-50 for vertebral classification | Dice score, public set | 0.917 |
| Hausdorff distance, public set | 6.14 mm | ||||
| Dice score, hidden set | 0.912 | ||||
| Hausdorff distance, hidden set | 7.15 mm | ||||
| Payer et al.[29] (2020) | Cervical, thoracic, lumbar | VerSe 2020 challenge | U-Net for spine localization SpatialConfiguration-Net for vertebral localization U-Net for vertebral segmentation | Dice score, public set | 0.909 |
| Hausdorff distance, public set | 6.35 mm | ||||
| Dice score, hidden set | 0.898 | ||||
| Hausdorff distance, hidden set | 7.08 mm | ||||
| Tao et al.[30] (2022) | Cervical, thoracic, lumbar | VerSe 2019 challenge MICCAI-CSI 2014 challenge | Spine-Transformers: ResNet50 backbone, Transformer | Dice score, public set | 0.911 |
| Hausdorff distance, public set | 6.34 mm | ||||
| Dice score, hidden set | 0.901 | ||||
| Hausdorff distance, hidden set | 6.68 | ||||
| Van der Graaf et al.[31] (2024) | Lumbar | Public Multicenter: 447 sagittal T1 and T2 MRI series | Iterative instance segmentation nnU-Net | Dice score | 0.93 ± 0.05 |
| Absolute surface distance | 0.49 ± 0.95 mm | ||||
| Zhang et al.[32] (2025) | Thoracic, lumbar, sacrum | CT-Verse: 41 CT scans CT-Liver: 150 CT scans MRSpineSeg: 215 T2-weighted MRIs | Visual state space blocks within nnU-Net | Dice score, CT-Verse | 0.9440 +/- 0.04 |
| Dice score, CT-Liver | 0.8833 +/- 0.05 | ||||
| Dice score, MRI dataset | 0.8828 +/- 0.03 |
Segmentation performance was markedly boosted by the advent of CNNs, which quickly proved effective for vertebra and spinal cord segmentation. A pioneering study from Janssens et al. presented a cascaded 3D CNN (LocalizationNet + SegmentationNet) for lumbar CT imaging, achieving an average Dice score of 0.958. However, evaluation was limited as the study utilized only 15 spine CT images from the 2016 xVertSeg challenge[26]. Later work from Lessmann et al. introduced an iterative instance CNN designed to segment vertebrae sequentially, or “instance-by-instance”[27]. Across five public datasets totaling 118 CT and MRI images, the model achieved an impressive average Dice score of 0.949, with anatomical identification accuracy of 93%.
The field has since benefited greatly from the release of large public datasets. The International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) has contributed significantly through the Large Scale Vertebrae Segmentation challenge (VerSe). VerSe 2019 released 160 CT images with expert voxel-level annotations, which included images with spinal pathologies[28]. This was expanded for VerSe 2020, which included an unprecedented 319 CT images of 300 patients from multiple institutions[33]. These benchmarks pushed algorithms toward greater generalizability, as both versions of the dataset were intentionally designed to capture a broad range of anatomical variants. Many of the top-performing submissions leveraged multi-stage approaches. For instance, Chen et al. implemented a three-step pipeline: an initial U-Net to identify ROIs, a second U-Net to segment individual vertebrae, and a final CNN to perform multi-class classification of the segmented vertebrae, as reported by Sekuboyina et al.[28]. Their model achieved strong performance across key metrics including Dice score and Hausdorff distance on the public testing set. Similarly, Payer et al. employed a multi-stage approach consisting of a U-Net for spinal localization, SpatialConfiguration-Net for vertebrae localization, and another U-Net for segmentation[29]. This approach outperformed all other submissions on the final hidden test set, achieving high scores across both labeling and segmentation tasks. Since the release of the VerSe benchmarks, newer architectures such as transformers have also been trained and tested on this publicly accessible data[30,32].
Open-access datasets beyond VerSe are being developed, with van der Graaf et al. releasing a multicenter dataset comprising 447 lumbar MRIs from 218 patients[31]. Using an iterative instance segmentation approach, their baseline model achieved mean Dice scores of 0.93 for vertebrae, 0.85 for intervertebral discs, and 0.92 for the spinal canal. They also trained an nnU-Net on the same dataset, which produced “almost identical” performance metrics, underscoring the robustness of modern segmentation architectures. Ultimately, such efforts to open-source imaging datasets are crucial to continue accelerating algorithm deployment, while laying the groundwork for fair benchmarking and reproducibility across the field.
Advancing modeling capabilities
Computer vision techniques have been extended to other spinal structures and pathologies, including intervertebral discs and spinal tumors. One notable challenge of radiographic imaging is poor soft tissue contrast, which limits the visibility of intervertebral discs. To address this, Sa et al. employed a Faster-R-CNN that achieved an average precision of 0.91 for the detection of intervertebral discs via lateral lumbar radiographs, though they did not attempt any form of segmentation[34]. In the future, it is possible that collapsed discs and degenerative disc changes could be studied through radiographs with the use of advanced segmentation and spatial relation methods. In parallel, progress has been made in segmenting spinal tumors using MRI. For example, Lemay et al. built upon prior spinal cord segmentation models to create a two-stage U-Net model capable of multiclass spinal cord tumor segmentation from MRIs[35]. Their model achieved a Dice score of 0.618 for tumor segmentation alone and was released publicly as part of the open-source “Spinal Cord Toolbox.” Together, these studies illustrate the growing versatility of DL models across both bony and soft tissue structures in spinal imaging, enabling automated detection and characterization of complex features.
Beyond an expansion in the range of clinical tasks performed, there have been dramatic architectural advancements in recent years with the emergence of foundation models and ViTs. While the most striking improvements in segmentation performance are still unfolding, early innovations have laid critical groundwork. Tao et al. conducted one of the earlier studies to leverage transformer architectures for the spine with Spine-Transformers, which combined a transformer-based 3D object detector and a multi-task encoder-decoder network to label arbitrary field-of-view CT scans automatically[30]. This was followed by Li et al.’s VerFormer, which further improved segmentation accuracy by making the network “vertebrae-aware,” outperforming conventional CNNs on multicenter CT benchmarks[36]. However, the landscape is rapidly evolving as the field shifts toward large-scale, task-agnostic foundation models. Meta’s Segment Anything Model (SAM) was the first foundation model for general image segmentation, sparking considerable interest in its adaptation for medical imaging tasks[37]. Building on this momentum, Ma et al. released MedSAM, a model trained on over 1.5 million medical image-mask pairs, which demonstrated competitive performance with specialist models across diverse clinical use cases[38]. On the vertebrae segmentation task, MedSAM achieved a median Dice score of 0.914, which was higher than the specialist DeepLabV3+ model (83.5%) and only slightly lower than the specialist U-Net model (92.9%). These results suggest that generalist foundation models, when sufficiently scaled and fine-tuned, may soon rival and even surpass domain-specific architectures in spinal segmentation tasks.
DIAGNOSTIC AND LONGITUDINAL IMAGING
Modern computer vision systems can now perform sophisticated clinical assessment tasks that previously relied solely on clinical expertise. Beyond identifying and segmenting spinal structures, computer vision tools can detect pathological conditions, classify disease severity, and aid in decision making throughout the surgical workflow [Table 3]. Accurate imaging assessment is a crucial aspect of advancing computer-aided analysis, with the ultimate goal of enhancing physicians’ ability to diagnose diseases and assess their severity.
Summary of computer vision use cases for clinical decision support
| Domain | Predictive tasks with selected studies |
| Quantification of spinal alignment and deformity radiographic measurements | Cobb angle: Seoud et al.[39] (2010), Lin et al.[40] (2020), Chen et al.[41] (2020), Dubost et al.[42] (2020), Lin et al.[43] (2022), Alukaev et al.[44] (2022), Chen et al.[45] (2024) Spinopelvic parameters: Orosz et al.[46] (2022), Harake et al.[47] (2024), Joshi et al.[48] (2025), Kang et al.[49] (2025) Postoperative: Grover et al.[50] (2022), Suri et al.[51] (2023), Löchel et al.[52] (2024), Vogt et al.[53] (2025) |
| Automated classification of spinal pathology from imaging | Vertebral fractures: Derkatch et al.[54] (2019), Li et al.[55] (2021), Shen et al.[56] (2023), Liu et al.[57] (2023) Degenerative: Jamaludin et al.[58] (2017), Merali et al.[59] (2021), Hallinan et al.[60] (2021), Nguyen et al.[61] (2021) Oncology: Maki et al.[62] (2020), Haim et al.[63] (2024), Zhuo et al.[64] (2022) |
| Identification of instrumentation and postoperative monitoring | Huang et al.[65] (2019), Yang et al.[66] (2021), Dutt et al.[67] (2022), Park et al.[68] (2022), Schwartz et al.[69] (2022), Anand et al.[70] (2023), Chun et al.[71] (2025), Sun et al.[72] (2025), Waranusast et al.[73] (2025) |
| Multimodal predictive modeling for surgical outcomes | Raw imaging: De Silva et al.[74] (2020) Radiomics: Saravi et al.[75] (2023), Wu et al.[76] (2025) Complications: Durand et al.[77] (2021), Rudisill et al.[78] (2022), Johnson et al.[79] (2023), Brigato et al.[80] (2025), Zheng et al.[81] (2025) |
Spinopelvic parameter detection
Initial spinal vertebrae segmentation efforts laid the groundwork for subsequent models to aid in measuring key spinal alignment parameters[82]. Commonly evaluated metrics include coronal and sagittal balance, Cobb angle, pelvic incidence (PI), pelvic tilt (PT), sacral slope (SS), lumbar lordosis (LL), and sagittal vertical axis (SVA)[7,8]. These are essential quantitative measurements of spinal alignment and deformity for guiding management prior to surgery, confirming alignment targets intraoperatively, and informing postoperative assessments of alignment over time. However, they can be tedious and laborious to perform manually, and their frequent use throughout the perioperative continuum highlights a need for automated approaches.
The Cobb angle is one such spinal parameter that is critical in the management of patients with coronal deformity, for which ML models have made significant advancements in automated measurement. An early study by Seoud et al. in 2010 demonstrated that a classifier composed of three support vector machines (SVMs) could predict scoliotic curvature from trunk surface radiographs with a prediction rate of 72.2%[39]. In 2019, there was an influx of studies following the MICCAI AASCE (Accurate Automated Spinal Curvature Estimation) challenge. All three of the highest performing submissions implemented some variation of a two-stage CNN system, typically starting with a segmentation model[40-42]. The highest-performing submission, Seg4Reg, from Lin et al., consisted of a two-stage CNN segmentation and regression pipeline, achieving a 21.71 symmetric mean absolute percentage error (SMAPE)[40]. In 2022, the authors released an updated Seg4Reg+ model that further implements an attention regularization module with class activation maps, achieving SOTA performance with a SMAPE of 7.32[43].
Other work has expanded beyond CNN architectures, with a study by Alukaev et al. utilizing an ensemble of U-Nets to compute vertebral body heights, intervertebral disc heights, and coronal and sagittal Cobb angles based on 3D CT spine images[44]. The model was found to have mean absolute errors in the computation of vertebral body heights, intervertebral disc heights, and coronal and sagittal Cobb angles of 1.17 mm, 0.54 mm, and 3.42 degrees as compared with manual measurements. There is now commercially available software to measure Cobb angles, with one called “IB Lab SQUIRREL” trained on an extensive dataset of over 17,000 images, though limited to a single site. An external study of the software found strong agreement with the reference standard generated by four experienced radiologists, with a mean difference of 0.16 degrees[45].
As a mark of the rapid evolution of computer vision systems, the most recent studies have pivoted toward multitask learning frameworks capable of simultaneously predicting multiple spinal alignment parameters. Rather than focusing solely on a single metric such as the Cobb angle, these models are now trained to output a suite of spinopelvic parameters from a single lateral radiograph. A 2022 study by Orosz et al. used three DL models to automatically compute spinopelvic parameters, applied to both preoperative and postoperative spine radiographs[46]. This model was, however, limited by its minimal exposure to patients with severe coronal deformities. Building on this work, Harake et al. (2024) presented the SpinePose algorithm containing three parallel CNNs to predict spinopelvic parameters from both sagittal whole-spine and sagittal lumbosacral radiographs[47]. The median parameter errors of SpinePose for landmarks of SVA, LL, and PT were 2.2 mm, 2.6 degrees, and 1.3 degrees, respectively. The model was recently validated in an out-of-state, external cohort[48]. The median parameter error was not reported in the publication; however, there were no statistically significant differences between SVA, PT, PI, and LL metrics and the ground-truth annotations.
