Applications of speech analysis in diseases’ assessment, prediction and diagnosis: a scoping review
Abstract
Background: Speech production is a coordinated physiological process and a vital digital biomarker for health assessment. Recent advances in artificial intelligence (AI), particularly in representation learning, have substantially expanded the application of speech analysis across diverse clinical domains.
Methods: This review was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses Extension for Scoping Reviews (PRISMA-ScR). Five major bibliographic databases were systematically searched for studies published between 2015 and 2025. Eligible studies applied AI-driven speech analysis for clinical diagnosis or monitoring, while those lacking quantitative evaluation or sufficient methodological detail were excluded.
Results: A total of 124 studies were analyzed, covering neurological, psychiatric, and respiratory disorders. The field has transitioned from traditional machine learning with handcrafted features to deep learning and foundation models. Parkinson’s disease, Alzheimer’s disease, depression, and coronavirus disease 2019 (COVID-19) are the most frequently investigated conditions. The included studies were charted and synthesized to map disease coverage, methodological trends, and clinical application scenarios.
Conclusion: Speech analysis offers a non-invasive approach for early disease detection and remote monitoring in telemedicine. To support clinical translation, future research should prioritize model robustness and interpretability across diverse clinical populations.
Keywords
INTRODUCTION
Speech production constitutes a precisely coordinated physiological process. The larynx, vocal cords, oral cavity, nasal cavity, and other vocal organs, together with their subsystems, work in close coordination with respiratory muscle groups[1]. Functional abnormalities in any component directly manifest as alterations in speech patterns. Owing to this close coupling between speech production and physiological function, speech provides a unique, non-invasive, and information-rich window for disease assessment, prediction, and diagnosis. However, traditional speech assessment methods heavily rely on manual evaluation by specialized physicians. This approach is not only cumbersome and inefficient but also susceptible to subjective influences, leading to significant inter-observer variability and limited scalability in routine clinical practice.
In recent years, the rapid advancement of artificial intelligence (AI) technology has introduced new paradigms for disease assessment, prediction, and diagnosis[2]. As a result, speech has emerged as a highly promising digital health biomarker in the medical field. As an information-rich data source, speech possesses inherent advantages including ease of acquisition, non-invasiveness, and high information density, encompassing multi-dimensional features such as prosody, pitch, rhythm, and spectral characteristics. AI-driven analysis of speech signals enables sensitive and objective characterization of subtle acoustic changes that may reflect early disease manifestations, progression trajectories, or treatment responses. Speech characteristics such as loudness, pitch, speech rate, and pausing patterns are linked to emotional, cognitive, and pathological states[3]. These findings suggest that speech analysis supports not only diagnostic decision-making but also disease severity assessment and predictive modeling of health outcomes[4,5].
Driven by these methodological advances, speech-based approaches have been increasingly applied across a wide range of medical domains. In neurological disorders, speech analysis has been used for early screening, disease staging, and progression monitoring in Parkinson’s disease (PD) and Alzheimer’s disease (AD). In mental health, speech features have been explored for the assessment and prediction of depressive and anxiety disorders by modeling emotional tone, rhythm, and prosodic dynamics. In respiratory diseases, acoustic cues derived from speech, coughing, and breathing sounds have been investigated for disease screening and symptom monitoring in asthma and chronic obstructive pulmonary disease (COPD). Representative studies have demonstrated the feasibility and clinical relevance of these approaches across diverse disease contexts[6-8]. Collectively, these studies demonstrate that speech analysis has evolved beyond isolated diagnostic tasks to support longitudinal assessment, risk stratification, and outcome prediction across diverse clinical scenarios.
Despite these encouraging applications, existing research on speech-based disease analysis remains fragmented. Many studies focus on individual diseases, isolated feature representations, or specific modeling paradigms, while experimental settings, datasets, and evaluation protocols vary substantially. This fragmentation hampers cross-study comparability and limits the ability to derive unified insights into methodological robustness, clinical generalizability, and translational readiness across disease domains. As a result, there is a clear need for integrative syntheses that systematically connect speech features, AI methodologies, and clinical tasks across diseases.
From a technological perspective, AI methods for speech-based disease analysis have progressed from traditional machine learning (ML) to deep learning (DL), and more recently to foundation models. Traditional ML approaches, such as support vector machine (SVM)[9], random forest (RF)[10], and k-nearest neighbors (KNN)[11], rely on handcrafted acoustic features and demonstrate good interpretability on small-scale datasets[12,13]. However, manually engineered features often fail to adequately represent complex temporal dependencies and subtle pathological variations in real-world clinical speech analysis. This limitation is especially pronounced when analyzing spontaneous or continuous speech. DL models, including convolutional neural networks (CNNs)[14] and transformers[15], enable automated feature learning and have shown improved performance and robustness under noisy conditions[16]. Nevertheless, most DL approaches depend heavily on large labeled clinical datasets, which are difficult to obtain due to privacy constraints, annotation costs, and inter-patient heterogeneity. Recently, foundation models pretrained on large-scale unlabeled data [e.g., Generative Pre-trained Transformer (GPT)[17] and Bidirectional Encoder Representations from Transformers (BERT)[18]] have attracted increasing attention by enabling transferable representations, cross-task generalization, and data-efficient adaptation, thereby offering new opportunities for robust disease assessment and prediction in clinically constrained settings.
In parallel with these methodological developments, several review articles have summarized speech analysis research from different perspectives, including classical ML techniques, disease-specific applications, and early DL methods[19-24]. While these reviews provide valuable foundations, they typically focus on limited disease categories or earlier methodological stages. Recent breakthroughs since 2023, particularly those involving foundation-model-based pretraining (e.g., Whisper[25], wav2vec 2.0[26]) and cross-domain generalization, have not yet been systematically integrated. As summarized in Table 1, existing reviews vary substantially in disease coverage, time scope, and methodological focus, and few offer a unified synthesis that spans multiple disease categories while incorporating recent advances in foundation-model-based speech analysis. Accordingly, Table 1 provides a concise comparison between representative prior reviews and the present work in terms of disease scope, time scope, method scope, and primary focus, thereby clarifying the positioning and complementary contribution of this review.
Comparison between existing reviews and our review
| Reference | Disease scope | Time scope | Method scope | Focus |
| Idrisoglu et al., 2023[19] | Systematic conditions, nonlaryngeal aerodigestive disorders, and neurological disorders | 2012-2022 | ML | Cross-disease synthesis; machine learning focus |
| De Silva et al., 2025[20] | Neurological disorders | 2010-2022 | ML, DL | Clinical decision support; neurological focus |
| Hecker et al., 2022[21] | Neurological disorders | 2001-2021 | ML, DL | Acoustic features; traditional pipelines |
| Khaskhoussy and Ben Ayed, 2022[22] | Parkinson’s disease only | 2004-2022 | ML, DL | Early PD detection; speech features |
| Ding et al., 2024[23] | Alzheimer’s disease only | 2018-2023 | ML, DL | AD-focused analysis; datasets and challenges |
| Moell et al., 2025[24] | Speech disorders (cross-disease) | 2000-2023 | ML | Method taxonomy; speech classification |
| Our review | Neurological, psychiatric, and respiratory diseases | 2015-2025 | ML, DL, foundation models | Cross-disease synthesis; task-level perspective |
Motivated by the above gaps, this review aims to provide a comprehensive and up-to-date synthesis of speech analysis for disease assessment, prediction, and diagnosis. Our contributions are summarized as follows: (1) We synthesize AI-based speech analysis frameworks across major disease categories, highlighting the methodological evolution from handcrafted features to DL and foundation models. (2) We compare modeling strategies, datasets, and evaluation protocols across diseases to identify shared technical challenges and disease-specific characteristics. (3) We critically examine key barriers to clinical translation, including interpretability, robustness, and cross-population generalization, and outline future research directions. In addition, we introduce a conceptual framework that organizes disease domains, clinical tasks and methodological categories, as shown in Figure 1. This knowledge graph serves as an organizational framework to facilitate efficient navigation and cross-domain understanding of the review.
Figure 1. Conceptual framework of the review showing the organization of disease domains, clinical tasks and methodological categories. AI: Artificial intelligence; PD: Parkinson’s disease; AD: Alzheimer’s disease; COPD: chronic obstructive pulmonary disease; COVID-19: coronavirus disease 2019; SCZ: Schizophrenia; SHAP: SHapley additive explanations; LIME: local interpretable model-agnostic explanations; MDVR-KCL: Mobile Device Voice Recordings at King’s College London; EWA-DB: early warning of Alzheimer speech database; DAIC-WOZ: Distress Analysis Interview Corpus-Wizard of Oz; MODMA: Multi-modal Open Dataset for Mental-disorder Analysis; CWLSD: Chest Wall Lung Sound Database.
METHOD
This review adopts a scoping review methodology to systematically map and synthesize existing literature on speech-based AI for disease analysis. The methodological framework of this review was informed by the Preferred Reporting Items for Systematic Reviews and Meta-Analyses Extension for Scoping Reviews (PRISMA-ScR)[27], to enhance transparency and reproducibility of the literature mapping process. For more information, please refer to Figure 2.
Figure 2. PRISMA-ScR flow diagram of the literature identification and selection process. PRISMA-ScR: Preferred Reporting Items for Systematic Reviews and Meta-Analyses Extension for Scoping Reviews.
Information sources and search strategy
Based on the venues represented in the final reference list, the literature search covered both biomedical and engineering-oriented databases, including PubMed, Web of Science, IEEE Xplore, ScienceDirect, and Scopus. The search covered studies published between January 2015 and December 2025. Only articles published in English were considered.
The search strategy was structured around four complementary conceptual dimensions: speech-related data modalities, clinical disease contexts, clinical tasks, and AI methodologies. Speech-related terms included speech, voice, audio, and acoustics. Disease-related terms encompassed both general descriptors such as disease and disorder and higher-level disease categories (e.g., neurological, psychiatric, and respiratory disorders). Task-related terms focused on clinically relevant objectives, including diagnosis, assessment, severity grading, prediction, and screening. Methodological terms included ML, DL, AI, neural networks, foundation models, and large models (LMs).
These concepts were combined using Boolean operators, with database-specific adaptations applied where necessary. A representative search query followed the structure:
(“speech” OR “voice” OR “audio”) AND (“disease” OR “disorder” OR “clinical” OR “medical”) AND (“diagnosis” OR “assessment” OR “severity” OR “prediction” OR “screening”) AND (“ML” OR “DL” OR “AI” OR “LM”)
Study selection and eligibility criteria
All retrieved records were imported into a reference management system, and duplicate entries were removed. Two independent reviewers screened titles and abstracts to identify potentially eligible studies, followed by full-text assessment of selected articles. Disagreements were resolved first through consensus or by consultation with another author.
Studies were included if they utilized human speech or voice data as a primary modality, addressed clinically meaningful tasks such as disease diagnosis, assessment, severity grading, screening, or prediction, applied AI-based modeling approaches, and reported quantitative experimental results (e.g., accuracy, sensitivity, specificity, or area under the curve). Both traditional ML, DL, and foundation-model-based approaches were considered.