The latest DL model by Kang et al. implemented a U-Net for spine segmentation and a Keypoint Region-based CNN for landmark detection[49]. This model was trained on 1,011 standing lumbar lateral radiographs and was found to achieve near-perfect intraclass correlation coefficients (ICCs) across all spinopelvic parameters, including LL, PI, PT, and SS, for even patients with severe coronal deformity. Furthermore, Kang’s model demonstrated ICCs ranging from 0.96 to 0.99, surpassing the performance of orthopedic surgeons. These advances reflect the maturation of computer vision toward clinically validated tools for comprehensive spinal alignment analysis.
Spinopelvic monitoring
Postoperative imaging presents distinct technical challenges, as large constructs and hardware can obscure bony landmarks, introduce imaging artifacts, or alter expected spatial relationships between anatomical structures. Only a subset of alignment measurements have been validated specifically in postoperative, instrumented cohorts. Grover et al. evaluated an unspecified DL framework on 170 paired preoperative and postoperative full-spine radiographs, measuring parameters of sagittal balance such as PI, PT, SS, and LL[50]. Postoperative ICCs ranged from 0.72 to 0.89 with a landmark detection rate of 91%, slightly lower than preoperative performance but within clinically acceptable ranges. Löchel et al. extended this analysis to an adult spinal deformity (ASD) cohort of 141 patients, reporting ICCs of 0.72-0.96 across postoperative PI, PT, SS, LL, and SVA, with a landmark detection rate of 84% postoperatively compared to 91.5% preoperatively[52]. The authors also explicitly noted that the presence of long instrumented constructs was the primary driver of the observed accuracy reduction relative to preoperative imaging. However, more recent work has demonstrated that hardware-invariant performance is achievable. Vogt et al. validated an AI algorithm on 129 paired pre- and postoperative lateral cervical radiographs, measuring five cervical sagittal balance parameters including C2-C7 lordosis, C1-C7 SVA, C2-C7 SVA, C7 slope, and T1 slope[53]. The model achieved postoperative ICCs of 0.86-0.99, which was comparable to preoperative outcomes. Critically, the authors attributed this result to deliberate enrichment of the training set with a broad variety of implant types combined with data augmentation. Similarly, Suri et al. developed a SpineTK-based pipeline for automated Cobb angle measurement on 1,310 AP radiographs from six centers and showed no statistically significant difference in accuracy between cases with and without surgical hardware (P = 0.80), with an overall ICC of 0.96[51]. Taken together, these findings are encouraging as they suggest that with appropriate model design and deliberate curation of postoperative training data, the presence of instrumentation need not be a barrier to accurate alignment surveillance.
Disease classification
In a similar vein, computer vision tools can be used directly as diagnostic agents to identify spinal pathologies from medical imaging. Many of these models build upon upstream segmentation of key anatomical regions to extract features for downstream classification. In contrast, some models leverage well-annotated data to classify pathology from raw images directly. DL tools are now demonstrating improved diagnostic performance across a range of spinal conditions in both emergent trauma and routine elective spine surgery settings.
Among spinal pathologies, vertebral fractures have been one of the most successfully identified diagnoses by DL algorithms. Early work by Derkatch et al. implemented an ensemble of CNNs, including InceptionResNetV2 and DenseNet, to detect vertebral fractures on dual X-ray absorptiometry, resulting in a respectable area under the curve (AUC) of 0.94, sensitivity of 87.4%, and specificity of 88.4%[54]. The utilization of ensemble methods and more advanced models in later work yielded further performance improvements. A study by Li et al. utilized a You Only Look Once (YOLO) v3 model for object detection and an ensemble of ResNet34, DenseNet121, and DenseNet201 models for detecting lumbar vertebral fractures[55]. The total accuracy was 93%, with a sensitivity of 91% and a specificity of 93%, on an internal validation dataset. Subsequent work from Shen et al. in 2023 demonstrated a multitask detection network combining YOLOv5 for segmentation and a grid search optimization algorithm for grading of osteoporotic vertebral fractures from thoracic and lumbar radiographs[56]. This YOLO-XRAY system had an even higher overall accuracy of 96.9% for all fractures on an external validation set. Researchers have more recently extended their efforts beyond fracture detection to the classification of vertebral fracture subtypes. For example, Liu et al. demonstrated impressive results using a two-stream compare-and-contrast network to classify benign versus malignant vertebral compression fractures, which at times reportedly outperformed the diagnostic accuracy of individual radiologists[57].
There have also been significant efforts to develop models for a broader range of degenerative pathologies. One of the earliest and most influential examples was the SpineNet model, introduced by Jamaludin et al. in 2017, which trained CNNs to classify multiple lumbar MRI abnormalities simultaneously, including disc narrowing, spondylolisthesis, and central stenosis[58]. Although the accuracy of disease classification tasks left room for improvement (ranging widely from 60% to 90%), the researchers demonstrated near-human performance on radiological grading and showed that multi-task training actually improved performance. A later study by Merali et al. on AO Spine patients undergoing surgery for degenerative cervical myelopathy used a modified ResNet-50 model to detect cervical spinal cord compression on MRI scans[59]. The model was trained on 6588 images and achieved 88% sensitivity and 89% specificity on the test set. More recent DL models have only continued to improve, now frequently matching the performance of radiologists. A dual-component CNN framework created by Hallinan et al. was able to grade lumbar central canal stenosis and foraminal narrowing on MRI almost as reliably as subspecialist radiologists, reaching near-perfect agreement [κ (Cohen's Kappa) of 0.95-0.96] on an external validation set[60]. There have since been concerted efforts to increase the availability of open-access spine imaging, with Nguyen et al. releasing the VinDr-SpineXR dataset, which contains over 10,000 manually annotated spinal radiographs[61]. Their initial results, training a large-scale CNN for an abnormality detection task, achieved an AUC of 0.886.
Spinal oncology has emerged as another active area for the application of diagnostic computer vision. In a study by Maki et al., researchers utilized a CNN to differentiate between spinal schwannoma and meningiomas in baseline MRI imaging, achieving an AUC of 0.876 on T2 and 0.870 on T1 imaging[62]. The performance of the model was comparable to that of two board-certified radiologists, though its evaluation was limited by a small dataset (50 schwannoma and 34 meningioma cases) and the absence of external validation. Building on this, Haim et al. developed a multi-class classification framework using Fast.ai to distinguish between the spinal pathologies of carcinoma, infection, meningioma, and schwannoma[63]. The algorithm reached an accuracy of 0.93 on the test set. However, this study similarly lacked external validation and was limited to a cohort of 231 patients. Finally, Zhuo et al. addressed the diagnostic task of differentiating intramedullary spinal cord tumors from demyelinating lesions such as multiple sclerosis[64]. Using a combination of 2D MultiResUNet and DenseNet121 architectures, their model was trained on a cohort of 490 patients and prospectively tested on an independent set of 157 patients. The system achieved 96% accuracy in distinguishing tumors from demyelinating lesions, and 82% accuracy in classifying tumor subtypes, including astrocytomas and ependymomas. Altogether, these studies underscore the potential of computer vision in spinal oncology while highlighting the need for larger, more diverse datasets and external validation to support clinical translation.
Implant assessment
Beyond characterization of anatomical disease states, instrumentation can be segmented and classified from clinical images, which is particularly relevant when planning revision surgeries where prior hardware may be unfamiliar, or for monitoring of implant integrity in the postoperative period. For pedicle screw identification, early work by Anand et al. applied a bag-of-visual-words technique, KAZE feature extractor, and SVM to classify thoracolumbar pedicle screws on radiographs[70]. Their classifier achieved accuracy rates of 93.15% on lateral views, 88.98% on anteroposterior views, and 91.08% on fused images, notably including cases with multiple spinal levels and overlapping instrumentation. Critically, in a direct head-to-head comparison, the model classified instrumentation in 14 s versus 20 min for humans, while also outperforming humans in accuracy (79% vs. 44%). Another study by Yang et al. focusing on deeper architectures compared a custom ResNet-34, Google AutoML, and Apple CreateML on a spinal implant detection task. Using 2,894 lumbar radiographs spanning five different pedicle screw types, the highest-performing custom model achieved precision of 97.0%-98.7% and recall of 96.7%-98.2%[66]. Most recently, Waranusast et al. developed a fully automated end-to-end pipeline for pedicle screw manufacturer identification on plain radiographs, achieving 100% accuracy on AP views and 98.9% on lateral views, comparable to spine surgeon performance of 97.1%[73].
Parallel efforts have addressed cervical hardware, with a foundational study by Huang et al. using the bag-of-visual-words technique and KAZE feature detection techniques to classify the manufacturer and implant system placed in patients receiving anterior cervical discectomy and fusion (ACDF)[65]. The classifier selected the correct ACDF system as the top choice in 91.5% of cases and as one of the top two choices in 97.1% of cases. Similar work by Dutt et al. used a weakly supervised approach combining EfficientDet for implant localization and DenseNet121 for brand classification across 984 patients and 10 hardware models, achieving a localization overlap of 86.8% and a classification F1 of 93.5%-98.7%[67]. Notably, these computer vision tools do not need to be confined to dedicated imaging workstations, as Schwartz et al. demonstrated that a CNN could identify anterior cervical implant types from smartphone photographs of radiographs with high accuracy across 15 plate types, with 85.8% top-1 accuracy and 94.4% top-3 accuracy[69].
Whereas implant identification is a reasonably well-studied task, the clinical problem of detecting hardware failure on postoperative imaging remains comparatively underdeveloped. For example, in patients with ASD, rod fracture affects 6.8% to 14.9% of patients and often necessitates revision but remains challenging to detect or predict[83,84]. To address this, Chun et al. developed a patch-wise ViT pipeline applied to 9,924 spinal radiographs from 798 patients, first detecting instrumentation and then classifying each 224 × 224 patch for implant fractures[71]. The model alone achieved an accuracy of 0.83 and recall of 0.94 overall, though precision was poor with an F1 score of 0.54. Notably, when radiologists used the model as an assistive tool, their recall rose from 0.70 to 0.95, with substantial gains from less experienced readers (0.49 to 0.92). Computer vision has also been applied to fusion status assessment, a related but distinct postoperative endpoint. Park et al. trained a CNN on postoperative cervical radiographs to classify fusion status after ACDF, achieving an accuracy of 89.5% and an AUC of 0.89[68]. For cage subsidence, a study from Sun et al. developed a LightGBM (Light Gradient Boosting Machine) classifier trained on 620 patients that incorporated imaging features such as cage height and postoperative intervertebral height, achieving an AUC of 0.98[72]. These findings suggest that vision-based screening could serve as a practical triage tool for postoperative implant surveillance.
Outcome prediction
Of significant interest to spine surgeons is outcome forecasting, which has been a predominant focus of ML models in the surgical literature. The ability to predict whether patients will demonstrate weak or strong outcomes after surgery has substantial implications for patient counseling and decision making. There has long been interest in whether spinal imaging might hold useful biomarkers, as numerous studies have assessed the correlations between spinopelvic alignment parameters and postoperative measures of disability, pain, and functional status[85-87]. However, the majority of published outcome models in spine surgery do not incorporate imaging features, in part due to the challenges of image availability, preprocessing, and feature extraction. Exceptions are typically retrospective studies in which radiographic features were already collected and manually annotated, though these remain more uncommon[88,89]. Where imaging features have been incorporated, they are almost invariably combined with clinical and demographic variables, a multimodal fusion approach that is likely to define the next generation of predictive frameworks in spine surgery.
Full-blown computer vision frameworks that directly integrate raw imaging data into predictive modeling frameworks are particularly rare, though a few efforts have recently emerged. The SpineCloud framework, developed by De Silva et al., extracted spinal parameters from CT imaging and fed a combination of demographic, clinical, and radiographic features into a boosted decision tree to predict patient-reported outcomes (PROs) after surgery[74]. Model performance improved after incorporating intraoperative and immediate postoperative imaging features, with prediction of the 3-month modified Japanese Orthopaedic Association (mJOA) score rising from an AUC of 0.54 to 0.71. Although a promising initial proof-of-concept, the method was evaluated on only a limited set of 64 patients.