Studies were excluded if they were limited to abstracts, reviews, editorials, or commentaries; focused exclusively on generic speech tasks (e.g., speech recognition) without disease relevance; relied solely on speech-to-text semantic analysis without acoustic modeling; or failed to report methodological details and experimental performance.
Data extraction and study categorization
For each included study, information regarding disease category, clinical task, speech modality, modeling methodology, dataset characteristics, and evaluation metrics was extracted. To facilitate systematic synthesis, studies were categorized along three primary dimensions: disease domain, clinical task, and AI methodology. This structured organization enabled comparative analysis across diseases and modeling paradigms, supporting a comprehensive assessment of methodological trends and clinical applicability. The distribution of papers per disease task is shown in Figure 3.
SPEECH DATA
As an important digital biomarker of health, speech data has the characteristics of non-invasiveness, ease of acquisition, and convenient collection. Data collection plays a crucial role in the development of ML models for disease analysis and is the cornerstone of building and training these models. In this section, we will introduce several mainstream speech datasets. For more detailed information about the datasets, please consult Table 2.
The summary of speech datasets
| Dataset | Year | Disease | Disease category | Access link |
| Oxford | 2008 | PD | PD:23, HC:6 | https://www.kaggle.com/datasets/thecansin/parkinsons-data-set |
| NeuroVoz | 2024 | PD | PD:54, HC:58 | https://zenodo.org/doi/10.5281/zenodo.10777656 |
| MDVR-KCL | 2017 | PD | PD:16, HC:21 | https://zenodo.org/records/2867216 |
| DementiaBank | 2005 | AD | AD:117, HC:93 | https://tensorflow.google.cn/datasets/catalog/dementiabank |
| EWA-DB | 2017 | PD&AD | AD:87, PD:175, MCI:62, AD&PD:2, HC:1323 | https://zenodo.org/records/10952480 |
| iFLYTEK | 2019 | AD | AD:84, MCI:179, HC:138 | https://challenge.xfyun.cn/2019/gamedetail?blockId=978 |
| DAIC-WOZ | 2014 | Depression | Depression:56, HC:133 | https://dcapswoz.ict.usc.edu/ |
| MODMA | 2022 | Depression | Depression:23, HC:29 | https://modma.lzu.edu.cn/data/index/ |
| The SCZ dataset | 2022 | SCZ | SCZ:34, HC:38 | ____________________ |
| Coswara | 2023 | COVID-19 | COVID-19:674, HC:1819, and 142 recovered subjects | https://coswara.iisc.ac.in/ |
| COUGHVID | 2021 | COVID-19 | COVID-19 over 25,000 recordings (1,155 samples tested positive) | https://opendatalab.org.cn/OpenDataLab/COUGHVID |
| COVID-19 Sounds | 2021 | COVID-19 | COVID-19 36,116 participants (2,106 samples tested positive) | https://openreview.net/forum?id=9KArJb4r5ZQ |
| CWLSD | 2021 | COPD | COPD:77, HC:35 | https://data.mendeley.com/datasets/jwyy9np4gv/3 |
PD. The Oxford Parkinson’s Disease Detection Dataset was created by Little et al.[28] in 2008. By asking subjects to continuously pronounce specific vowel sounds and using the Multi-Dimensional Voice Program (MDVP) and advanced mathematical analysis methods, 22 acoustic features were extracted to distinguish between control participants and PD patients. The creation of this dataset is of great significance for the remote monitoring and early diagnosis of PD. NeuroVoz was jointly recorded by the Bioengineering and Optoelectronics (ByO) group from Universidad Politécnica de Madrid (UPM) and the Otorhinolaryngology and Neurology Services of Hospital General Universitario Gregorio Marañón (HGUGM) and Hospital Universitario de Fuenlabrada (HUF), Madrid, Spain[29]. It provides rich resources for scientific research on the impact of PD on speech and is currently the most complete public speech corpus for PD. The Mobile Device Voice Recordings at King’s College London (MDVR-KCL) dataset was collected by King’s College London (KCL) Hospital in 2017 using smartphones to conduct voice calls with subjects, and all calls were made in a quiet indoor environment[30].
AD. DementiaBank was created by Boller and Becker in 2005. DementiaBank is a shared database of multimedia interactions for the study of communication in dementia[31]. The dataset contains 117 people diagnosed with AD and 93 participants from a control group reading a description of an image. Early Warning of Alzheimer speech database (EWA-DB) is a speech database that contains data from 3 clinical groups: AD, PD, mild cognitive impairment (MCI), as well as a control group of cognitively unimpaired participants[32]. iFLYTEK is a Chinese dataset created in 2019. It contains the speech and text data of 138 control participants, 179 people with MCI, and 84 AD patients. These datasets provide researchers with comprehensive data support for in-depth exploration of speech features during the progression of AD, significantly advancing the application of speech analysis in AD diagnostic research.
Depression. The Distress Analysis Interview Corpus-Wizard of Oz (DAIC-WOZ) database is part of the Distress Analysis Interview Corpus (DAIC)[33]. This corpus mainly contains clinical interview records and aims to support the diagnosis of psychological distress conditions such as anxiety, depression, and post-traumatic stress disorder. The dataset contains 189 interview data, with an average interview duration of 16 minutes. Each interview includes the transcribed text of the interview, the audio file of the participant, and facial feature information. This dataset is commonly used in text-based detection, speech-based detection, and multimodal architecture research. The Multi-modal Open Dataset for Mental-disorder Analysis (MODMA) dataset is a multi-modal open dataset for mental disorder analysis, released and continuously updated by the Key Laboratory of Wearable Devices of Gansu Province, Lanzhou University[34]. It contains data of clinical depression patients and control participants.
Schizophrenia (SCZ). The Schizophrenia dataset (also named SCZ dataset) is recruited by the Psychiatry Department of the Mental Health Center, Sichuan University[35]. It comprises 34 first-episode drug-naive patients with SCZ and 38 participants from a control group. All participants are asked to read text with neutral, positive, and negative sentiments. All recordings were made in 16-bit format using SONY ICD-TX650, and the sampling frequency is 44.1 kHz. Specifically, SCZ dataset comprises 720 utterances (340 schizophrenic patients and 380 control participants) with a neutral sentiment, 569 utterances (271 schizophrenic patients and 298 control participants) with a positive sentiment (emotional state of happiness), and 216 utterances (102 schizophrenic patients and 114 control participants) with a negative sentiment (emotional state of anger).
Coronavirus disease 2019. Coswara is one of the most widely used datasets for sound-based coronavirus disease 2019 (COVID-19) detection[36]. Its sound samples were crowdsourced globally via a web application, recorded and stored at a sampling frequency of 16kHz, comprising over 140,000 audio files. This dataset contains a diverse set of respiratory sounds and rich metadata, recorded between April 2020 and February 2022 from 2,635 individuals [1,819 SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2) negative, 674 positive, and 142 recovered subjects]. The respiratory sounds encompass nine categories associated with variations of breathing, cough, and speech. The rich metadata includes demographic information such as age, gender, and geographic location, as well as health information relating to symptoms, pre-existing respiratory ailments, comorbidities, and SARS-CoV-2 test status. The COUGHVID crowdsourcing dataset is an extensive, validated, and publicly available dataset of cough recordings[37]. With over 20,000 recordings (including 1,010 self-reported COVID-19 cases) sourced globally. Notably, Experienced pulmonologists labeled more than 2,000 recordings to diagnose medical abnormalities present in the coughs, thereby contributing one of the largest expert-labeled cough datasets in existence that can be used for a plethora of cough audio classification tasks. To the best of our knowledge, COVID-19 Sounds is the largest multimodal COVID-19 respiratory sound dataset: it comprises three modalities, namely breathing, cough, and voice recordings[38]. Crowd-sourced from 36,116 global participants via the COVID-19 Sounds App, the dataset contains 53,449 audio samples (including 2,106 COVID-19 positive samples). As the dataset was crowd-sourced from various platforms, it contains diverse audio file formats (e.g., .ogg, .m4a, .wav, and .webm) and sampling rates (specifically: 2.6% at 8KHz, 0.3% at 12KHz, 50.3% at 16KHz, 36.7% at 44.1KHz, and 10.1% at 48KHz).
COPD. The Chest Wall Lung Sound Database (CWLSD)[39] comprises sounds from seven medical conditions (namely asthma, heart failure, pneumonia, bronchitis, pleural effusion, lung fibrosis, and COPD) and normal breathing sounds. This dataset features audio recordings collected from chest wall examinations at multiple vantage points. It includes respiratory sounds from 112 subjects (35 control participants and 77 patients with ailments, among whom 9 are COPD patients).
AI FRAMEWORK
In this section, we introduce the framework for speech-based disease analysis (see Figure 4), including main steps such as pre-processing, feature engineering, and model construction. Firstly, pre-processing the original speech data eliminates irrelevant information such as noise and interference. Then, speech feature parameters that can reflect the characteristics of diseases are extracted from the pre-processed speech data, converting the original speech signal into a representative and discriminative feature vector. Next, feature selection is used to remove redundant and irrelevant features, selecting the most valuable and discriminative feature subset from the numerous extracted features. Finally, a model is constructed to leverage computational power and ML to automatically identify the relationships between speech features and diseases, enabling the assessment, prediction, and diagnosis of new speech data.
Pre-processing
In the reviewed literature, speech pre-processing served as a task-driven step designed to enhance signal quality and adapt raw recordings to disease-specific diagnostic scenarios. Most studies applied a standardized preprocessing pipeline tailored to clinical speech tasks such as sustained phonation, reading passages, or spontaneous speech.
Pre-emphasis was widely used to amplify high-frequency components that are typically attenuated during speech production and recording. In PD studies relying on cepstral and linear predictive features, pre-emphasis was applied prior to Mel-Frequency Cepstral Coefficients (MFCC) or Linear Predictive Coding (LPC) extraction to better capture articulatory imprecision and phonatory instability associated with hypokinetic dysarthria[4,13,40]. Similar preprocessing strategies were also reported in early AD speech analysis to enhance subtle spectral changes related to cognitive decline[41,42].
Framing and windowing were universally adopted to exploit the short-term quasi-stationary nature of speech signals. Across neurological and psychiatric disorder studies, speech signals were typically segmented into frames of 20-40 ms with partial overlap, and Hamming windows were the most frequently employed windowing function[16,43,44]. This configuration ensured stable short-term spectral representations and reduced spectral artifacts caused by abrupt signal truncation.
Endpoint detection was primarily used to isolate effective speech segments and exclude silence or low-energy regions. This step was particularly relevant in PD and dysarthria studies using sustained vowel phonation, where accurate isolation of voiced segments improved robustness against background noise and recording variability[45,46]. In spontaneous speech tasks for AD and depression detection, endpoint detection also helped remove non-informative pauses and recording artifacts[6,7].
Overall, preprocessing in the reviewed studies was closely aligned with the characteristics of the speech task and the targeted disease, rather than being treated as a purely technical routine.
Feature engineering
Feature engineering serves as a critical link between raw speech signals and downstream diagnostic models. In the reviewed studies, this process generally comprised two stages: feature extraction, which transforms speech signals into acoustic representations, and feature selection, which aims to retain disease-relevant features while reducing redundancy and dimensionality. The choice of these strategies was closely influenced by disease characteristics, speech tasks, and dataset scale.
Feature extraction. Feature engineering remained a central component in speech-based disease analysis. The reviewed studies consistently employed combinations of prosodic, spectral, and voice quality-related features, with the choice of features reflecting both disease pathology and data availability.
Prosodic features, including fundamental frequency, speaking rate, pause duration, and intensity-related measures, were widely used to characterize temporal and rhythmic abnormalities in speech. These features were particularly prevalent in studies on PD, depression, and SCZ, where altered speech rhythm and prosody are clinically observable symptoms[47-49].
Spectral features constituted the most commonly used feature category across all disease domains. MFCCs, LPC, and log-Mel spectrogram representations were repeatedly adopted in PD and AD studies due to their effectiveness in encoding vocal tract characteristics and phonatory patterns[50]. In several works, spectral features formed the primary input for both traditional ML and DL models.
Voice quality-related features, such as jitter, shimmer, and harmonics-to-noise ratio (HNR), were frequently employed to capture micro-instabilities in vocal fold vibration. These features were especially prominent in PD and voice disorder studies, reflecting impairments in phonatory control and vocal stability[40,45,46]. Many studies reported improved diagnostic performance when voice quality features were combined with spectral representations.
Feature selection. Given the limited sample sizes typical of clinical speech datasets, feature selection was widely applied to reduce dimensionality and mitigate overfitting. Most reviewed studies favored statistically motivated filter methods or embedded feature selection strategies integrated within classifiers, such as L1-regularized models and tree-based approaches[51,52]. Wrapper-based methods were less frequently adopted due to their high computational cost and sensitivity to small datasets.
Rather than emphasizing methodological taxonomy, feature selection in the reviewed literature primarily aimed to identify disease-sensitive features and improve model generalizability. Representative feature selection strategies are summarized in Table 3.
The classification of feature selection methods
| Category | Definition | Common methods |
| Filter | Evaluate the importance of features by assessing the statistical properties of features (such as variance, correlation, mutual information, etc.), and select features accordingly | Chi-square test, mutual information, Pearson correlation coefficient, etc. |
| Wrapper | Regard feature selection as an optimization problem, and find the optimal feature combination by searching different feature subsets | Recursive Feature Elimination (RFE), Forward Selection, etc. |
| Embedded | Embedded feature selection incorporates feature selection into the model training process. By evaluating and adjusting the importance of features during the model training process, the optimal feature subset is selected | Lasso regression, decision trees, etc. |
Model construction
In the process of using speech data for disease diagnosis, model construction is the core link. It determines how to extract key features related to diseases from complex speech signals and make accurate predictions or diagnoses based on them. Currently, the construction methods of AI-assisted diagnosis models can be mainly divided into three categories: traditional ML, DL and LMs. For the specific comparison of the advantages and disadvantages of these three methods, please refer to Figure 5.
Machine learning. Traditional ML methods remain widely adopted in speech-based disease diagnosis, particularly in studies relying on handcrafted acoustic features and limited clinical datasets. Algorithms such as support vector machines (SVMs) and random forests (RFs) are frequently used due to their relatively low data requirements and stable performance in small-sample scenarios. From a methodological perspective, ML models operate on explicitly defined feature spaces, allowing them to exploit disease-sensitive acoustic descriptors such as prosodic irregularities or voice quality perturbations. This property makes ML approaches particularly suitable for clinical studies where interpretability and feature-level analysis are important. However, their reliance on predefined features also constrains their ability to capture complex temporal dependencies and subtle nonlinear patterns in speech, which may limit performance in more heterogeneous disease populations.
Deep learning. With the advancement of DL techniques, CNNs have been widely applied to disease-related speech analysis. Recurrent neural networks (RNNs)[53], including long short-term memory (LSTM)[54] networks and gated recurrent units (GRUs)[55], as well as Transformer-based models, have also been increasingly explored in this domain. Unlike traditional ML approaches, DL models can learn hierarchical representations directly from time-frequency features, such as spectrograms, thereby reducing dependence on manual feature design. CNN-based architectures are particularly effective in modeling local spectral patterns, while RNNs and Transformers are better suited for capturing temporal dynamics and long-range dependencies in speech signals. These capabilities are advantageous for diseases characterized by progressive or task-dependent speech impairments. Nevertheless, the reviewed studies indicate that DL-based models typically require larger datasets and careful regularization to avoid overfitting, which remains a practical limitation in many clinical speech datasets.
Large models. LMs, including foundation models pretrained on massive speech or multimodal corpora, represent an emerging paradigm in speech analysis. Models such as Qwen-Audio[56], LlaMA2[57] and Viola[58] have demonstrated strong general-purpose representation learning capabilities. In theory, such models offer the potential to capture complex and subtle speech patterns that may be difficult to learn from limited clinical data alone.
However, based on the reviewed literature, the application of LMs to disease-specific speech diagnosis remains largely exploratory. Most existing studies focus on feasibility analysis, transfer learning, or hypothetical clinical potential rather than validated diagnostic deployment. Challenges such as domain mismatch, limited labeled medical speech data, and concerns regarding interpretability currently restrict their widespread clinical adoption. As a result, LMs should be regarded as a promising future direction rather than a mature solution for speech-based disease diagnosis.
Overall, the reviewed studies suggest that no single modeling paradigm is universally optimal for speech-based disease diagnosis. Traditional ML methods remain effective in small-scale, feature-driven clinical settings, while DL approaches offer improved representation capacity at the cost of higher data demands. LMs introduce new opportunities for scalable and transferable speech modeling, but their clinical readiness has yet to be fully demonstrated. The choice of modeling strategy therefore depends on disease characteristics, dataset scale, and the balance between diagnostic performance and interpretability.
NEUROLOGICAL DISORDERS
Neurological disorders are the leading cause of the global disease burden, significantly affecting both the central and peripheral nervous systems[59]. In 2021, neurological disorders affected approximately 3.4 billion people worldwide, accounting for 43% of the global population[60]. The research on the application of speech processing technology in the diagnosis of neurological disorders, particularly in the context of PD and AD, has witnessed significant advancements in recent times. In this section, we will comprehensively and elaborately review the relevant research achievements of speech in the diagnosis of these two neurological disorders. The summary of research literature is shown in Table 4.
The Summary of applications of speech analysis in neurological disorders
| Reference | Disease | Task | Method | Modality | Performance |
| Upadhya et al.[40] | PD | Classification | NN | Speech | Accuracy: 98% Specificity: 96.6% Sensitivity: 99.4% |
| Haider et al.[42] | AD | Detection | DT | Speech | Accuracy: 78.7% |
| Faragó et al.[44] | PD | Identification in noise | CNN | Speech | Accuracy: 96% |
| Bayesiehtashk et al.[46] | PD | Severity Assessment | Ridge regression | Speech | - |
| Hason et al.[50] | AD | Prediction and stages classification | RF | Speech | Accuracy: 82.2% Precision: 81.6% Recall: 81.4% AUC: 89.3% |
| Karan[51] | PD | Prediction | XGBoost | Speech | Accuracy: 95.07% AUC: 96% Specificity: 89.57% Sensitivity: 95.07% |
| Tunc et al.[52] | PD | Severity Estimation | XGBoost | Speech | MAE: 7.13±1.07 |
| Moro-Velazquez et al.[61] | PD | Detection | GMM-UBM classifiers | Speech | Accuracy: 75%-82% AUC: 84%-95% |
| Shastry[62] | PD | Prediction | KNN + GB | Speech | Accuracy: 75.48% Precision: 75.63% Recall: 74.92% F1: 75.06% AUC: 74.92% |
| Mohammadi et al.[63] | PD | Diagnosis | LR | Speech | Accuracy: 97.22% |
| Govindu et al.[64] | PD | Early Detection | RF | Speech | Accuracy: 91.83% Sensitivity: 95% |
| Mahesh et al.[65] | PD | Prediction | XGBoost-RF | Speech | Accuracy: 98% Precision: 97.24% F1: 97.4% Specificity: 97% Sensitivity: 97.56% |
| Al Mudawi et al.[66] | PD | Detection | LightGBM | Speech | Accuracy: 98.3051% |
| Jain et al.[67] | PD | Detection | KNN | Speech | Accuracy: 97.33% Precision: 96.2963% Recall: 1% F1: 98.1132% |
| Wang et al.[68] | PD | Diagnosis | EMSFE | Speech | Accuracy: 92.5% Precision: 94.7% Specificity: 95% Sensitivity: 90% |
| Deepa et al.[69] | PD | Detection & Classification | ERT | Speech | Accuracy: 87% |
| Abedinzadeh Torghabeh et al.[70] | PD | Severity Assessment | WMV | Speech | Accuracy: 98.62% Precision: 98.62% F1: 98.62% Specificity: 99.54% Sensitivity: 98.62% |
| Laudis et al.[71] | PD | Classification | SVM | Speech | Accuracy: 92.5% Precision: 91% Recall: 94% F1: 92.5% AUC: 93% Specificity: 93% Accuracy: 91% |
| Yuan et al.[72] | PD | Prediction | DNN | Speech | Accuracy: 95% F1: 97% |
| Wrobel et al.[73] | PD | Diagnosis | MLP | Speech | Accuracy: 90.6% |
| Liu et al.[74] | PD | Prediction | ANN | Speech | Precision: 96% F1: 98% AUC: 91% Specificity: 82% Sensitivity: 99% |
| Quan et al.[75] | PD | Detection | CNN | Speech | Accuracy: 92% |
| Tayebi Arasich et al.[76] | PD | Detection | wav2vec 2.0 | Speech | Accuracy: 83.2% AUC: 85% Specificity: 89.2% Sensitivity: 90.8% |
| Wang et al.[77] | PD | Recognition | SVM | Speech | Accuracy: 87.5% Specificity: 86.11% Sensitivity: 88.89% |
| Xu et al.[78] | PD | Diagnosis | DNN | Speech | Accuracy: 89.5% |
| Pandey et al.[79] | PD | Prediction | CNN-LSTM | Speech | Accuracy: 97% |
| Mishra et al.[80] | PD | Severity Assessment | DNN | Speech | Accuracy: 96.2% Specificity: 96.15% Sensitivity: 94.15% |
| Jeancoias et al.[81] | PD | Early Detection | DNN + X-vectors | Speech | |
| Chronowski et al.[82] | PD | Diagnosis | wav2vec 2.0 | Speech | Accuracy: 97.92% AUC: 99% |
| Khaskhoussy et al.[83] | PD | Diagnosis | MLP | Speech | Accuracy: 95.52% |
| Hireš et al.[84] | PD | Detection | CNN | Speech | Accuracy: 99% AUC: 89.6% Specificity: 93.3% Sensitivity: 86.2% |
| Akila et al.[85] | PD | Classification | CNN | Speech | Accuracy: 99.1% Precision: 97.8% Recall: 94.7% F1: 99.5% |
| Palakayala et al.[86] | PD | Detection | DCNN + KNN | Speech | Accuracy: 93.7% |
| Skibińska et al.[87] | PD | Diagnosis | XGBoost[177] | audio and video | Accuracy: 83% Specificity: 78% Sensitivity: 88% |
| Khan Tusar et al.[88] | PD | Early Diagnosis | GB/AdaBoost | Speech | Accuracy: 92% F1: 92% AUC: 81% |