Other recent studies have employed a radiomics-based approach, in which handcrafted quantitative imaging features, such as shape, texture, and spatial relationships, are extracted from ROIs. Saravi et al. extracted 3D radiomics features from the preoperative T2 MRI scans of 172 patients and then evaluated a combined ML model using both clinical and radiomics features[75]. The authors found a minimal increase in accuracy, from 87.69% to 88.17%, when incorporating radiomics features. However, a more recent study by Wu et al., which utilized a larger dataset of 1,520 patients, performed segmentation of paraspinal muscles from CT and MRI scans[76]. The study then conducted feature extraction from the isolated ROIs using the Pyradiomics package. A combined model was developed using clinical variables and a radiomics score to predict functional status after degenerative lumbar spondylolisthesis surgery. On the derivation dataset, the combined model outperformed both the clinical and radiomics models, with an AUC of 0.85 compared to AUCs of 0.73 and 0.77, respectively. The combined model also demonstrated strong performance with AUCs of 0.82, 0.79, and 0.80 on three separate external test sets.
The utility of these predictive models extends beyond functional recovery to the assessment of postoperative complications. Cage subsidence, which can occur anywhere from 3.3% to 51.2% depending on the technique used, is a common yet unpredictable adverse event, underscoring the need for objective, imaging-based predictive markers[90]. A study by Zheng et al. extracted Pyradiomics features from preoperative CT and combined them with clinical variables from 253 patients undergoing ACDF[81]. A combined radiomic-clinical classifier outperformed clinical features alone in predicting cage subsidence of ≥ 3 mm at final follow-up, with an AUC of 0.81. In a separate cohort of 366 patients undergoing ACDF, Rudisill et al. developed an ML algorithm to predict early-onset adjacent segment degeneration, achieving an AUC of 0.794[78]. Presence of C4/C5 disc bulges, C6/C7 disc bulges, and osteophytes on preoperative imaging were the most important predictive features.
Computer vision models have also been developed extensively to predict dynamic postoperative complications, including proximal junctional kyphosis (PJK), which similarly affects 20%-40% of patients operated for ASD[91]. A study from Durand et al. applied an unsupervised self-organizing map to the lateral radiographs of 915 ASD patients from the International Spine Study Group database, identifying six morphological clusters with significantly different rates of PJK[77]. Whether this clustering reflects underlying biological differences that drive junctional failure, or simply correlates with surgical decision-making patterns, remains an open question, but it establishes that image-derived phenotyping has a role in PJK risk stratification. A subsequent 2025 systematic review from Brigato et al. synthesized seven ML studies encompassing 2,179 patients and reported model accuracies of 72.5%-100% and AUCs up to 1.0 for PJK prediction, with age, bone mineral density, and preoperative alignment parameters emerging as consistent predictors across studies[80]. Additional studies have shed insight on which specific imaging modalities best capture subtle drivers of junctional failure. Johnson et al. compared a classical SVM using clinical and radiographic features, a CNN trained on raw scoliosis radiographs, and a CNN trained on preoperative thoracic MRIs on 191 ASD patients with at least two-year follow-up[79]. The MRI-based CNN demonstrated the highest performance, achieving sensitivity of 73.1% and specificity of 79.5%. Attention maps further revealed that soft tissue characteristics, rather than bony alignment alone, were the most informative predictors of subsequent junctional failure. For certain tasks, reliance on radiographic inputs may represent a ceiling, and multimodal imaging approaches may be necessary to increase accuracy and clinical utility of model predictions.
APPLICATIONS IN THE OPERATING ROOM
The growing convergence of preoperative imaging and intraoperative sensors has opened a new frontier for computer vision tools inside the operating room. These capabilities create opportunities not only for more accurate navigation but also for decision support, safety alerts, and robotic assistance. As computing units become smaller and more readily available, intraoperative AI will be poised to augment surgeon perception and optimize surgical efficiency.
Spine navigation
Traditional open procedures rely on large incisions and extensive intraoperative fluoroscopy to achieve proper anatomical localization and guide pedicle screw placement. By contrast, modern navigation systems, such as the StealthStation S8 (Medtronic), Valence (Alphatec), 7D Surgical System (SeaSpine, now Orthofix), and Brainlab Spine & Trauma Navigation, can generate an image-guided roadmap that tracks instrument tips relative to patient anatomy with submillimetric accuracy[92]. Navigation has been shown to increase screw placement accuracy, lower revision rates, and reduce intraoperative radiation exposure to both staff and patients[93-95]. However, despite these gains, current navigation workflows still depend heavily on user-initiated registration and manual validation. There are numerous avenues for improvement by computer vision techniques, whether to increase accuracy or efficiency.
Computer vision can be used to accelerate the registration process that links initial imaging to the actual anatomy, a critical step for accurate surgical navigation. Many platforms currently require the placement of external tracking devices, such as infrared markers, which can be cumbersome and increase setup time. To streamline this process, researchers have begun exploring marker-less, vision-based alternatives. Liebmann et al. presented a proof-of-concept system for marker-less and radiation-free registration, leveraging a U-Net architecture to segment the lumbar spine from preoperative imaging and generate an initial anatomical pose[96]. This pose is then refined in real time during the procedure, and the system integrates with an augmented reality (AR) navigation platform for pedicle screw placement. In an initial cadaveric study involving 10 screws, the system achieved a median registration error of 1 mm and placement accuracy of 100%. Although preliminary, these results demonstrate the feasibility of fully marker-less, vision-based navigation. There have been efforts to leverage even minimal imaging for registration tasks, with Abumoussa et al. developing a self-supervised ML algorithm for 2D-to-3D registration using a single fluoroscopic image of the spine[97]. After simulating 10,160 radiographs from 127 cone-beam CT studies, the model demonstrated a convergence success rate of 82% with a mean compute time of 1.96 s.
Another key computer vision application is the intraoperative detection and verification of spinal instrumentation, building upon methods described earlier in this review. A study by Doerr et al. applied a Faster R-CNN architecture with a VGG16 backbone to fit bounding boxes around pedicle screws on intraoperative imaging[98]. The model was trained using a dataset of 12,500 images derived from intraoperative O-arm cone-beam CT scans obtained before and after pedicle screw placement. Detection accuracy and precision were 86.6% and 92.6% respectively, and localization accuracy was within 1.5 mm. In parallel, researchers have begun integrating AI models directly into intraoperative workflows to support automated pedicle screw planning and placement. Burström et al. described the use of an (unspecified) segmentation algorithm to perform real-time 3D vertebrae segmentation from intraoperative cone-beam CT images, for the ultimate purpose of generating automated pedicle screw trajectories[99]. On cadavers, the vertebra segmentation error was a root-mean-squared distance of 0.7 mm. Across 20 real clinical cases, the model achieved “clinically adequate” pedicle segmentation for 86.1% of the pedicles, with frequent failure points requiring manual intervention, including those with high Cobb angles over 75 degrees and vertebrae with severe degeneration.
Even beyond the core tasks of segmentation and classification, AI techniques can enable cross-modality translation, allowing for the extraction of volumetric information from sparse or lower-dimensional imaging. Jecklin et al. developed the X23D DL algorithm to intraoperatively estimate the 3D shape of lumbar vertebrae using a small number of fluoroscopic X-ray images[100]. Trained and evaluated using the CTSpine1K dataset, X23D achieved an average F1 score of 0.88 and a surface reconstruction score of 71%, approximately 22% higher than SOTA methods at the time. In a subsequent feasibility study of 49 pedicle screws placed in human cadavers, surgeons assigned to freehand with fluoroscopy versus X23D-based navigation did not show significant differences in breach rate, placement time, or radiation exposure[101].
The synthesis of CT-equivalent images from MRI scans has been another area of focus. Although MRI reduces exposure to ionizing radiation, CT is often preferred by surgeons for assessing bony structures and for use in neuronavigation. While quantitative lumbar CT is generally considered low-dose [approximately 0.1 millisieverts (mSv) per scan], patients may undergo multiple imaging studies across their surgical course[102]. The lifetime attributable cancer risk from spinal CT is estimated to range from approximately 1 in 200,000 to 1 in 3,200, depending on factors such as anatomic region, coverage, and imaging protocol[102]. The BoneMRI technology was one of the first DL tools developed to address this need by enabling 3D MRI-to-CT mapping[103]. An early study found that the synthetic CT scans were non-inferior to traditional CT, with a voxel-wise comparison showing a Dice score of 0.84[104]. Building on these initial validations, Staartjes et al. implemented their own prototype version of BoneMRI by deploying a patch-based CNN with Keras. In a case study of three patients, the authors stated that synthetic CT scans could also enable precise surgical planning and neuronavigation in the same manner as traditional CTs[105]. The BoneMRI technology was later tested in combination with the ExcelsiusGPS robot by Davidar et al. on a human cadaver[106]. When comparing the use of synthetic CT versus routine CT scan during the placement of 14 lumbosacral screws, the researchers did not find significant differences in mean tip distance, tail distance, and angular deviation. As these technologies continue to mature, MRI-to-CT tools may reduce cumulative radiation exposure while maintaining the granularity required for surgical planning and navigation, though future studies are needed to quantify their impact at both the patient and population levels.
Augmented reality
AR is the next major technology being introduced to the operating room, and there is significant interest in the ability of AR to change how surgeons visualize anatomy and interact with instruments during procedures[107-109]. Notably, AR technologies overlay directly onto the surgeon’s view of the operative field, delivering navigational guidance within the natural line of sight rather than on a remote monitor. Thus far, pedicle screw placement has been the most extensively validated application. The seminal in-human study by Elmi-Terander et al. reported 94.1% accuracy across 253 screws with no grade 3 misplacements[110]. This result was confirmed across multiple subsequent single-center series using the xVision head-mounted display platform, with one of the largest follow-ups documenting a 0.49% intraoperative replacement rate and zero instrumentation-related reoperations for 606 screws across two institutions[111]. Level I evidence has recently emerged from a randomized multicenter trial by Ma et al. enrolling 150 patients and approximately 700 screws, which demonstrated statistically significant superiority of AR navigation over freehand placement (difference of 6.3%, P = 0.0003)[112]. Beyond screw placement, AR navigation has been applied to a variety of minimally invasive and microscope-assisted interventions. A randomized trial from Auloge et al. of AR-guided vertebroplasty demonstrated equivalent procedural accuracy to fluoroscopy, along with reductions in delivered radiation dose of approximately 50%[113]. For open procedures, Carl et al. prospectively applied microscope-integrated AR across 42 consecutive spinal surgeries, confirming initial technical feasibility at all levels with low effective radiation doses[114].
The broader applications of AR in spine surgery have been extensively documented elsewhere and are not the primary focus of this paper[115]. It is worth noting, however, that computer vision is foundational to AR systems, as many of the same technologies that enable automated landmark detection, anatomical segmentation, and spatial registration on static radiographs also underpin real-time AR rendering. Whether AR ultimately changes patient outcomes in a meaningful and cost-effective way remains to be established. Notably, no randomized trial has yet compared AR to conventional CT-based navigation, the current standard of care for complex instrumented fusion. Addressing these gaps in the literature will be essential to establishing the role of AR in spine surgery, and the convergence of AR and AI-driven computer vision may be key to ultimately paving the way for intelligent surgical systems. Next-generation AR tools will be unlocked by real-time computer vision models capable of dynamically interpreting the surgical field, identifying critical structures, and adjusting visualizations based on the current phase of surgery.
Endoscopic surgery
Intraoperative integration of computer vision tools will require algorithms to accurately identify objects within the surgical field. This capability becomes especially critical in minimally invasive spine surgery, where the operative corridor and visual field are significantly restricted. While computer vision applications for endoscopic scene understanding have gained substantial traction in general surgery, their application in spine surgery remains comparatively underexplored[4]. Early work from Cui et al. employed the YOLOv3 architecture to label nerves and dura mater within endoscopic images. In their first study, using data from 15 different endoscopic spinal surgeries, the model identified images containing nerve and dural tissue with an accuracy of 0.95, a sensitivity of 0.94, and a specificity of 0.98[116]. In their second study, using data from a larger cohort of 65 patients who underwent percutaneous transforaminal endoscopic discectomy, images with nerve and dural tissue were well-identified with an accuracy of 0.92, a sensitivity of 0.91, and a specificity of 0.94[117].
Despite their strong performance in binary classification tasks, these early models have critical limitations. The analyses were conducted on selected still images extracted from video rather than continuous video streams, precluding real-time application. Furthermore, the models provided image-level labels without pixel-wise localization or segmentation of anatomical structures, thereby offering no spatial specificity that could guide intraoperative decision-making. Addressing these gaps, Peng et al. assembled the first large-scale dataset of spinal endoscopy with finely annotated nerves, resulting in approximately 10,000 consecutive frames captured on SPINENDOS systems (Munich, Germany)[118]. Scenes were selected to reflect typical “visual and motion characteristics” of surgery, and each frame was specifically annotated with pixel-level masks of spinal nerves. The researchers developed a Frame-UNet model, enhanced with both inter-frame and channel self-attention, which achieved a Dice score of 0.89 and mean absolute error of 0.016 on the spinal-nerve benchmark, significantly outperforming the baseline U-Net and ResNet architectures across all evaluation metrics.
Overall, such techniques highlight the potential of real-time computer vision tools to augment intraoperative awareness, particularly for less experienced surgeons and trainees who may benefit from automated alerts when sensitive structures, including spinal nerves or vessels, enter the visual field. As these tools become more reliable in identifying critical tissue types, they could be more seamlessly integrated into AR platforms and intraoperative workflows to enhance surgical precision and safety.
Commercial adaptation
These computer vision technologies are rapidly transitioning from research to commercial implementation in the operating room. In 2024, the ATEC EOS Insight platform demonstrated a new industry focus on “AI-driven” imaging analytics, boasting alignment measurements to support surgical planning, intraoperative verification, and postoperative analysis[119]. Similarly, the Carlsmed Aprevo technology platform leverages AI segmentation techniques to support surgical planning[120]. These platforms are being leveraged to design patient-specific rods and interbody devices, respectively, for spinal surgery. Medtronic’s UNiD Adaptive Spine Intelligence platform also supports custom rods, with data suggesting reductions in rod fracture rates to as low as 2.2% from the historical rate of approximately 9%[121]. Furthermore, patient-specific cages have shown a decreased risk of subsidence and can be used to achieve targeted alignment goals[122,123].
The following representative case illustrates the clinical utility of integrated AI planning platforms. A 77-year-old woman with scoliosis and multiplanar malalignment presented after failing extensive conservative management. She underwent a two-stage circumferential minimally invasive surgery via stage 1 L4-S1 anterior lumbar interbody fusion and T12-L4 lateral lumbar interbody fusion along with stage 2 percutaneous posterior instrumentation from T9-pelvis.
Preoperative planning utilized the Carlsmed Aprevo and Medtronic UNiD platforms in a sequential workflow. Anterior column correction was first planned with Aprevo to engineer patient-specific interbody cages [Figure 1], after which the correction plan was exported to UNiD to fabricate matched patient-specific posterior rods [Figure 2]. This integrated planning sequence is helpful to achieve a reliable circumferential minimally invasive deformity correction.
Figure 1. Carlsmed Aprevo preoperative planning for adult spinal deformity correction. (A) Baseline pre-operative versus planned parameter measurements from L1-S1 (B) 3D CT-based reconstruction comparing baseline and planned post-interbody construct configurations; (C) Patient-specific implant planning shown for L4-5 with annotated screw trajectories, implant footprint, and multiplanar construct visualization. 3D: Three-dimensional; CT: computed tomography; ALIFX: anterior lumbar interbody fusion, interfixated; AP: antero-posterior.
Figure 2. Medtronic UNiD preoperative planning for adult spinal deformity correction. (A) Representative preoperative radiographs and surgical plan generated using dedicated spine surgical planning software. Left and center panels show anteroposterior and lateral radiographs demonstrating the patient’s baseline alignment. The right panel illustrates the surgical plan, with proposed instrumentation for T9 to pelvis fusion; (B) Corresponding strategy table summarizing preoperative and planned spinopelvic parameters, with targeted corrections including SVA from 87 to 19 mm, PI-LL from 23° to 1°, and LL from -36° to -58°. The planned cage sizes and segmental corrections are imported from the Carlsmed Aprevo platform. SVA: Sagittal vertical axis; LLIF: lateral lumbar interbody fusion; ALIF: anterior lumbar interbody fusion; TK: thoracic kyphosis; TL: thoracolumbar.
At approximately six weeks postoperatively, postoperative full-length EOS radiographs confirmed successful deformity correction with restoration of sagittal and coronal alignment [Figure 3]. The patient notably reported clinical improvement with complete resolution of back pain without opioid use. Overall, this case illustrates how commercially available AI-driven planning platforms can translate preoperative deformity correction targets into patient-specific implants that are directly actionable in the operating room.
Figure 3. Preoperative and postoperative full-length EOS radiographs. Anteroposterior views obtained before (A) and after (B) posterior instrumented fusion with custom titanium rods, alongside corresponding lateral views before (C) and after (D) surgery.
Recent advances are further extending computer vision into real-time intraoperative guidance. The previously discussed BoneMRI technology has been successfully used in human spine surgeries and, as of April 2025, has been integrated into the Brainlab Spine Navigation software. Also in April 2025, the company Proprio received Food and Drug Administration (FDA) clearance for what it describes as the “world’s first AI surgical guidance platform.” The platform builds on a June 2023 white paper that introduced a method for intraoperative measurement of sagittal alignment using what the company refers to as “radiation-free volumetric intelligence”[124]. The core of the Proprio technology lies in its ability to derive 3D vertebral coordinates in real time, which can be immediately leveraged to compute regional and segmental sagittal angles. In a preliminary validation, the AI-derived sagittal angle measurements demonstrated strong agreement with those of a human expert, with a mean difference of ±3.77° and a Pearson correlation coefficient of r = 0.983. This represents a significant departure from traditional workflows reliant on fluoroscopic or radiographic imaging for alignment assessment. In parallel, Medivis Inc. obtained FDA clearance in April 2025 for a new Spine Navigation system that boasts “AI segmentation”[125]. While technical details remain limited and no peer-reviewed studies have been published to date, the platform reportedly enables holographic navigation during spine surgery.
To complement these surgical intelligence platforms, commercial AR systems are being introduced to the operative suite. The xVision system, developed by Augmedics and recently acquired by VB Spine, was the first FDA-approved AR navigation system, with the initial cadaveric proof-of-concept demonstrating a reported placement accuracy of 96.7% and non-inferiority to the baseline computer-navigated pedicle placement[125,126]. Medivis’ Spine Navigation system also now claims AR-enabled instrumentation with universal tool adaptors that can be utilized across open and minimally invasive surgical (MIS) procedures[125].
Collectively, these commercial developments mark a pivotal transition in which computer vision, once largely confined to research settings, is now integrated directly into the operative environment. While published, peer-reviewed validation remains nascent for many of these platforms, their rapid commercial emergence suggests that the operating room of the next decade may look fundamentally different, with computer vision providing a continuous, intelligent layer to augment human surgical expertise.
CHALLENGES & LIMITATIONS
Though the integration of computer vision in spine surgery promises wide-ranging advancements, several barriers remain before we can see seamless incorporation into clinical practice. A recurring methodological limitation across the body of literature described above is the predominance of retrospective studies derived from single institutional cohorts, constraining the generalizability of reported performance metrics and underscoring the need for prospective, multicenter studies. This constraint is a manifestation of the "data problem" that models may only be as good as the datasets used to train them. Data availability is a common bottleneck for medical classification models, as even the larger publicly available datasets of spinal imaging are small within the larger context of AI[31,127]. This issue is even greater for intraoperative models, where there remains a significant dearth of surgical videos available for training, particularly within spine surgery and neurosurgery. However, the growing adoption of commercial imaging platforms, such as Medtronic UNiD and ATEC EOS Insight, means that industry partners can now accumulate large volumes of high-quality imaging and surgical data[119,128]. This trend presents a significant opportunity to enhance model performance, an impact that can be accelerated through industry-academic collaboration and the development of open-access datasets.
Other than the most straightforward approach of amassing more data, which can be a labor-intensive process especially for surgical video repositories, researchers have developed novel techniques to address challenges with data availability. One commonly used method is to artificially “augment” datasets through rotation, zooming, blurring, and mirroring of images in the original training set[129]. Another recent and powerful development is transfer learning, which involves using existing models pre-trained on massive datasets of general images and then fine-tuning these models on medical data. Models such as VGG-Net, Inception, and ResNet variants have already been leveraged for tasks such as retinal disease detection and brain tumor segmentation[130,131]. These techniques are also being implemented in spine surgery, although they are still limited in their ability to overcome the inherent sampling biases within the training data.
When patients in training data do not accurately reflect the population on which the model will be later evaluated, performance can drop precipitously. When working with institutional clinical datasets, participants may be disproportionately represented by those who are easy to find and recruit, or biased toward specific sociodemographic groups. This problem is somewhat mitigated with increased sample sizes, such as when leveraging electronic health record (EHR) data across large healthcare institutions, but even large, multicenter trials have demonstrated bias due to the systemic underrepresentation of women and ethnic minorities in receiving medical services. For example, one study by Seyyed-Kalantari et al. showed that SOTA AI models consistently underdiagnosed chest radiograph pathologies in patients from underserved populations[132]. These findings occurred despite models being trained on a large dataset comprising millions of images and hundreds of thousands of patients. Similarly, a systematic review by Loftus et al. identified the potential for bias across sociodemographic groups when using AI to support surgical patients[133]. This underrepresentation in data can have targeted and damaging effects on model performance. Beyond patient characteristics, differences in data collection can introduce bias, such as the use of different brands of endoscopic cameras or MRI/CT scanners[134]. Careful preprocessing can homogenize the input data, but even slight differences can propagate down the pipeline to introduce idiosyncratic features.
At the point of care, algorithm performance may be further degraded by intraoperative conditions that are difficult to anticipate from retrospective training data. Patient positioning can alter spinal alignment relative to preoperative imaging[135], while surgical smoke, bleeding, and irrigation fluid can obscure anatomical landmarks critical to real-time scene understanding[136]. Even the presence of implanted hardware, a challenge previously discussed in the context of postoperative alignment monitoring above, can introduce beam-hardening artifacts and landmark occlusion that degrades detection of spinopelvic parameters. In endoscopic approaches, these difficulties are further compounded by lens fogging and variable illumination.
The translation of computer vision models from research into routine clinical use poses additional practical and operational barriers. One fundamental obstacle is technical interoperability, as diagnostic data often remains siloed within disparate EHR and Picture Archiving and Communication Systems (PACS) that were not designed to accommodate AI-generated outputs[137]. Achieving seamless integration across platforms requires conformance to interoperability standards such as DICOM (Digital Imaging and Communications in Medicine) and FHIR (Fast Healthcare Interoperability Resources)[137]. This fragmentation is a meaningful limitation in the context of the spine surgery workflow, where we can imagine that outputs from a preoperative deformity-planning model may lack a standardized pathway to interface with intraoperative navigation systems or postoperative monitoring dashboards.
Implementation is also significantly shaped by a multitude of ethical questions and regulatory frameworks. Physicians must be able to understand and explain algorithmic decisions from patient imaging, while maintaining clear protocols for when to override automated designations. Trustworthiness of AI algorithms is thus a legitimate concern that extends beyond mere technical performance metrics to encompass issues of consent, interpretability, and clinical accountability[138]. In response to these challenges, new efforts have been made to standardize the robust clinical evaluation of AI models for computer vision. CARES is a recently proposed framework for benchmarking medical vision language models across categories of “trustworthiness, fairness, safety, privacy, and robustness”[139]. An initial assessment by the researchers of SOTA models, including LLaVa-Med and RadFM (Radiology Foundation Model), on the CARES benchmark revealed shortcomings in trustworthiness and fairness disparities across demographic groups. Further work is thus needed to rigorously test models, ensuring their safe and ethical use in clinical practice. Moreover, future efforts must ensure that the deployment of these technologies does not inadvertently formalize or exacerbate existing healthcare disparities.