| Yousif et al.[89] | PD | Early Diagnosis | KNN+SVM | Images and Speech | Accuracy: 99.94% |
| Pappagari et al.[93] | Assessment and detection | X-vectors, BERT | Speech | Precision: 70% Recall: 88% F1: 78% | |
| Khodabakhsh et al.[95] | AD | Detection | SVM | Speech | Accuracy: 83.5% |
| König et al.[96] | AD | Assessment and detection of early stage dementia and MCI | ML | Speech | Accuracy: 92% |
| Li et al.[97] | AD | dementia Detection | LR | Speech | Accuracy: 77% |
| Ben Ammar et al.[98] | AD | Early Detection | SVM | Speech | Accuracy: 91(± 0.5)% Recall: 91(± 0.5)% |
| Nasrolahzadeh et al.[99] | AD | Diagnosis | GP | Speech | Accuracy: 99.09% |
| König[100] | AD | MCI/AD Classification | SVM | Speech | Accuracy: 79% ± 5%(MCI vs. HC) 87% ± 3%(AD vs. HC) 80% ± 5%(MCI vs. AD) |
| López-de-Ipiña et al.[101] | AD | Diagnosis | SVM | Speech | Accuracy: 93.79% |
| García-Gutiérrez et al.[102] | AD | SCD/MCI/ADD Identification | ML | Speech | F1: 92 |
| Kim et al.[103] | AD | Classification | Speech | Accuracy: 78% Precision: 82% Recall: 76% F1: 79% | |
| Chien et al.[104] | AD | Assessment | RNN | Speech | Accuracy: 83.8% AUC: 83.8 ± 3% Specificity: 76.4 ± 6% Sensitivity: 75.6 ± 7% |
| Roshanzamir et al.[105] | AD | Risk Assessment | Transformer+LR | Speech | Accuracy: 88.08% Precision: 90.57% Recall: 84.34% F1: 87.23% |
| Dong et al.[106] | AD | Detection | HAFFormer | Speech | Accuracy: 82.6% F1: 82.6% |
| Liu et al.[107] | AD | Detection | Transformer | Speech | Accuracy: 93.5% Precision: 94% Recall: 89% F1: 91.19% |
| Liu et al.[108] | AD | Detection | CNN + BiLSTM | Speech | Accuracy: 82.59% Precision: 85.24% Recall: 81.48% F1: 82.94% |
| Ahn et al.[109] | AD | Early Detection | Densenet121 | Speech | Accuracy: 90% F1: 91.39% AUC: 92.43% Specificity: 83.33% Sensitivity: 95.5% |
| Farazi et al.[110] | AD | Detection | CNN | Speech | Accuracy: 85% |
| Mittal et al.[111] | AD | AD detection | BERT + CNN | Speech and Text | Accuracy: 85.3% |
| Haulcy et al.[112] | AD | Classification | BERT + SVM/RF | Audio and Text | Accuracy: 85.4% F1: 84.4% Specificity: 82.3% Sensitivity: 89.2% |
| Li et al.[113] | AD | Prediction | BERT | Audio and Text | Accuracy: 83.69% |
| Jang et al.[114] | AD | Classification | LR | Speech and Video | Accuracy: 83 ± 1% |
| Ablimit et al.[115] | AD | Detection | CNN-GRU | Speech | Recall: 70.6%UAR |
| Martinc et al.[116] | AD | Diagnosis | BERT | Audio and Text | Accuracy: 93.75% |
| Mahajan et al.[117] | AD | Detection | CNN-LSTM | Speech | Accuracy: 72.92% Precision: 78.94% Recall: 62.5% F1: 69.76% |
| Zhang et al.[118] | AD | Detection | wav2vec 2.0 | Speech | Accuracy: 85.45% |
| Li et al.[119] | AD | Classification | Whisper | Speech | Accuracy: 84.51% Precision: 83.33% Recall: 85.71% F1: 84.5% |
| Bang et al.[120] | AD | Prediction | ChatGPT | Speech, Text, and Opinions | Accuracy: 87.3% Precision: 88.06% Recall: 87.32% F1: 87.25% Specificity: 94.44% |
| Cui et al.[121] | AD | Detection | WavLM + BERT | Speech and Text | F1: 92.8% |
PD
PD is a progressive neurodegenerative disorder characterized by motor and non-motor impairments. Speech disorders are among the most prevalent symptoms, affecting approximately 70%-90% of patients and substantially reducing quality of life[45]. Compared with the control group, individuals with PD exhibit consistent abnormalities across multiple acoustic dimensions, including phonation stability, articulation, and prosody[43]. These speech alterations have motivated extensive research into speech-based PD diagnosis, positioning speech as a non-invasive and cost-effective digital biomarker.
Early studies focused on identifying discriminative acoustic markers between PD patients and control groups. For example, Wang et al.[49] reported systematically lower formant frequencies in PD patients across multiple syllable phonation tasks, suggesting impaired articulatory control. Moro-Velazquez et al.[61] further demonstrated that specific phoneme groups - particularly plosives, vowels, and fricatives - carry higher diagnostic relevance for PD detection across multiple speech corpora. Similarly, Upadhya et al.[40] showed that articulatory features, either alone or combined with cepstral features, provide strong discriminative power, highlighting the importance of motor speech dysfunction in PD-related vocal impairment.
Traditional ML models with handcrafted features
Traditional ML approaches constitute the earliest and most extensively studied paradigm for speech-based PD diagnosis. These methods typically rely on handcrafted acoustic features, such as fundamental frequency (F0), jitter, shimmer, HNR, formant-related measures, and MFCCs, combined with classical classifiers including SVM, RF, KNN, Logistic Regression (LR), and gradient boosting machines (GBMs)[62,63].
Across multiple studies, ensemble-based and tree-based models frequently demonstrate strong diagnostic performance. Govindu et al.[64] and Mahesh et al.[65] reported that RF-based or boosting-based classifiers outperform other ML models when trained on perturbation-related vocal features. Similar trends were observed in comparative studies by Al Mudawi et al.[66] and Jain et al.[67], where Light Gradient Boosting Machine (LightGBM) and KNN achieved high classification accuracy. Wang et al.[68] designed a novel ensemble learning algorithm (EMSFE) based on a Sparse Fusion Ensemble Learning Mechanism (SFELM). Experimental results showed that the proposed algorithm achieved an accuracy of 93.75%. These findings suggest that non-linear ensemble learners are particularly well-suited for capturing subtle interactions among handcrafted speech features.
Beyond binary diagnosis, several studies explored severity estimation and continuous monitoring[69]. Abedinzadeh Torghabeh et al.[70] demonstrated that optimized ML classifiers could robustly stratify PD severity levels, while Bayesiehtashk et al.[46] and Tunc et al.[52] showed that speech-derived features correlate with clinical severity scores across multiple speech tasks. These results support the feasibility of speech-based models for longitudinal PD monitoring rather than solely binary classification.
Feature selection plays a crucial role in improving model efficiency and robustness. Laudis et al.[71] showed that reducing MFCC dimensionality via feature selection not only improved classification accuracy but also reduced computational cost. Similar observations were reported by Yuan et al.[72] and Karan[51], indicating that a compact subset of well-chosen acoustic features often outperforms high-dimensional representations[73].
Overall, handcrafted-feature-based ML models consistently achieve strong performance on benchmark datasets and offer relatively high interpretability. However, many studies rely on small or single-center datasets, raising concerns about overfitting and limited generalizability to real-world clinical populations.
DL models
DL approaches have increasingly been adopted to overcome the limitations of manual feature engineering by enabling automatic representation learning from raw or minimally processed speech signals[74]. CNNs, RNNs, and Transformer-based architectures have been explored across various speech tasks.
Quan et al.[75] proposed an end-to-end DL framework combining temporal modeling with spectrogram-based representations, demonstrating improved performance over expert-crafted features on both sustained vowel and reading tasks. Tayebi Arasich et al.[76] extended DL-based PD diagnosis to a federated learning (FL) setting, showing that privacy-preserving collaboration across multilingual datasets can outperform monolingual models without data sharing, addressing a key barrier to multi-center clinical adoption.
Several studies focused on improving data efficiency and robustness. Wang et al.[77] introduced a deep speech sample learning strategy to reconstruct high-quality prototype samples, while Xu et al.[78] employed a Deep Neural Network (DNN) framework using voiceprint features to enhance PD discrimination. Hybrid DL architectures, such as CNN-LSTM combinations, were explored by Pandey et al.[79], aiming to capture both spectral patterns and temporal dynamics. These approaches generally outperform traditional ML baselines, particularly in continuous or spontaneous speech scenarios.
Noise robustness and real-world applicability were addressed by Faragó et al.[44], who demonstrated effective PD detection from noisy continuous speech, and Mishra et al.[80], who implemented a cloud-based home monitoring framework. Early-stage detection was further explored by Jeancoias et al.[81], showing that speaker embedding techniques such as X-vectors are particularly effective for detecting early PD, especially in text-independent settings.
Recent studies have leveraged transfer learning from large-scale speech models. Chronowski et al.[82] utilized wav2vec 2.0 representations for PD classification, achieving strong performance under cross-validation. Other works explored autoencoders and optimized CNN architectures to further push diagnostic accuracy[83-86].
Overall, DL-based methods demonstrate strong potential for learning disease-relevant speech representations and generalizing across tasks and languages. Nevertheless, many reported results are obtained under controlled experimental settings, and their robustness across heterogeneous clinical populations remains to be fully validated.
Multimodal fusion
In addition to speech-only analysis, several studies investigated multimodal fusion strategies to enhance diagnostic robustness. Skibińska et al.[87] combined audio and video data, while Khan Tusar et al.[88] integrated vocal features with clinical and motor examination data. Yousif et al.[89] further explored fusion of speech and handwriting modalities, reporting very high diagnostic accuracy under experimental conditions.
While multimodal fusion consistently improves classification performance, many of these studies rely on limited datasets and task-specific protocols. Consequently, although multimodal approaches are promising, their clinical applicability requires standardized acquisition protocols, larger cohorts, and prospective validation before deployment in routine clinical practice.
Methodological synthesis for PD
From a clinical perspective, many discriminative speech features identified across studies align with known PD pathophysiology. Perturbation-based features (e.g., jitter and shimmer) reflect impaired phonatory control and hypophonia, while articulation-related features and vowel formants correspond to motor speech dysfunction and dysarthria. Temporal speech abnormalities further mirror bradykinesia and reduced motor coordination.
However, a recurring concern is the prevalence of extremely high reported accuracies, often exceeding 95%. Such results are frequently derived from small datasets, repeated use of benchmark corpora, or cross-validation without external validation, which may overestimate real-world performance. Speech-based PD diagnosis should therefore be viewed as a complementary tool to established clinical assessments, such as the Unified Parkinson’s Disease Rating Scale (UPDRS), rather than a standalone diagnostic replacement.
AD
Language impairment is one of the most prominent and early manifestations of AD, reflecting progressive cognitive decline across multiple domains, including memory, attention, and executive function[90,91]. With advances in speech processing and AI, speech has emerged as a promising non-invasive biomarker for AD screening and monitoring. Numerous studies have demonstrated that acoustic and linguistic characteristics extracted from spontaneous and task-based speech can effectively capture disease-related cognitive impairment[41,92-94].