At the regulatory level, the FDA now reviews “AI/ML-enabled medical devices” before their commercial use. This classification subjects these tools to the FDA’s testing and approval processes, ideally ensuring they meet established safety and efficacy standards before deployment. This includes gaining approval through premarket approval, 510(k), or De Novo regulatory pathways, depending on the risk associated with the computer vision application[140]. Furthermore, all AI models must use de-identified data or obtain explicit patient consent to use identifiable data in accordance with the Health Insurance Portability and Accountability Act (HIPAA). However, given the rapid pace of AI innovation, FDA oversight must be complemented by coordinated efforts across the healthcare ecosystem, with all stakeholders approaching these technologies with the care, rigor, and shared responsibility that they demand[141]. At the legislative level, there has been limited clarity on how to govern the use of AI in clinical care. Recent bills proposed in states including New York and Georgia have sought to prohibit AI systems from serving as the sole decision-makers in healthcare, reflecting growing concerns about algorithmic accountability[142,143]. In contrast, the federal 2025 Healthy Technology Act, introduced in the U.S. House of Representatives, would permit FDA-approved AI tools to prescribe medications[144]. These divergent approaches underscore the current lack of consensus and regulatory certainty around how AI technologies should be integrated into medical decision-making. At a time when computer vision applications are increasingly being used to guide diagnosis and operative decision-making in spine surgery, more explicit guidance will be essential to support the safe integration of these applications into high-stakes surgical workflows.
Finally, the financial and institutional infrastructure required for AI deployment can pose additional barriers, particularly for smaller practices and resource-limited settings. AI platforms are not free, and capital expenditures are required for initial acquisition, long-term technical maintenance, and specialized personnel. Substantial upfront investment may be difficult to justify in the absence of clear reimbursement pathways, and the reimbursement landscape for computer vision tools continues to evolve[145]. While the Centers for Medicare & Medicaid Services (CMS) has begun developing payment policies and Current Procedural Terminology (CPT) codes for specific AI-enabled services, the absence of established coverage determinations often requires institutions to absorb deployment costs without a defined path to cost recovery[146,147].
As similarly observed with established technologies such as spine navigation and robotics, the value proposition of AI must be underpinned by robust outcomes data to justify widespread implementation. Currently, the financial case for deploying computer vision tools in spine surgery remains largely unestablished, as formal cost-effectiveness analyses are strikingly sparse. A 2025 systematic review by Brin et al. identified only ten studies with formal economic analyses across the general radiology literature, all of which relied on theoretical modeling rather than real-world data[148]. Within spine surgery specifically, the literature is even more limited. We identified a cost-utility analysis from Hiligsmann et al. of AI-based opportunistic screening for osteoporotic vertebral compression fractures, estimating an incremental cost-effectiveness ratio of $72,085 per quality-adjusted life year gained for women over 50[149]. However, opportunistic screening inherently diverges from the perioperative applications central to this review. Addressing these gaps, whether by validating computer vision in prospective multicenter cohorts, establishing interoperability standards, or generating real-world economic evidence, will be essential prerequisites for translating the performance benchmarks described in this review into meaningful improvements in surgical care.
CONCLUSION
The rapid evolution of computer vision is poised to transform spine surgery. Advances in model architectures and the release of large-scale datasets are enabling faster, more accurate analysis of patient data. The resulting techniques will continue to enhance diagnostic precision, streamline alignment measurement, and improve prognostic modeling. However, the impact of computer vision extends beyond operative planning, as real-time navigation systems, marker-less registration, and AR platforms are already being integrated into intraoperative workflows. These tools are enabling new levels of surgical precision and safety, laying the foundation for a future in which intelligent, image-driven systems support every stage of spine surgery. While this progress is promising, important concerns remain, particularly around data privacy and algorithmic bias. Rigorous clinical validation will become increasingly necessary to ensure the safe and equitable adoption of computer vision technologies, especially given the high stakes associated with surgical decision-making. Looking ahead, the future of spine surgery will likely be shaped by multimodal systems capable of integrating complex visual and clinical data streams across the operative workflow. With continued investment in infrastructure, open-access datasets, and cross-disciplinary collaboration, computer vision will play an essential role in a new era of data-driven surgical practice.
DECLARATIONS
Authors’ contributions
Made substantial contributions to the overall design of the study: Yang E, Howell HJ, Chou D, Chan AK
Made substantial contributions to the review of the existing literature: Yang E, Howell HJ, Schonfeld E, Fuller J, Ayloo B
Made substantial contributions to the writing of this manuscript: Yang E, Howell HJ, Schonfeld E, Fuller J, Khan F, Ayloo B, Izima C, Sampath SG, Tang AJ, Rolfe NW, Green T, Chou D, Chan AK
Provided supervision, critical feedback, and guidance as senior author: Chou D, Chan AK
All authors reviewed and approved the final version of the manuscript.
Availability of data and materials
Not applicable.
AI and AI-assisted tools statement
Not applicable.
Financial support and sponsorship
None.
Conflicts of interest
Chou D reported personal fees from Globus and Medtronic outside the submitted work. Chan AK reported grants from the Cervical Spine Research Society and personal fees from ATEC, Carlsmed, and Spineart outside the submitted work.
All other authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Weiss AJ, Elixhauser A, Andrews RM. Characteristics of operating room procedures in U.S. hospitals, 2011. In: Healthcare Cost and Utilization Project (HCUP) Statistical Briefs. Rockville (MD): Agency for Healthcare Research and Quality (US); 2006. Available from https://pubmed.ncbi.nlm.nih.gov/24716251/ [accessed 16 April 2026].
2. Martin BI, Mirza SK, Spina N, Spiker WR, Lawrence B, Brodke DS. Trends in lumbar fusion procedure rates and associated hospital costs for degenerative spinal diseases in the United States, 2004 to 2015. Spine 2019;44:369-76.
3. Philipp LR, Leibold A, Mahtabfar A, Montenegro TS, Gonzalez GA, Harrop JS. Achieving value in spine surgery: 10 major cost contributors. Global Spine J 2021;11:14S-22S.
4. Mascagni P, Alapatt D, Sestini L, et al. Computer vision in surgery: from potential to clinical value. NPJ Digit Med 2022;5:163.
5. Maier-Hein L, Wagner M, Ross T, et al. Heidelberg colorectal data set for surgical data science in the sensor operating room. Sci Data 2021;8:101.
6. Zia A, Berniker M, Nespolo RG, et al. Intuitive surgical SurgToolLoc challenge results: 2022-2023. arXiv 2023;arXiv:2305.07152. Available from: http://arxiv.org/abs/2305.07152 [accessed 16 April 2026].
7. Wybier M, Bossard P. Musculoskeletal imaging in progress: the EOS imaging system. Joint Bone Spine 2013;80:238-43.
8. Santiago F, Láinez Ramos-Bossini AJ, Wáng YXJ, Martínez Barbero JP, García Espinosa J, Martínez Martínez A. The value of magnetic resonance imaging and computed tomography in the study of spinal disorders. Quant Imaging Med Surg 2022;12:3947-86.
9. Chai J, Zeng H, Li A, Ngai EW. Deep learning in computer vision: a critical review of emerging techniques and application scenarios. Mach Learn Appl 2021;6:100134.
10. Han K, Wang Y, Chen H, et al. A survey on vision transformer. IEEE Trans Pattern Anal Mach Intell 2023;45:87-110.
11. Esteva A, Chou K, Yeung S, et al. Deep learning-enabled medical computer vision. NPJ Digit Med 2021;4:5.
12. Minaee S, Boykov Y, Porikli F, Plaza A, Kehtarnavaz N, Terzopoulos D. Image segmentation using deep learning: a survey. arXiv 2020;arXiv:2001.05566. Available from: http://arxiv.org/abs/2001.05566 [accessed 16 April 2026].
13. Zhou N, Wen H, Wang Y, Liu Y, Zhou L. Review of deep learning models for spine segmentation. In: Proceedings of the 2022 International Conference on Multimedia Retrieval; 2022 Jun 27-30; Newark, NJ, USA. New York: ACM; 2022. pp. 498-507.
14. Bhairnallykar ST, Narawade V. Exploration of image segmentation: a comprehensive review of traditional segmentation techniques. In: 2023 7th International Conference on Electronics, Materials Engineering & Nano-Technology (IEMENTech); 2023 Dec 18-20; Kolkata, India. New York: IEEE; 2023. pp. 1-6.
15. Zamora G, Sari-Sarraf H, Long LR. Hierarchical segmentation of vertebrae from X-ray images. In: Sonka M, Fitzpatrick JM, Editors. Medical Imaging 2003: Image Processing; 2003; San Diego, CA, USA. Bellingham: SPIE; 2003.
16. Koompairojn S, Hua KA, Bhadrakom C. Automatic classification system for lumbar spine X-ray images. In: 19th IEEE Symposium on Computer-Based Medical Systems (CBMS’06); 2006 Jun 22-23; Salt Lake City, UT, USA. New York: IEEE; 2006. pp. 213-8.
17. Lecron F, Benjelloun M, Mahmoudi S. Cervical spine mobility analysis on radiographs: a fully automatic approach. Comput Med Imaging Graph 2012;36:634-42.
18. Li Y, Liang W, Zhang Y, An H, Tan J. Automatic lumbar vertebrae detection based on feature fusion deep learning for partial occluded C-arm X-ray images. Annu Int Conf IEEE Eng Med Biol Soc 2016;2016:647-50.
19. Al Arif SMMR, Knapp K, Slabaugh G. Fully automatic cervical vertebrae segmentation framework for X-ray images. Comput Methods Programs Biomed 2018;157:95-111.
20. Cho BH, Kaji D, Cheung ZB, et al. Automated measurement of lumbar lordosis on radiographs using machine learning and computer vision. Global Spine J 2020;10:611-8.
21. Shin Y, Han K, Lee YH. Temporal trends in cervical spine curvature of South Korean adults assessed by deep learning system segmentation, 2006-2018. JAMA Netw Open 2020;3:e2020961.
22. Kim KC, Cho HC, Jang TJ, Choi JM, Seo JK. Automatic detection and segmentation of lumbar vertebrae from X-ray images for compression fracture evaluation. Comput Methods Programs Biomed 2021;200:105833.
23. Chen Y, Mo Y, Readie A, et al. VertXNet: an ensemble method for vertebral body segmentation and identification from cervical and lumbar spinal X-rays. Sci Rep 2024;14:3341.
24. Bertels J, Eelbode T, Berman M, et al. Optimizing the Dice score and Jaccard index for medical image segmentation: theory and practice. In: Shen D, Liu T, Peters TM, Staib LH, Essert C, Zhou S, Yap PT, Khan A, Editors. Medical Image Computing and Computer Assisted Intervention - MICCAI 2019; 2019 Oct 13-17; Shenzhen, China. Cham: Springer; 2019. pp. 92-100.
25. Ronneberger O, Fischer P, Brox T. U-Net: convolutional networks for biomedical image segmentation. arXiv 2015;arXiv:1505.04597. Available from: http://arxiv.org/abs/1505.04597 [accessed 16 April 2026].
26. Janssens R, Zeng G, Zheng G. Fully automatic segmentation of lumbar vertebrae from CT images using cascaded 3D Fully Convolutional Networks. arXiv 2017;arXiv:1712.01509. Available from: http://arxiv.org/abs/1712.01509 [accessed 16 April 2026].
27. Lessmann N, van Ginneken B, de Jong PA, Išgum I. Iterative fully convolutional neural networks for automatic vertebra segmentation and identification. Med Image Anal 2019;53:142-55.
28. Sekuboyina A, Husseini ME, Bayat A, et al. VerSe: a Vertebrae labelling and segmentation benchmark for multi-detector CT images. Med Image Anal 2021;73:102166.
29. Payer C, Štern D, Bischof H, Urschler M. Coarse to fine vertebrae localization and segmentation with SpatialConfiguration-net and U-net. In: Farinella GM, Radeva P, Braz J, Editors. Proceedings of the 15th International Joint Conference on Computer Vision, Imaging and Computer Graphics Theory and Applications; 2020 Feb 27-29; Valletta, Malta. Setúbal: SciTePress; 2020. pp. 124-33.
30. Tao R, Liu W, Zheng G. Spine-transformers: vertebra labeling and segmentation in arbitrary field-of-view spine CTs via 3D transformers. Med Image Anal 2022;75:102258.
31. van der Graaf JW, van Hooff ML, Buckens CFM, et al. Lumbar spine segmentation in MR images: a dataset and a public benchmark. Sci Data 2024;11:264.
32. Zhang Z, Liu T, Fan G, et al. SpineMamba: enhancing 3D spinal segmentation in clinical imaging through residual visual Mamba layers and shape priors. Comput Med Imaging Graph 2025;123:102531.
33. Liebl H, Schinz D, Sekuboyina A, et al. A computed tomography vertebral segmentation dataset with anatomical variations and multi-vendor scanner data. Sci Data 2021;8:284.
34. Sa R, Owens W, Wiegand R, et al. Intervertebral disc detection in X-ray images using faster R-CNN. Annu Int Conf IEEE Eng Med Biol Soc 2017;2017:564-7.
35. Lemay A, Gros C, Zhuo Z, et al. Automatic multiclass intramedullary spinal cord tumor segmentation on MRI with deep learning. Neuroimage Clin 2021;31:102766.
36. Li X, Hong Y, Xu Y, Hu M. VerFormer: vertebrae-aware transformer for automatic spine segmentation from CT images. Diagnostics 2024;14:1859.
37. Kirillov A, Mintun E, Ravi N, et al. Segment anything. arXiv 2023;arXiv:2304.02643. Available from: http://arxiv.org/abs/2304.02643 [accessed 16 April 2026].
38. Ma J, He Y, Li F, Han L, You C, Wang B. Segment anything in medical images. Nat Commun 2024;15:654.
39. Seoud L, Adankon MM, Labelle H, Dansereau J, Cheriet F. Prediction of scoliosis curve type based on the analysis of trunk surface topography. In: 2010 IEEE International Symposium on Biomedical Imaging: From Nano to Macro; 2010 Apr 14-17; Rotterdam, Netherlands. New York: IEEE; 2010. pp. 408-11.
40. Lin Y, Zhou H, Ma K, Yang X, Zheng Y. Seg4Reg networks for automated spinal curvature estimation. In: Cai Y, Wang L, Audette M, Zheng G, Li S, Editors. Proceedings of the 7th International Workshop and Challenge on Computational Methods and Clinical Applications for Spine Imaging, CSI 2019; 2019 Oct 17; Shenzhen, China. Cham: Springer; 2020. pp. 69-74.
41. Chen K, Peng C, Li Y, Cheng D, Wei S. Accurate automated keypoint detections for spinal curvature estimation. In: Cai Y, Wang L, Audette M, Zheng G, Li S, Editors. Proceedings of the 7th International Workshop and Challenge on Computational Methods and Clinical Applications for Spine Imaging, CSI 2019; 2019 Oct 17; Shenzhen, China. Cham: Springer; 2020. pp. 63-8.
42. Dubost F, Collery B, Renaudier A, et al. Automated estimation of the spinal curvature via spine centerline extraction with ensembles of cascaded neural networks. In: Cai Y, Wang L, Audette M, Zheng G, Li S, Editors. Proceedings of the 7th International Workshop and Challenge on Computational Methods and Clinical Applications for Spine Imaging, CSI 2019; 2019 Oct 17; Shenzhen, China. Cham: Springer; 2020. pp. 88-94.
43. Lin Y, Liu L, Ma K, Zheng Y. Seg4Reg+: consistency learning between spine segmentation and Cobb angle regression. arXiv 2022;arXiv:2208.12462. Available from: http://arxiv.org/abs/2208.12462 [accessed 16 April 2026].
44. Alukaev D, Kiselev S, Mustafaev T, Ainur A, Ibragimov B, Vrtovec T. A deep learning framework for vertebral morphometry and Cobb angle measurement with external validation. Eur Spine J 2022;31:2115-24.
45. Chen K, Stotter C, Klestil T, Mitterer JA, Lepenik C, Nehrer S. Fully automated measurement of Cobb angles in coronal plane spine radiographs. J Clin Med 2024;13:4122.
46. Orosz LD, Bhatt FR, Jazini E, et al. Novel artificial intelligence algorithm: an accurate and independent measure of spinopelvic parameters. J Neurosurg Spine 2022;37:893-901.
47. Harake ES, Linzey JR, Jiang C, et al. Development and validation of an artificial intelligence model to accurately predict spinopelvic parameters. J Neurosurg Spine 2024;41:88-96.
48. Joshi RS, Harake ES, Jiang C, et al. Validation of a novel artificial intelligence model (SpinePose) to automatically and accurately predict spinopelvic parameters using scoliosis radiographs in an external cohort. Neurosurg Focus 2025;58:E10.
49. Kang DH, Jeong YJ, Kim ST, et al. Automated measurement of pelvic parameters using convolutional neural network in complex spinal deformities: overcoming challenges in coronal deformity cases. Spine J 2025;25:1688-97.
50. Grover P, Siebenwirth J, Caspari C, et al. Can artificial intelligence support or even replace physicians in measuring sagittal balance? Eur Spine J 2022;31:1943-51.
51. Suri A, Tang S, Kargilis D, et al. Conquering the Cobb angle: a deep learning algorithm for automated, hardware-invariant measurement of Cobb angle on radiographs in patients with scoliosis. Radiol Artif Intell 2023;5:e220158.
52. Löchel J, Putzier M, Dreischarf M, et al. Deep learning algorithm for fully automated measurement of sagittal balance in adult spinal deformity. Eur Spine J 2024;33:4119-24.
53. Vogt S, Scholl C, Grover P, et al. Novel AI-based algorithm for the automated measurement of cervical sagittal balance parameters. A validation study on pre- and postoperative radiographs of 129 patients. Global Spine J 2025;15:1155-65.
54. Derkatch S, Kirby C, Kimelman D, Jozani MJ, Davidson JM, Leslie WD. Identification of vertebral fractures by convolutional neural networks to predict nonvertebral and hip fractures: a registry-based cohort study of dual X-ray absorptiometry. Radiology 2019;293:405-11.
55. Li YC, Chen HH, Horng-Shing Lu H, Hondar Wu HT, Chang MC, Chou PH. Can a deep-learning model for the automated detection of vertebral fractures approach the performance level of human subspecialists? Clin Orthop Relat Res 2021;479:1598-612.
56. Shen L, Gao C, Hu S, et al. Using artificial intelligence to diagnose osteoporotic vertebral fractures on plain radiographs. J Bone Miner Res 2023;38:1278-87.
57. Liu B, Jin Y, Feng S, Yu H, Zhang Y, Li Y. Benign vs malignant vertebral compression fractures with MRI: a comparison between automatic deep learning network and radiologist’s assessment. Eur Radiol 2023;33:5060-8.
58. Jamaludin A, Kadir T, Zisserman A. SpineNet: automated classification and evidence visualization in spinal MRIs. Med Image Anal 2017;41:63-73.
59. Merali Z, Wang JZ, Badhiwala JH, Witiw CD, Wilson JR, Fehlings MG. A deep learning model for detection of cervical spinal cord compression in MRI scans. Sci Rep 2021;11:10473.
60. Hallinan JTPD, Zhu L, Yang K, Makmur A, Algazwi DAR, Thian YL, et al. Deep learning model for automated detection and classification of central canal, lateral recess, and neural foraminal stenosis at lumbar spine MRI. Radiology 2021;300:130-8.
61. Nguyen HT, Pham HH, Nguyen NT, et al. VinDr-SpineXR: a deep learning framework for spinal lesions detection and classification from radiographs. arXiv 2021;arXiv:2106.12930. Available from: http://arxiv.org/abs/2106.12930 [accessed 16 April 2026].
62. Maki S, Furuya T, Horikoshi T, et al. A deep convolutional neural network with performance comparable to radiologists for differentiating between spinal schwannoma and meningioma. Spine 2020;45:694-700.
63. Haim O, Agur A, Gabay S, et al. Differentiating spinal pathologies by deep learning approach. Spine J 2024;24:297-303.
64. Zhuo Z, Zhang J, Duan Y, et al. Automated classification of intramedullary spinal cord tumors and inflammatory demyelinating lesions using deep learning. Radiol Artif Intell 2022;4:e210292.
65. Huang KT, Silva MA, See AP, et al. A computer vision approach to identifying the manufacturer and model of anterior cervical spinal hardware. J Neurosurg Spine 2019;31:844-50.
66. Yang HS, Kim KR, Kim S, Park JY. Deep learning application in spinal implant identification. Spine 2021;46:E318-24.
67. Dutt R, Mendonca D, Phen HM, et al. Automatic localization and brand detection of cervical spine hardware on radiographs using weakly supervised machine learning. Radiol Artif Intell 2022;4:e210099.
68. Park S, Kim JK, Chang MC, Park JJ, Yang JJ, Lee GW. Assessment of fusion after anterior cervical discectomy and fusion using convolutional neural network algorithm. Spine 2022;47:1645-50.
69. Schwartz JT, Valliani AA, Arvind V, et al. Identification of anterior cervical spinal instrumentation using a smartphone application powered by machine learning. Spine 2022;47:E407-14.
70. Anand A, Flores AR, McDonald MF, Gadot R, Xu DS, Ropper AE. A computer vision approach to identifying the manufacturer of posterior thoracolumbar instrumentation systems. J Neurosurg Spine 2023;38:417-24.
71. Chun KS, Lee S, Choi H, Park H, Lee S, Jung JY. Patch-wise approach with vision transformer for detecting implant failure in spinal radiography. J Imaging Inform Med 2025.
72. Sun H, Tang W, Deng L, et al. Development and validation of interpretable machine learning models incorporating paraspinal muscle quality to predict cage subsidence risk following posterior lumbar interbody fusion. Spine 2025;50:1375-85.
73. Waranusast R, Riyamongkol P, Weerakul S, Chaibhuddanugul N, Laoruengthana A, Mahatthanatrakul A. Fully automated pedicle screw manufacturer identification in plain radiograph with deep learning methods. Eur Spine J 2025;34:3940-9.
74. De Silva T, Vedula SS, Perdomo-Pantoja A, et al. SpineCloud: image analytics for predictive modeling of spine surgery outcomes. J Med Imaging 2020;7:031502.
75. Saravi B, Zink A, Ülkümen S, et al. Clinical and radiomics feature-based outcome analysis in lumbar disc herniation surgery. BMC Musculoskelet Disord 2023;24:791.
76. Wu J, Li J, Zhang H, Wu L, Shen X, Lv W. Predicting functional outcome after open lumbar fusion surgery: a retrospective multicenter cohort study. Eur J Radiol 2025;182:111836.
77. Durand WM, Lafage R, Hamilton DK, et al.; International Spine Study Group (ISSG). Artificial intelligence clustering of adult spinal deformity sagittal plane morphology predicts surgical characteristics, alignment, and outcomes. Eur Spine J 2021;30:2157-66.
78. Rudisill SS, Hornung AL, Barajas JN, et al. Artificial intelligence in predicting early-onset adjacent segment degeneration following anterior cervical discectomy and fusion. Eur Spine J 2022;31:2104-14.
79. Johnson GW, Chanbour H, Ali MA, et al. Artificial intelligence to preoperatively predict proximal junction kyphosis following adult spinal deformity surgery: soft tissue imaging may be necessary for accurate models. Spine 2023;48:1688-95.
80. Brigato P, Vadalà G, De Salvatore S, et al. Harnessing machine learning to predict and prevent proximal junctional kyphosis and failure in adult spinal deformity surgery: a systematic review. Brain Spine 2025;5:104273.
81. Zheng B, Yu P, Ma K, Zhu Z, Liang Y, Liu H. Radiomics-based machine learning model integrating preoperative vertebral computed tomography and clinical features to predict cage subsidence after single-level anterior cervical discectomy and fusion with a zero-profile anchored spacer. J Orthop Surg Res 2025;20:851.
82. Bishara A, Patel S, Warman A, et al. Artificial intelligence automated measurements of spinopelvic parameters in adult spinal deformity - a systematic review. Spine Deform 2025;13:1289-304.
83. Smith JS, Shaffrey CI, Ames CP, et al.; International Spine Study Group. Assessment of symptomatic rod fracture after posterior instrumented fusion for adult spinal deformity. Neurosurgery 2012;71:862-7.
84. Hamilton DK, Buza JA 3rd, Passias P, et al.; International Spine Study Group. The fate of patients with adult spinal deformity incurring rod fracture after thoracolumbar fusion. World Neurosurg 2017;106:905-11.
85. Schwab FJ, Blondel B, Bess S, et al.; International Spine Study Group (ISSG). Radiographical spinopelvic parameters and disability in the setting of adult spinal deformity: a prospective multicenter analysis. Spine 2013;38:E803-12.
86. Divi SN, Goyal DKC, Bowles DR, et al. How do spinopelvic parameters influence patient-reported outcome measurements after lumbar decompression? Spine J 2020;20:1610-7.