Traditional ML models with handcrafted features
Early research on speech-based AD diagnosis primarily relied on handcrafted acoustic and linguistic features combined with classical ML classifiers. Commonly used feature sets include prosodic measures, MFCCs, Computational Paralinguistics ChallengE (ComParE), extended Geneva Minimalistic Acoustic Parameter Set (eGeMAPS), and linguistic indicators derived from transcripts, analyzed using LR, RF, and SVM classifiers.
Several studies focused on identifying core speech biomarkers associated with AD. Khodabakhsh et al.[95] showed that prosodic features extracted from conversational speech significantly outperformed linguistic features in distinguishing AD patients from control groups. Similarly, König et al.[96] demonstrated that fluency-related speech tasks recorded via mobile applications were particularly effective for differentiating AD, MCI, and subjective cognitive impairment, highlighting the feasibility of remote speech-based screening.
Cross-lingual robustness emerged as an important research direction. Li et al.[97] demonstrated that mapping engineered speech features across languages enables AD detection in low-resource settings, while Haider et al.[42] introduced an Active Data Representation framework that improved purely acoustic-based AD recognition. Linguistic feature-based approaches further showed strong discriminative power. Ben Ammar et al.[98] and Nasrolahzadeh et al.[99] demonstrated that language features and higher-order spectral representations can serve as effective biomarkers, although such methods often depend on high-quality transcripts.
Clinical feasibility has been explored in multiple studies, where traditional ML classifiers combined acoustic and linguistic features to analyze spontaneous speech, typically achieving diagnostic accuracies of approximately 80% to over 90% across early detection, staging, and severity assessment tasks[100-102]. These findings suggest the potential of non-invasive, low-cost, and remotely deployable speech-based screening approaches, rather than confirmed large-scale clinical implementation.
From a clinical perspective, traditional ML approaches offer interpretability and modest computational cost. However, their performance is sensitive to feature design and often degrades when applied to spontaneous or noisy speech, limiting generalizability across populations.
Overall, handcrafted-feature-based ML methods provide interpretable insights into speech and language impairment in AD but rely heavily on manual feature engineering and task-specific design.
DL models
DL techniques have increasingly been adopted to automatically learn discriminative representations from speech, reducing reliance on handcrafted features. CNNs, RNNs, and Transformer-based architectures dominate this research stream.
Comparative studies have consistently shown DL methods outperform traditional ML across languages. Kim et al.[103] demonstrated that DL models trained on raw acoustic representations achieved higher accuracy and faster inference than ML models using handcrafted features. Temporal modeling of spontaneous speech was further explored using RNN-based architectures, with Chien et al.[104] reporting improved discrimination through sequence-level representations.
Transformer-based models addressed challenges associated with long and heterogeneous speech sequences. Roshanzamir et al.[105] utilized a Transformer- based DL model (specifically BERT Large combined with a logistic regression classifier) for early AD risk assessment based on speech transcripts from picture description tests, achieving 88.08% classification accuracy. Dong et al.[106] proposed the Hierarchical Attention-Free Transformer (HAFFormer) framework to reduce computational complexity while maintaining performance, and Liu et al.[107] further refined Transformer representations through feature purification mechanisms. These studies highlight the importance of modeling long-range dependencies in spontaneous speech for AD detection.
Other works focused on improving data efficiency and reducing annotation dependency. Liu et al.[108] leveraged automatic speech recognition (ASR)-derived acoustic features to eliminate the need for manual transcription, while Ahn et al.[109] and Farazi et al.[110] demonstrated the effectiveness of CNN-based architectures on MFCC and Mel-spectrogram representations.
Overall, DL-based approaches show strong potential for capturing complex speech patterns associated with cognitive decline. However, many studies still rely on relatively small benchmark datasets, and external validation across diverse clinical cohorts remains limited.
Multimodal fusion
To enhance robustness, several studies integrated speech with complementary modalities, such as text, eye-tracking, or demographic information. Multimodal fusion consistently outperformed unimodal approaches by capturing both acoustic and cognitive-linguistic aspects of AD.
Mittal et al.[111] and Haulcy et al.[112] combined audio and textual representations, demonstrating that speech-text fusion improves classification accuracy over single-modality systems. Li et al.[113] proposed a multi-feature fusion model combining MFCC, audio, and text features, achieving 83.69% accuracy in AD detection. Beyond language, Jang et al.[114] incorporated eye-tracking features, while Ablimit et al.[115] integrated demographic information, improving interpretability and clinical relevance. Similarly, Martinc et al.[116] fused spontaneous speech audio with textual features and achieved high diagnostic accuracy for AD, while Mahajan et al.[117] demonstrated that bimodal deep learning models integrating acoustic and transcript-derived linguistic features consistently improved precision compared with acoustic-only baselines.
Despite promising performance, multimodal approaches often require complex data acquisition protocols and standardized task design. Consequently, their translation into large-scale clinical screening remains challenging.
Foundation and pre-trained models
Recent studies have explored pre-trained foundation models to leverage large-scale speech and language knowledge for AD detection. Zhang et al.[118] demonstrated that wav2vec 2.0 representations significantly improve AD recognition performance. Li et al.[119] further showed that incorporating global semantic context using Whisper-based transcripts enhances diagnostic accuracy.
Large language models have also been investigated. Bang et al.[120] and Cui et al.[121] explored LLM-based and sequential knowledge transfer frameworks, demonstrating improved detection of AD and related affective disorders. These approaches represent a shift toward generalized, adaptable diagnostic frameworks.
Nevertheless, most foundation-model-based studies remain retrospective and experimental. Their clinical readiness requires careful validation, transparency, and assessment of failure modes.
Methodological synthesis for AD
Speech and language impairments in AD reflect underlying cognitive deficits in memory, executive function, and semantic processing. Acoustic degradation, reduced fluency, and linguistic simplification observed across studies align with known neuropathological mechanisms.
However, as with PD, many studies report high diagnostic accuracy under controlled conditions. These results should be interpreted cautiously, as real-world deployment requires robustness to demographic variability, spontaneous speech, and recording conditions. Speech-based AD diagnosis is best positioned as a complementary screening tool rather than a replacement for clinical neuropsychological assessment.
PSYCHIATRIC DISORDERS
Psychiatric disorders have long been a major public health concern and a primary contributor to the global health-related burden. From 1990 to 2021, the global incidence of psychiatric disorders showed a significant upward trend, making timely diagnosis and treatment crucial for improving patients’ quality of life[122]. In this section, we focus on two major conditions, depression and SCZ, to examine the applications of speech analysis in mental health. The summary of research literature is shown in Table 5.
Summary of applications of speech analysis in psychiatric disorders
| Reference | Disease | Task | Method | Modality | Performance |
| Shin et al.[7] | Depression | Detection | MLP | Speech | F1: 58.9% AUC: 65.9% Specificity: 66.2% Sensitivity: 65.6% |
| Dumpala et al.[8] | Depression | Detection and Severity Estimation | CNN/LSTM | Speech | Accuracy: 66% (balanced accuracy) F1: 51% (DAIC-WOZ) |
| Berardi et al.[47] | SCZ and Depression | Classification | SVM | Speech | Accuracy: 94.7% (HC vs SSD) Accuracy: 92% (HC vs MDD) Accuracy: 93.2% (SSD vs MDD) |
| Sahu et al.[48] | Depression | Detection | SVM | Speech | Accuracy: 77.8% |
| Scibelli et al.[123] | Depression | Detection | SVM | Speech | Accuracy: over 75% |
| Jiang et al.[124] | Depression | Detection | LR | Speech | Accuracy: 75% (females) Accuracy: 81.82% (males) Specificity: 70.59% (females) Specificity: 85.29% (males) Sensitivity: 79.25% (females) Sensitivity: 78.13% (males) |
| Xu et al.[125] | SCZ and Depression | Classification and Severity Assessment | Ensemble | Audio and video | Accuracy: 82.3% (SCZ vs. HC) Accuracy: 82.3% (MDD vs. HC) Accuracy: 84.7% (MDD vs. SCZ) |
| Shankayi et al.[126] | Depression | Recognition | SVM | Speech | Accuracy: 92.85% |
| Zulfiker et al.[127] | Depression | Prediction | AdaBoost | Text | Accuracy: 92.56% Precision: 95.77% Recall: 91.89% F1: 93.79% AUC: 96% Specificity: 93.62% Sensitivity: 91.89% |
| König et al.[128] | Depression | Detection | SVM | Speech + Text | Accuracy: 93% |
| Kim et al.[129] | Depression | Prediction | RF | Text | Accuracy: 93.1% AUC: 82.3% Specificity: 99.3% |
| He et al.[130] | Depression | Severity Assessment | DCNN | Speech | - |
| Chlasta et al.[131] | Depression | detection | ResNet-34 | Speech | Accuracy: 77% Precision: 57.14% Recall: 66.67% F1: 61.54% |
| Muzammel et al.[132] | Depression | Diagnosis | CNN | Speech | Accuracy: 86.06% Precision: 81% Recall: 73% F1: 77% AUC: 83% |
| Srimadhur et al.[133] | Depression | Detection and Assessment | CNN | Speech | Accuracy: 74.64% Precision: 79% Recall: 74% F1: 77% |
| Kim et al.[134] | Depression | Classification | CNN | Speech | Accuracy: 78.14% Precision: 76.83% Recall: 77.9% F1: 77.27% AUC: 86% |
| Ishimaru et al.[135] | Depression | Classification and Severity Assessment | GCNN | Audio | Precision: 94.75% |
| Das et al.[136] | Depression | Detection | CNN | Audio | Accuracy: 90.26% (DAIC-WOZ) Accuracy: 90.47% (MODMA) |
| Zhang et al.[137] | Depression | Detection | wav2vec 2.0+1D- CNN + LSTM | Speech | Precision: 84.49% (DAIC-WOZ) Precision: 94.83% (MODMA) F1: 79% (DAIC-WOZ) F1: 90.53% (MODMA) |
| Gupta et al.[138] | Depression | Detection | DAttn Conv 2D LSTM | Speech | Accuracy: 97.82% (DAIC-WOZ) Accuracy: 98.91% (SH2-FS) |
| Lin et al.[139] | Depression | Detection | CNN + Transformer | Speech | Specificity: 80.35% Sensitivity: 82.14% |
| Pandey et al.[140] | Depression | Recognition | TFNN | Speech | - |
| Huang et al.[141] | Depression | Recognition | wav2vec 2.0 + Transformer | Speech | Accuracy: 94.81% |
| Wang et al.[142] | Depression | Detection | HuBERT | Speech | Precision: 70.59% Recall: 85.71% F1: 83.15% |
| Tian et al.[143] | Depression | Recognition | CNN | Speech | Accuracy: 87.5% (females) Accuracy: 87% (males) |
| Harati et al.[144] | Depression | Severity Classification | DNN | Audio | AUC: 80% |
| Liu et al.[146] | Depression | Detection | Resnet X-vectors | Speech | Accuracy: 74.72% F1: 76.9% |
| Ravi et al.[147] | Depression | Detection | CNN-LSTM | Speech | F1: 80% |
| Wang et al.[148] | Depression | Detection | Transformer | Speech | Accuracy:96.43% F1: 96.63% |
| Yang et al.[149] | Depression | Diagnosis | BERT + BiLSTM | Audio, text and image | Accuracy: 81.1% Precision: 80.2% Recall: 81% F1: 80.6% |
| Rejaibi et al.[150] | Depression | Recognition and Assessment | LSTM | Audio and video | Accuracy: 95.6% F1: 94% |
| Zhang et al.[151] | Depression | Detection | LLaMA 2 | Speech and text | F1: 84% |
| Tank et al.[152] | Depression | Detection | Whisper + BiLSTM | Audio, text and video | - |
| Patapati et al.[153] | Depression | Classification | GPT-4 + BiLSTM | Audio, text and video | Accuracy: 91.01% Precision: 80% Recall: 92.86% F1: 85.95% |
| He et al.[155] | SCZ | Detection | Attention-based CNN | Speech | Accuracy: 97.37% Precision: 94.99% Recall: 99.25% F1: 96.95% |
| He et al.[156] | SCZ | Detection | DT | Speech | Accuracy: 91.1% ~ 94.6% |
| Chakraborty et al.[157] | SCZ | Prediction | SVM | Speech | Accuracy: 86.36% |
| Premanamin et al.[158] | SCZ | Assessment | CNN | Audio and video | Accuracy: 75% F1: 76.41% AUC: 91.52% |
Depression
Depression is a prevalent psychiatric disorder characterized by persistent negative mood states, psychomotor retardation, and cognitive impairment. These alterations manifest not only in emotional expression but also in speech production, making speech a promising non-invasive biomarker for depression assessment and monitoring.
Traditional ML models with handcrafted features
Early studies predominantly relied on handcrafted acoustic and prosodic features - such as pitch variation, speaking rate, pause patterns, and spectral descriptors - combined with classical ML classifiers. These approaches demonstrated that depressive states are associated with reduced speech energy, slower articulation, and altered prosody.
Across multiple datasets, traditional ML models achieved moderate to strong performance in depression detection and severity estimation[47,123-129]. Ensemble learners and SVM-based classifiers frequently outperformed simpler linear models, indicating that non-linear interactions among speech features are informative for mood state discrimination. Several studies further extended this paradigm to non-clinical or at-risk populations, suggesting that subtle speech changes may precede clinical diagnosis[128].
Despite their interpretability and low computational cost, handcrafted-feature-based approaches are sensitive to feature design, recording conditions, and task variability. Their generalizability across spontaneous speech and diverse populations remains limited.
Overall, traditional ML models provide interpretable insights into depression-related speech alterations but struggle to scale across heterogeneous clinical settings.
DL models
To overcome the limitations of manual feature engineering, DL methods have been increasingly adopted for automatic representation learning. CNN, LSTM, and attention-based architectures enable end-to-end modeling of spectral-temporal patterns in speech.
A broad range of DL architectures demonstrated improved performance over traditional ML baselines, particularly on benchmark datasets such as DAIC-WOZ and MODMA[130-143]. Harati et al.[144] utilized the link between short-term emotions and long-term depressive mood states to construct a predictive model based on emotion-derived features. Experimental results demonstrate that their approach can effectively classify depressive and remission phases during DBS treatment, with an AUC of 0.80. Hybrid CNN-LSTM and attention-based models were especially effective in capturing both local acoustic cues and long-term temporal dependencies. Graph-based and tensor-based models further explored structured representations of speech dynamics, reflecting increasing methodological sophistication.
Speaker embedding techniques, including X-vectors and Residual Neural Network(ResNet)[145] based representations, emerged as a powerful tool for depression detection and severity estimation[146]. However, recent studies cautioned that speaker identity information may introduce bias and privacy risks, motivating efforts to disentangle depression-relevant features from speaker-specific traits[147,148].
Overall, DL-based approaches substantially improve representation learning capacity and robustness but remain vulnerable to dataset bias and limited external validation.
Multimodal fusion
Several studies incorporated multimodal information - such as visual cues, facial action units, and textual content - to enhance diagnostic robustness. Multimodal fusion consistently outperformed unimodal speech-based systems by jointly modeling emotional, behavioral, and linguistic signals[149,150].
Nevertheless, multimodal systems often rely on complex acquisition protocols and controlled experimental settings. Their scalability and feasibility for routine clinical screening require further validation.
Foundation and pre-trained models
Recent work explored foundation models and large language models to leverage large-scale pretraining for depression diagnosis. Pre-trained speech models such as wav2vec 2.0 and Whisper enabled more robust feature extraction from raw audio, while LLM-based frameworks incorporated acoustic cues into multimodal reasoning pipelines[151-153].
These approaches represent a shift toward generalized and adaptable diagnostic frameworks. However, most remain exploratory and retrospective, and their clinical readiness requires careful evaluation.
Methodological synthesis for depression
Speech alterations associated with depression, including reduced prosodic variability, slowed articulation, and diminished emotional expressiveness, are consistent with well-established neurophysiological and psychomotor changes observed in this disorder. Speech-based models should therefore be viewed as complementary tools for screening and monitoring rather than standalone diagnostic systems.
As with other psychiatric applications, many reported results are derived from small or benchmark datasets. Robust validation across diverse populations, recording conditions, and longitudinal settings remains essential before clinical deployment.
SCZ
SCZ is a severe psychiatric disorder with a lifetime prevalence of approximately 1%, characterized by disorganized speech, impaired semantic coherence, and diminished emotional expressivity. These speech abnormalities reflect underlying cognitive and motor dysfunctions, making speech a clinically meaningful behavioral marker for early detection and symptom assessment[154,155].
Existing speech-based studies on SCZ remain relatively limited in number and scope, with most work focusing on handcrafted acoustic features combined with ML models. He et al.[156] demonstrated that carefully designed acoustic markers are strongly associated with negative symptom severity, achieving substantially higher accuracy than traditional pitch- or energy-based features. Their results suggest that SCZ-related speech impairment is multifaceted and cannot be captured by single low-level descriptors alone. Similarly, Chakraborty et al.[157] applied multiple classical classifiers to Open-Source Speech & Music Interpretation by Large-space Extraction (openSMILE) acoustic features and achieved reasonable discrimination between patients and control participants, with further performance gains observed when behavioral signals beyond speech were incorporated.
More recent efforts have begun to move beyond binary classification toward representation learning and severity-oriented modeling. Premanamin et al.[158] proposed a multimodal representation learning framework to estimate SCZ severity scores, marking an important shift from disease detection to clinically relevant symptom quantification. The improved performance achieved through multimodal integration highlights the limitation of speech-only models in capturing the full complexity of SCZ, which manifests across cognitive, affective, and motor domains.
Overall, current speech-based SCZ research suggests that while handcrafted acoustic features remain informative and interpretable, their diagnostic scope is inherently limited. Emerging learning-based and multimodal approaches show promise for modeling symptom severity and heterogeneity, but their clinical applicability is constrained by small sample sizes, heterogeneous protocols, and the absence of large, standardized datasets. Compared with depression and neurological disorders, the adoption of large-scale pre-trained or foundation models in SCZ remains nascent, indicating a clear direction for future research.
RESPIRATORY DISORDERS
Respiratory diseases such as COVID-19, COPD, asthma, and tuberculosis (TB) are often accompanied by characteristic abnormalities in cough, speech, and breathing sounds. These acoustic changes reflect underlying airway obstruction, inflammation, and impaired respiratory control, making audio signals a valuable non-invasive source for screening and monitoring respiratory conditions. Existing studies increasingly explore speech, cough, and breathing sounds - often in a multimodal manner - to extract discriminative features for disease detection and severity assessment. The related literature is summarized in Table 6.
Summary of applications of speech analysis in respiratory disorders
| Reference | Disease | Task | Method | Modality | Performance |
| Xia et al.[159] | COVID-19 | Prediction | VGGish | Audio | AUC: 71% Specificity: 69% Sensitivity: 65% |
| Dash et al.[160] | COVID-19 | Detection | GBM | Speech | Accuracy: 97.8% AUC: 97.6% |
| Zhu et al.[161] | COVID-19 | Detection | SVM | Speech and patient metadata | UAR: 79.9% Specificity: 87.6% Sensitivity: 72.3% |
| Xia et al.[162] | COVID-19 | Detection | VGGish + CNN | Cough, breath, and speech | AUC: 74% Specificity: 69% Sensitivity: 68% |
| Zhang et al.[163] | COVID-19 | Cough detection | ResNet-18 | Cough | AUC: 85.91% Specificity: 91.16% Sensitivity: 59.89% |
| Cai et al.[164] | COVID-19 | Detection | Transformer | Cough | AUC: 83.2% Specificity: 87% Sensitivity: 63% |
| Liu et al.[165] | COVID-19 | Detection | ResNet-18 | Speech | AUC: 76.87% |
| Reiter et al.[166] | COVID-19 | Detection | ResNet-18 + MIL | Cough, speech, breath, vowel phonation, and patient metadata | Accuracy: 92.4% F1: 64.2% AUC: 92.2 ± 0.5% |
| Chen et al.[167] | COVID-19 | Detection | wav2vec 2.0 + BiLSTM | Breath, speech and cough | AUC: 88.44% |
| Dutta et al.[168] | COVID-19 | Detection | BiLSTM | Breath and speech | AUC: 80.5% (Breathing) AUC: 81.5% (Speech) |
| Nallanthighal et al.[169] | COPD | Detection | SVM | Speech | Accuracy: 75.12% Sensitivity: 85% |
| Claxton et al.[170] | COPD | Diagnosis | TDNN + LR | Text and Cough audio | AUC: 89% Specificity: 91% Sensitivity: 82.6% |
| Roy et al.[171] | Asthma | Classification | MLP | Wheezing of Lung Sounds | Accuracy: 98.54% F1: 98.27% Specificity: 98.73% Sensitivity: 98.27% |
| Frost et al.[172] | TB | Cough Classification | BiLSTM | Audio | Accuracy: 80% AUC: 85% Specificity: 81.3% Sensitivity: 77.8% |
COVID-19
Research on speech- and sound-based COVID-19 detection has progressed rapidly, largely driven by the availability of large-scale open datasets and the urgent need for scalable screening tools. The release of the COVID-19 Sounds dataset by Xia et al.[159] marked a key milestone, enabling systematic benchmarking across cough, breathing, and speech modalities and demonstrating that ML models can achieve meaningful discrimination performance even under heterogeneous data collection conditions.
Early studies primarily relied on handcrafted acoustic features combined with classical classifiers, reporting high accuracies under controlled settings[160,161]. However, subsequent work highlighted the limitations of such approaches when applied across datasets or in real-world environments. To address these challenges, learning-based methods increasingly adopted end-to-end and representation learning paradigms, including CNNs, Transformers, and attention-based architectures[162-168]. These models showed improved robustness to noise, data imbalance, and modality variation, particularly when integrating cough, breathing, and speech signals.