87. Ye J, Gupta S, Farooqi AS, et al. Predictive role of global spinopelvic alignment and upper instrumented vertebra level in symptomatic proximal junctional kyphosis in adult spinal deformity. J Neurosurg Spine 2023;39:774-84.
88. Wondra JP 2nd, Kelly MP, Greenberg J, et al. Validation of adult spinal deformity surgical outcome prediction tools in adult symptomatic lumbar scoliosis. Spine 2023;48:21-8.
89. Schönnagel L, Caffard T, Vu-Han TL, et al. Predicting postoperative outcomes in lumbar spinal fusion: development of a machine learning model. Spine J 2024;24:239-49.
90. Parisien A, Wai EK, ElSayed MSA, Frei H. Subsidence of spinal fusion cages: a systematic review. Int J Spine Surg 2022;16:1103-18.
91. Bui TT, Joseph K, Yahanda AT, Vogl S, Ruiz-Cardozo M, Molina CA. How much variance exists among published definitions of proximal junctional kyphosis? J Clin Med 2025;14:5469.
92. Wilson JP Jr, Fontenot L, Stewart C, Kumbhare D, Guthikonda B, Hoang S. Image-guided navigation in spine surgery: from historical developments to future perspectives. J Clin Med 2024;13:2036.
93. Gebhard FT, Kraus MD, Schneider E, Liener UC, Kinzl L, Arand M. Does computer-assisted spine surgery reduce intraoperative radiation doses? Spine 2006;31:2024-7; discussion 2028.
94. Amiot LP, Lang K, Putzier M, Zippel H, Labelle H. Comparative results between conventional and computer-assisted pedicle screw installation in the thoracic, lumbar, and sacral spine. Spine 2000;25:606-14.
95. Verma R, Krishan S, Haendlmayer K, Mohsen A. Functional outcome of computer-assisted spinal pedicle screw placement: a systematic review and meta-analysis of 23 studies including 5,992 pedicle screws. Eur Spine J 2010;19:370-5.
96. Liebmann F, von Atzigen M, Stütz D, et al. Automatic registration with continuous pose updates for marker-less surgical navigation in spine surgery. Med Image Anal 2024;91:103027.
97. Abumoussa A, Gopalakrishnan V, Succop B, et al. Machine learning for automated and real-time two-dimensional to three-dimensional registration of the spine using a single radiograph. Neurosurg Focus 2023;54:E16.
98. Doerr SA, Uneri A, Huang Y, et al. Data-driven detection and registration of spine surgery instrumentation in intraoperative images. Proc SPIE Int Soc Opt Eng 2020;11315:113152P.
99. Burström G, Buerger C, Hoppenbrouwers J, et al. Machine learning for automated 3-dimensional segmentation of the spine and suggested placement of pedicle screws based on intraoperative cone-beam computer tomography. J Neurosurg Spine 2019;31:147-54.
100. Jecklin S, Jancik C, Farshad M, Fürnstahl P, Esfandiari H. X23D-intraoperative 3D lumbar spine shape reconstruction based on sparse multi-view X-ray data. J Imaging 2022;8:271.
101. Luchmann D, Jecklin S, Cavalcanti NA, et al. Spinal navigation with AI-driven 3D-reconstruction of fluoroscopy images: an ex-vivo feasibility study. BMC Musculoskelet Disord 2024;25:925.
102. Richards PJ, George J, Metelko M, Brown M. Spine computed tomography doses and cancer induction. Spine 2010;35:430-3.
103. van Stralen M, van der Kolk BYM, Zijlstra F, et al. BoneMRI of the cervical spine: deep learning-based radiodensity contrast generation for selective visualization of osseous structures. In: ISMRM 27th Annual Meeting and Exhibition; 2019 May 11-16; Montréal, Canada. Hoboken: John Wiley & Sons, Inc.; 2019. Available from https://ismrm.gitlab.io/2019/1142.html [accessed 16 April 2026].
104. van der Kolk BYMM, Slotman DJJ, Nijholt IM, et al. Bone visualization of the cervical spine with deep learning-based synthetic CT compared to conventional CT: a single-center noninferiority study on image quality. Eur J Radiol 2022;154:110414.
105. Staartjes VE, Seevinck PR, Vandertop WP, van Stralen M, Schröder ML. Magnetic resonance imaging-based synthetic computed tomography of the lumbar spine for surgical planning: a clinical proof-of-concept. Neurosurg Focus 2021;50:E13.
106. Davidar AD, Judy BF, Hersh AM, et al. Robot-assisted screw fixation in a cadaver utilizing magnetic resonance imaging-based synthetic computed tomography: toward radiation-free spine surgery. Illustrative case. J Neurosurg Case Lessons 2023;6:CASE23120.
107. Azad TD, Warman A, Tracz JA, Hughes LP, Judy BF, Witham TF. Augmented reality in spine surgery - past, present, and future. Spine J 2024;24:1-13.
108. Ghaednia H, Fourman MS, Lans A, et al. Augmented and virtual reality in spine surgery, current applications and future potentials. Spine J 2021;21:1617-25.
109. Pierzchajlo N, Stevenson TC, Huynh H, et al. Augmented reality in minimally invasive spinal surgery: a narrative review of available technology. World Neurosurg 2023;176:35-42.
110. Elmi-Terander A, Burström G, Nachabe R, et al. Pedicle screw placement using augmented reality surgical navigation with intraoperative 3D imaging: a first in-human prospective cohort study. Spine 2019;44:517-25.
111. Butler AJ, Colman MW, Lynch J, Phillips FM. Augmented reality in minimally invasive spine surgery: early efficiency and complications of percutaneous pedicle screw instrumentation. Spine J 2023;23:27-33.
112. Ma Y, Wu J, Dong Y, Tang H, Ma X. Augmented reality navigation system enhances the accuracy of spinal surgery pedicle screw placement: a randomized, multicenter, parallel-controlled clinical trial. Orthop Surg 2025;17:631-43.
113. Auloge P, Cazzato RL, Ramamurthy N, et al. Augmented reality and artificial intelligence-based navigation during percutaneous vertebroplasty: a pilot randomised clinical trial. Eur Spine J 2020;29:1580-9.
114. Carl B, Bopp M, Saß B, Nimsky C. Microscope-based augmented reality in degenerative spine surgery: initial experience. World Neurosurg 2019;128:e541-51.
115. McCloskey K, Turlip R, Ahmad HS, Ghenbot YG, Chauhan D, Yoon JW. Virtual and augmented reality in spine surgery: a systematic review. World Neurosurg 2023;173:96-107.
116. Cui P, Guo Z, Xu J, et al. Tissue recognition in spinal endoscopic surgery using deep learning. In: 2019 IEEE 10th International Conference on Awareness Science and Technology (iCAST); 2019 Oct 23-25; Morioka, Japan. New York: IEEE; 2019. pp 1-5.
117. Cui P, Shu T, Lei J, Chen W. Nerve recognition in percutaneous transforaminal endoscopic discectomy using convolutional neural network. Med Phys 2021;48:2279-88.
118. Peng S, Zhao P, Ye Y, Chen J, Chang Y, Zheng X. Spinal nerve segmentation method and dataset construction in endoscopic surgical scenarios. arXiv 2023;arXiv:2307.10955. Available from: http://arxiv.org/abs/2307.10955 [accessed 16 April 2026].
119. ATEC launches EOS insight, an end-to-end spine surgery platform powered by EOS imaging and AI. ATEC (Carlsbad, CA, USA), July 24, 2024. Available from https://investors.alphatecspine.com/press-releases/news-details/2024/ATEC-Launches-EOS-InsightTM-an-End-To-End-Spine-Surgery-Platform-Powered-by-EOS-Imaging-and-AI/default.aspx [accessed 16 April 2026].
120. Carlsmed. Technology Platform. Available from https://carlsmed.com/aprevo-technology-platform/ [accessed 16 April 2026].
121. Fiere V, Fuentès S, Burger E, et al. UNID Patient-Specific Rods show a reduction in rod breakage incidence. New York: Medicrea USA Corp. 2017. Available from https://api.semanticscholar.org/CorpusID:13695587 [accessed 16 April 2026].
122. Fernandes RJR, Gee A, Kanawati AJ, et al. Biomechanical comparison of subsidence between patient-specific and non-patient-specific lumbar interbody fusion cages. Global Spine J 2024;14:1155-63.
123. Sadrameli SS, Blaskiewicz DJ, Asghar J, et al. Predictability in achieving target intervertebral lordosis using personalized interbody implants. Int J Spine Surg 2024;18:S16-23.
124. Proprio - Technology. Available from https://www.propriovision.com/technology [accessed 16 April 2026].
125. Medivis receives FDA 510(k) clearance for spine navigation and announces commercial launch in the United States. Medivis (New York, NY, USA), April 02, 2025. Available from https://www.prnewswire.com/news-releases/medivis-receives-fda-510k-clearance-for-spine-navigation-and-announces-commercial-launch-in-the-united-states-302418834.html [accessed 16 April 2026].
126. Molina CA, Theodore N, Ahmed AK, et al. Augmented reality-assisted pedicle screw insertion: a cadaveric proof-of-concept study. J Neurosurg Spine 2019;31:139-46.
127. Deng Y, Wang C, Hui Y, et al. CTSpine1K: a large-scale dataset for spinal vertebrae segmentation in computed tomography. arXiv 2021;arXiv:2105.14711. Available from http://arxiv.org/abs/2105.14711 [accessed 16 April 2026].
128. Medtronic. UNiDTM adaptive spine intelligence. Available from: https://www.medtronic.com/en-us/healthcare-professionals/products/spinal-orthopedic/complex-spine/fixation-systems/unid-adaptive-spine-intelligence.html. [Last accessed on 16 Apr 2026].
129. Mumuni A, Mumuni F. Data augmentation: a comprehensive survey of modern approaches. Array 2022;16:100258.
130. Miladinović A, Biscontin A, Ajčević M, et al. Evaluating deep learning models for classifying OCT images with limited data and noisy labels. Sci Rep 2024;14:30321.
131. Basha NK, Ananth C, Muthukumaran K, Sudhamsu G, Mittal V, Gared F. Mask region-based convolutional neural network and VGG-16 inspired brain tumor segmentation. Sci Rep 2024;14:17615.
132. Seyyed-Kalantari L, Zhang H, McDermott MBA, Chen IY, Ghassemi M. Underdiagnosis bias of artificial intelligence algorithms applied to chest radiographs in under-served patient populations. Nat Med 2021;27:2176-82.
133. Loftus TJ, Altieri MS, Balch JA, et al. Artificial intelligence-enabled decision support in surgery: state-of-the-art and future directions. Ann Surg 2023;278:51-8.
134. Stanley EAM, Souza R, Wilms M, Forkert ND. Where, why, and how is bias learned in medical image analysis models? EBioMedicine 2025;111:105501.
135. Wessels L, Komm B, Bohner G, Vajkoczy P, Hecht N. Spinal alignment shift between supine and prone CT imaging occurs frequently and regardless of the anatomic region, risk factors, or pathology. Neurosurg Rev 2022;45:855-63.
136. Wang F, Sun X, Li J. Surgical smoke removal via residual Swin transformer network. Int J Comput Assist Radiol Surg 2023;18:1417-27.
137. Tejani AS, Cook TS, Hussain M, Sippel Schmidt T, O’Donnell KP. Integrating and adopting AI in the radiology workflow: a primer for standards and Integrating the Healthcare Enterprise (IHE) profiles. Radiology 2024;311:e232653.
138. Hasani N, Morris MA, Rhamim A, et al. Trustworthy artificial intelligence in medical imaging. PET Clin 2022;17:1-12.
139. Xia P, Chen Z, Tian J, Gong Y, Hou R, Xu Y, et al. CARES: a comprehensive benchmark of trustworthiness in medical vision language models. arXiv 2024;arXiv:2406.06007. Available from http://arxiv.org/abs/2406.06007 [accessed 16 April 2026].
140. Petrick N, Chen W, Delfino JG, et al. Regulatory considerations for medical imaging AI/ML devices in the United States: concepts and challenges. J Med Imaging 2023;10:051804.
141. Warraich HJ, Tazbaz T, Califf RM. FDA perspective on the regulation of artificial intelligence in health care and biomedicine. JAMA 2025;333:241-7.
142. Partisan Bill. Georgia House Bill 887. LegiScan. Available from: https://legiscan.com/GA/text/HB887/id/2877485. [Last accessed on 16 Apr 2026].