Notably, several studies emphasized cross-dataset generalization and noisy-environment robustness, revealing that models trained on curated data often degrade substantially under realistic conditions[162,165]. Multimodal fusion and self-supervised pre-training (e.g., wav2vec 2.0) have been shown to partially mitigate these issues[166-168], although reported performance remains highly sensitive to dataset composition and evaluation protocols. Overall, while audio-based COVID-19 detection demonstrates strong feasibility, its clinical deployment requires cautious interpretation due to dataset bias, variability in recording conditions, and limited prospective validation.
COPD
In COPD research, speech analysis has primarily been explored as a tool for detecting acute exacerbations rather than general diagnosis. Studies consistently report that acoustic features related to voice stability and respiratory control - such as shimmer and syllables per breath - are significantly altered during acute exacerbations compared to stable states[169]. Traditional ML models using handcrafted features achieve moderate accuracy, indicating that speech carries clinically relevant but incomplete information about disease status.
More recent work has incorporated learning-based breathing and speech models to estimate respiratory parameters directly from speech signals, providing a closer link to physiological dysfunction[169]. Importantly, Claxton et al.[170] demonstrated the practical feasibility of deploying speech-based COPD screening via smartphone platforms, combining audio signals with minimal patient-reported information. These findings suggest that, in COPD, speech-based systems may be most effective as decision-support or triage tools, rather than standalone diagnostic solutions.
Asthma
Compared with other respiratory conditions, speech and sound analysis for asthma has received relatively limited attention. Existing work focuses primarily on lung auscultation sounds, where persistent wheezing and abnormal airflow patterns provide strong acoustic cues. Roy et al.[171] showed that representation learning with supervised contrastive objectives can achieve high discrimination performance between asthmatic and healthy lung sounds. While these results are promising, they are derived from relatively constrained experimental settings, and their generalizability to broader clinical populations remains unclear.
Tuberculosis
Research on speech- or cough-based TB detection remains exploratory. Frost et al.[172] demonstrated that temporal modeling of cough signals using RNN-based architectures, particularly Bidirectional LSTM (BiLSTM) with attention mechanisms, can improve classification performance by focusing on diagnostically salient sound segments. These findings support the hypothesis that TB-related cough exhibits distinctive temporal patterns. However, the limited scale of available datasets and the variability of cough characteristics across disease stages constrain current conclusions.
Across respiratory disorders, speech, cough, and breathing sounds provide accessible and physiologically grounded signals for disease screening. While DL and representation learning methods have improved robustness and performance, most reported results are obtained under dataset-specific conditions. Future research should prioritize standardized data collection, cross-dataset evaluation, and clinically meaningful endpoints to ensure that audio-based respiratory assessment methods translate into real-world healthcare settings.
CHALLENGES AND FUTURE PROSPECTS
Despite significant achievements in disease diagnosis using voice analysis, substantial limitations persist across multiple dimensions. Challenges encountered in clinical translation constrain model performance enhancement, while inherent technological limitations remain unresolved. Investigating solutions to these constraints represents a critical future direction for this field. This section delineates the current challenges from both clinical and technical perspectives and proposes potential solutions.
Clinical translation challenges
While speech analysis has demonstrated substantial potential for disease assessment, prediction, and diagnosis, significant barriers continue to hinder its translation from research settings to routine clinical practice. These obstacles span inconsistent data governance, limited clinical validation, privacy concerns, and intrinsic algorithmic constraints. Addressing these challenges is not merely a matter of performance optimization but a prerequisite for safe, reliable, and equitable clinical deployment. This section examines critical challenges from both clinical and technical perspectives and outlines actionable future directions.
Lack of standardized data.
Data heterogeneity remains a fundamental bottleneck for clinical applicability. The absence of unified acquisition protocols leads to substantial variability in recording devices, acoustic environments, and sampling procedures. As a result, speech data exhibit inconsistent characteristics, including fluctuating signal-to-noise ratios (SNRs), frequency responses, and temporal resolutions. Such inconsistencies directly undermine model robustness and limit the generalizability of speech-based AI systems across clinical sites and populations.
Establishing a rigorous and standardized data acquisition framework is therefore essential. This includes defining unified recording parameters (e.g., a sampling rate of 16 kHz and SNR above 30 dB), standardizing acoustic environments, and specifying consistent vocal tasks such as sustained phonation or standardized sentence reading. In addition, automated preprocessing pipelines based on DL, including adaptive noise reduction and normalization, can further mitigate device- and environment-induced variability, thereby improving cross-center data comparability.
Lack of large-scale clinical validation
The current literature is dominated by single-center, retrospective studies with relatively small sample sizes. This design introduces unavoidable selection bias and often leads to optimistic performance estimates that may not generalize to broader clinical settings. Without large-scale, prospective, and multi-center validation, the true diagnostic and prognostic value of speech-based models remains uncertain.
A tiered validation strategy is therefore required. Early-stage studies may focus on single-center pilot cohorts targeting high-prevalence diseases to establish feasibility. Subsequent investigations should expand to large-scale, prospective, multicenter trials that incorporate real-world clinical heterogeneity. The integration of mobile health (mHealth) platforms for continuous, remote voice collection further enables longitudinal monitoring.
Privacy protection risks
Speech signals inherently contain sensitive biometric identifiers, such as voiceprints, as well as implicit information about an individual’s health status. The multi-stage lifecycle of clinical data - including acquisition, transmission, storage, and sharing - amplifies the risk of privacy breaches[173]. Inadequate protection mechanisms may result in ethical concerns and legal liabilities.
FL provides a privacy-preserving paradigm by enabling collaborative model training without sharing raw speech data. In this framework, institutions perform local model updates and transmit only encrypted parameters to a central aggregation server. The combination of FL with differential privacy (DP) mechanisms further reduces the risk of re-identification, ensuring that individual patient contributions cannot be reverse-engineered from shared updates.
Technical application challenges
From a technical perspective, speech-based disease analysis systems face three interrelated challenges: limited interpretability, insufficient generalization across diseases and populations, and inadequate modeling of long-term temporal dynamics. These challenges are largely disease-agnostic at the algorithmic level but manifest differently across clinical scenarios, directly affecting model trustworthiness, robustness, and longitudinal utility. Collectively, they constrain real-world deployment in diagnosis, severity assessment, and disease monitoring.
Lack of interpretability
Despite their strong predictive performance, DL-based speech models often function as opaque systems. The absence of transparent reasoning mechanisms makes it difficult for clinicians to determine which acoustic or linguistic features underpin specific predictions. This lack of interpretability undermines clinical trust and limits integration into evidence-based decision support systems.
SHapley Additive exPlanations Algorithm. SHapley Additive exPlanations (SHAP) is grounded in cooperative game theory and assigns each feature a Shapley value representing its marginal contribution to the model output. By accounting for feature interactions, SHAP provides a global explanation of model behavior. Its visualization tools - such as feature importance plots and dependence maps - enable clinicians to verify whether predictions rely on clinically meaningful speech biomarkers (e.g., jitter, shimmer, articulation rate) rather than spurious correlations. As a result, SHAP facilitates both model auditing and feature-level clinical interpretation. For instance, Shen et al.[174] effectively employed SHAP to elucidate the importance of individual acoustic features in their model's decision-making process.
Local Interpretable Model-agnostic Explanations Algorithm. Local Interpretable Model-Agnostic Explanations (LIME) focuses on explaining individual predictions. It approximates the complex decision boundary of a model using a locally interpretable surrogate, typically a linear model, in the vicinity of a specific input sample. Such instance-level explanations are particularly valuable in clinical contexts, where clinicians must justify and validate algorithmic outputs for individual patients, thereby supporting personalized diagnostic decision-making.
Insufficient generalization capability
Speech-based disease models frequently experience performance degradation when applied to new populations, languages, or clinical environments. Overlapping vocal manifestations across different diseases further complicate discrimination, while cross-lingual variability in prosody, phonetics, and articulation patterns exacerbates domain shift. These limitations significantly restrict scalability and equitable clinical deployment.
Multimodal fusion. Unimodal acoustic analysis captures only a subset of disease-relevant information. Multimodal fusion frameworks address this limitation by integrating complementary signals, including facial expressions, articulatory movements, textual transcripts, and clinical metadata, to construct more comprehensive and robust representations. By jointly modeling heterogeneous modalities, these approaches improve robustness under noisy recording conditions, reduce ambiguity across overlapping disease phenotypes, and enhance generalization across diverse clinical settings. For example, Fang et al.[175] introduced an enhanced Multimodal Fusion Model with multi-level Attention (MFM-Att) for depression detection, significantly improving performance through a comprehensive analysis of audiovisual and textual data.
Self-supervised learning and foundation models. Self-supervised learning (SSL) represents a paradigm shift in speech representation learning. By pre-training models on large-scale, unlabeled speech corpora, SSL methods learn high-level, language-agnostic representations that encode phonetic, prosodic, and temporal patterns. Foundation models such as wav2vec 2.0 and Hidden-Unit BERT (HuBERT) enable efficient adaptation to downstream clinical tasks with limited labeled data. This capability is critical for rare diseases and low-resource languages, substantially improving cross-disease and cross-population generalization.
Limitations in dynamic temporal modeling
Many neurological and psychiatric disorders, including PD and depression, are inherently progressive and require long-term monitoring rather than static classification. However, most existing speech-based approaches rely on short-duration recordings and cross-sectional analysis, neglecting disease trajectories. This limitation restricts clinical utility to screening or coarse assessment and hampers prognosis and treatment monitoring.
Longitudinal trajectory analysis. Longitudinal modeling emphasizes temporal changes in speech biomarkers rather than absolute values at a single time point. By quantifying rates of change - such as temporal slopes of fundamental frequency variability or articulatory precision - models can distinguish pathological progression from normal aging. Williamson et al.[176] demonstrated that tracking articulatory coordination trajectories provides more sensitive indicators of depression severity and treatment response than static features.
Advanced temporal architectures. Conventional classifiers struggle to capture long-range temporal dependencies in speech signals. Advanced sequence modeling approaches, including RNNs, temporal convolutional networks, and Transformer-based architectures, are better suited for this task. Through mechanisms such as gated memory units and attention, these models can capture latent temporal patterns across repeated recordings collected over extended periods. Such architectures enable early detection of disease progression and support longitudinal clinical decision-making.
Clinical implications of technical limitations
The technical limitations discussed above translate directly into constraints on clinical applicability. Limited interpretability weakens clinician confidence and hinders the integration of speech-based models into evidence-based decision support workflows. Insufficient generalization results in population- and context-dependent performance variability, reducing robustness when systems are deployed across languages, institutions, or disease subtypes. Inadequate temporal modeling further confines current approaches to cross-sectional screening, rather than enabling longitudinal disease monitoring and prognostic assessment. Figure 6 provides a schematic overview linking each technical challenge to its proposed solutions and associated clinical impacts, offering a clear roadmap for researchers and practitioners. As illustrated, advances in explainable AI, robust multimodal and foundation models, and longitudinal temporal modeling are not isolated algorithmic improvements, but essential enablers for reliable, clinically meaningful, and scalable deployment of speech-based diagnostic systems.