143. Gonzalez K. Senate Bill S7896. The New York State Senate. Available from: https://www.nysenate.gov/legislation/bills/2025/S7896. [Last accessed on 16 Apr 2026].
144. Schweikert D. Healthy Technology Act of 2025. Available from: https://www.congress.gov/bill/119th-congress/house-bill/238. [Last accessed on 16 Apr 2026].
145. Parikh RB, Helmchen LA. Paying for artificial intelligence in medicine. NPJ Digit Med 2022;5:63.
147. Dogra S, Silva EZ 3rd, Rajpurkar P. Reimbursement in the age of generalist radiology artificial intelligence. NPJ Digit Med 2024;7:350.
148. Brin D, Tau N. Cost-effectiveness of artificial intelligence tools in radiology: a systematic review. Eur Radiol 2025.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data








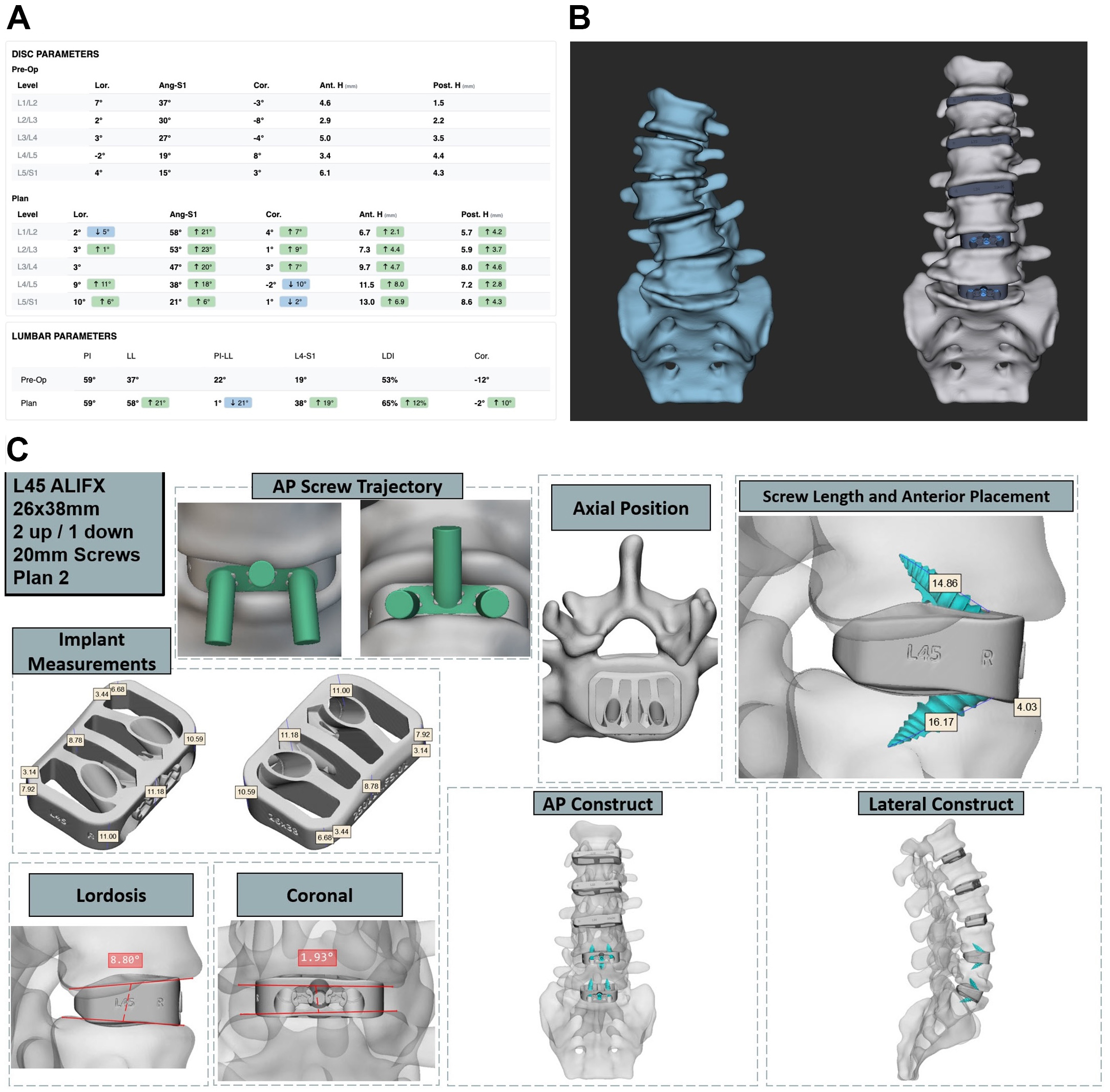
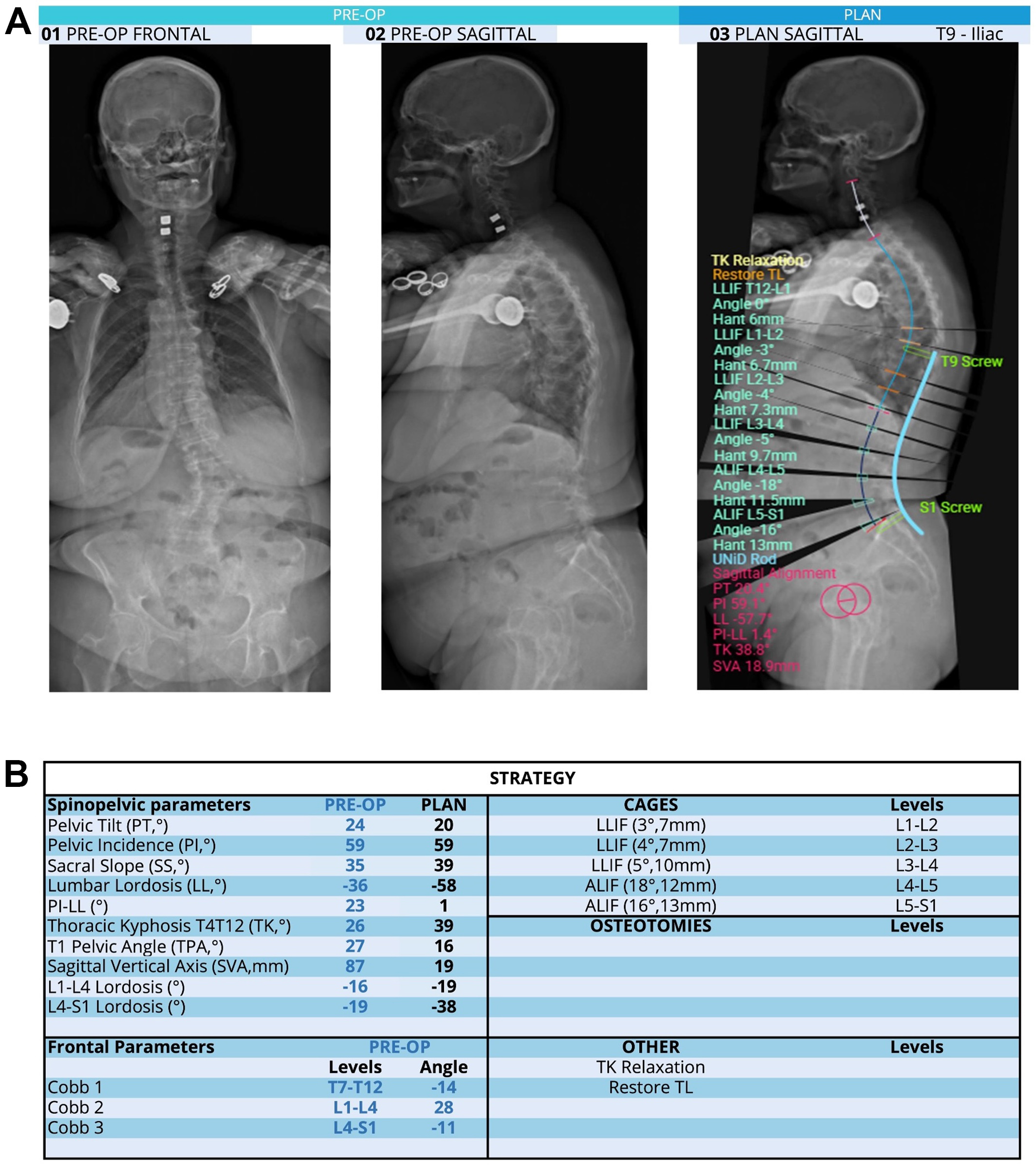
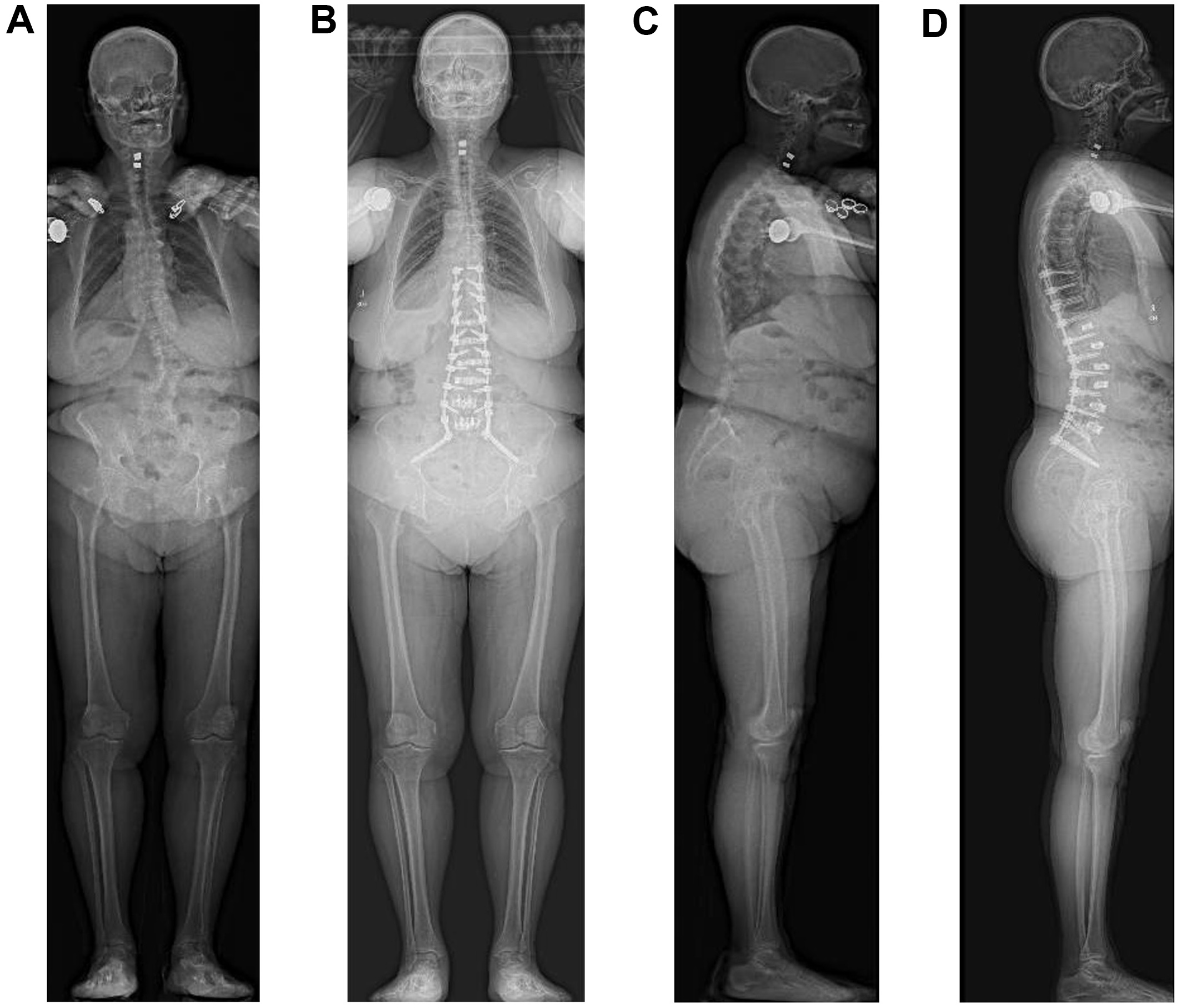











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].