CONCLUSION
This review summarizes recent research and application progress in speech analysis technology for neurological, psychiatric, and respiratory disorders. It aims to explore the potential of speech analysis in AI-assisted disease assessment, prediction, and diagnosis, and promote the in-depth application of this technology in the medical field. First, the article outlines speech data resources and public datasets. Subsequently, it systematically introduces the core components of the speech-based AI disease diagnosis framework, including data pre-processing, feature engineering, and model construction. Furthermore, the article reviews the current state of speech analysis in the aforementioned disease areas, identifies challenges, proposes potential solutions, and provides guidance for future research directions. Additionally, we hope to leverage existing technological pathways to extend speech analysis to broader disease diagnostic scenarios. Looking ahead, with continuous in-depth research and technological advancements, speech analysis is expected to play a more significant clinical role in AI disease assessment, prediction and diagnosis. It should be noted that although many studies report high diagnostic performance, such results are often obtained under controlled experimental settings and may not directly translate to routine clinical practice. Therefore, current speech-based diagnostic approaches should be regarded as supportive tools, and further large-scale, clinically validated studies are required to ensure robustness and generalizability.
DECLARATIONS
Authors’ contributions
Conceptualization: Xu X, Zhang Y, Zhao L
Methodology: Long, N, Niu Q
Investigation: Zhang Y, Long, N, Niu Q
Formal analysis: Xu X, Zhang Y
Writing - original draft preparation: Xu X, Zhang Y
Writing - review and editing: Xu X, Zhang Y, Zhao L
Supervision: Zhao L, Li J
Funding acquisition: Li J, Yin J, Yang J
Availability of data and materials
Not applicable.
Financial support and sponsorship
This work was supported by the National High-Level Hospital Clinical Research Funding (No. BJ-2023-111).
Conflicts of interest
Xu X and Li J serve as Guest Editors of the Special Issue “Multi-modal Data and AI Technologies in Medical Diagnosis and Surgery” of the journal Artificial Intelligence Surgery. They were not involved in any stage of the editorial process for this manuscript, including reviewer selection, manuscript handling, or decision-making. The other authors declare that they have no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Duffy JR. Motor speech disorders: Substrates, differential diagnosis, and management. Elsevier Health Sciences, 2012. Available from: URL:https://books.google.com/books/about/Motor_Speech_Disorders.html?id=M8t-KgGhjjwC [Last accessed on 13 Feb 2026].
2. Zhao Q, Xu H, Li J, Rajput FA, Qiao L. The application of artificial intelligence in Alzheimer's research. Tsinghua Sci. Technol. 2024;29:13-33.
3. Latif S, Qadir J, Qayyum A, Usama M, Younis S. Speech technology for healthcare: opportunities, challenges, and state of the art. IEEE Rev Biomed Eng. 2021;14:342-56.
4. Aryagopal HRT, Aryagopal
5. Szatloczki G, Hoffmann I, Vincze V, Kalman J, Pakaski M. Speaking in Alzheimer’s disease, is that an early sign? Front Aging Neurosci. 2015;7:195.
6. Gosztolya G, Vincze V, Tóth L, Pákáski M, Kálmán J, Hoffmann I. Identifying mild cognitive impairment and mild Alzheimer’s disease based on spontaneous speech using ASR and linguistic features. Comput Speech Lang. 2019;53:181-97.
7. Shin D, Cho WI, Park CHK, et al. Detection of minor and major depression through voice as a biomarker using machine learning. J Clin Med. 2021;10:3046.
8. Dumpala SH, Dikaios K, Rodriguez S, et al. Manifestation of depression in speech overlaps with characteristics used to represent and recognize speaker identity. Sci Rep. 2023;13:11155.
11. Cover T, Hart P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory. 1967;13:21-7.
12. Sangeetha KB, Mengani S, Shaik AB, Reddy Kasarla V. A Parkinson’s disease detection using support vector machine in machine learning. 2024 Eighth International Conference on Parallel, Distributed and Grid Computing (PDGC); 2024 Dec 18-20; Waknaghat, Solan, India. IEEE; 2024. pp. 640-4.
13. Haq AU, Li JP, Memon MH, et al. Feature selection based on L1-norm support vector machine and effective recognition system for Parkinson’s disease using voice recordings. IEEE Access. 2019;7:37718-34.
14. LeCun Y, Bottou L, Bengio Y, Haffner P. Gradient-based learning applied to document recognition. Proceedings of the IEEE. 2002;86:2278.
15. Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need. In The Thirty-First Annual Conference on Neural Information Processing Systems, Advances in Neural Information Processing Systems, Long Beach, USA. 2017 Dec 4-9; Neural Information Processing Systems Foundation, Inc.; 2017. Vol. 30, pp. 6000-10.
16. Frid A, Kantor A, Svechin D, Manevitz LM. Diagnosis of Parkinson’s disease from continuous speech using deep convolutional networks without manual selection of features. 2016 IEEE International Conference on the Science of Electrical Engineering (ICSEE); 2016 Nov 16-18; EILAT, Israel. IEEE; 2016. pp. 1-4.
17. Radford A, Narasimhan K, Salimans T, Sutskever I. Improving language understanding by generative pre-training. In: (2018). Available from: https://api.semanticscholar.org/CorpusID:49313245 [Last accessed on 13 Feb 2026].
18. Devlin J, Chang M, Lee K, Toutanova K. BERT: pre-training of deep bidirectional transformers for language understanding. In Proceedings of the 2019 Conference of the North; Minneapolis, MN, USA. 2019 Jun 2-7; Association for Computational Linguistics; 2019. Vol. 1, pp. 4171-86.
19. Idrisoglu A, Dallora AL, Anderberg P, Berglund JS. Applied machine learning techniques to diagnose voice-affecting conditions and disorders: systematic literature review. J Med Internet Res. 2023;25:e46105.
20. De Silva U, Madanian S, Olsen S, et al. Clinical decision support using speech signal analysis: systematic scoping review of neurological disorders. J Med Internet Res. 2025;27:e63004.
21. Hecker P, Steckhan N, Eyben F, Schuller BW, Arnrich B. Voice analysis for neurological disorder recognition-a systematic review and perspective on emerging trends. Front Digit Health. 2022;4:842301.
22. Khaskhoussy R, Ayed YB. Speech processing for early Parkinson’s disease diagnosis: machine learning and deep learning-based approach. Soc Netw Anal Min. 2022;12:73.
23. Ding K, Chetty M, Noori Hoshyar A, Bhattacharya T, Klein B. Speech based detection of Alzheimer’s disease: a survey of AI techniques, datasets and challenges. Artif Intell Rev. 2024;57:325.
24. Birger Moell et al. The order in speech disorder: a scoping review of state of the art machine learning methods for clinical speech classification. ArXiv 2025;arXiv:2503.04802. Available from https://doi.org/10.48550/arXiv.2503.04802.
25. Radford A, Kim JW, Xu T, Brockman G, Mcleavey C, Sutskever I. Robust speech recognition via large-scale weak supervision. In the 40th International Conference on Machine Learning: Proceedings of the 40th International Conference on Machine Learning, Honolulu, Hawaii, USA. 2023, Jul 23-29; MLResearch Press; 2023. Vol 202, pp. 28492-518. Available from https://proceedings.mlr.press/v202/radford23a.html [Last accessed on 13 February 2026].
26. Baevski A, Zhou Y, Mohamed A, Auli M. wav2vec 2.0: A framework for self-supervised learning of speech representations. Adv Neural Inf Process Syst (NeurIPS). 2020;33:12449-60. https://proceedings.neurips.cc/paper_files/paper/2020/file/92d1e1eb1cd6f9fba3227870bb6d7f07 [Last accessed on 13 February 2026].
27. Tricco AC, Lillie E, Zarin W, et al. PRISMA extension for scoping reviews (PRISMA-ScR): checklist and explanation. Ann Intern Med. 2018;169:467-73.
28. Little MA, McSharry PE, Hunter EJ, Spielman J, Ramig LO. Suitability of dysphonia measurements for telemonitoring of Parkinson’s disease. IEEE Trans Biomed Eng. 2009;56:1015.
29. Mendes-Laureano J, Gómez-García JA, Guerrero-López A, et al. Neurovoz: a Castillian Spanish corpus of parkinsonian speech. Sci Data. 2024;11:1367.
30. Jaeger H, Trivedi D, Stadtschnitzer M. Mobile Device Voice Recordings at King’s College London (MDVR-KCL) from both early and advanced Parkinson’s disease patients and healthy controls. Available from https://zenodo.org/record/2867216 [Last accessed on 13 February 2026].
31. Dementiabank database guide. Available from https://dementia.talkbank.org/.
32. Rusko M, Sabo R, Trnka M, et al. Slovak database of speech affected by neurodegenerative diseases. Sci Data. 2024;11:1320.
33. Gratch J, Artstein R, Lucas GM, et al. The distress analysis interview corpus of human and computer interviews In the Ninth International Conference on Language Resources and Evaluation, Proceedings of the Ninth International Conference on Language Resources and Evaluation (LREC'14), Reykjavik, Iceland 2014 May 26-31; European Language Resources Association: Luxembourg, Fance, 2014: pp. 3123-8. Available from https://dcapswoz.ict.usc.edu/wp-content/uploads/2022/02/DAICWOZDepression_Documentation.pdf [Last accessed on 13 February 2026].
34. Cai H, Yuan Z, Gao Y, et al. A multi-modal open dataset for mental-disorder analysis. Sci Data. 2022;9:178.
35. He L, Fu J, Li Y, Xiong X, Zhang J. WNSA-Net: an axial-attention-based network for schizophrenia detection using wideband and narrowband spectrograms. IEEE/ACM Trans. Audio Speech Lang. Process. 2023;31:721-33.
36. Bhattacharya D, Sharma NK, Dutta D, et al. Coswara: A respiratory sounds and symptoms dataset for remote screening of SARS-CoV-2 infection. Sci Data. 2023;10:397.
37. Orlandic L, Teijeiro T, Atienza D. The COUGHVID crowdsourcing dataset, a corpus for the study of large-scale cough analysis algorithms. Sci Data. 2021;8:156.
38. Xia T, Spathis D, Ch J, et al. COVID-19 sounds: a large-scale audio dataset for digital respiratory screening. Thirty-fifth conference on neural information processing systems datasets and benchmarks track (round 2). 2021. Available from https://openreview.net/forum?id=9KArJb4r5ZQ [Last accessed on 13 February 2026].
39. Fraiwan M, Fraiwan L, Khassawneh B, Ibnian A. A dataset of lung sounds recorded from the chest wall using an electronic stethoscope. Data Brief. 2021;35:106913.
40. Upadhya SS, Cheeran A. Discriminating parkinson and healthy people using phonation and cepstral features of speech. Procedia Comput Sci. 2018;143:197-202.
41. Meghanani A, C
42. Haider F, De La Fuente S, Luz S. An assessment of paralinguistic acoustic features for detection of Alzheimer’s Dementia in spontaneous speech. IEEE J. Sel. Top. Signal Process. 2020;14:272-81.
43. Vizza P, Tradigo G, Mirarchi D, et al. Methodologies of speech analysis for neurodegenerative diseases evaluation. Int J Med Inf. 2019;122:45-54.
44. Faragó P, Ștefănigă SA, Cordoș CG, et al. CNN-based identification of Parkinson’s disease from continuous speech in noisy environments. Bioengineering (Basel). 2023:10.
45. Gillivan-Murphy P, Miller N, Carding P. Voice tremor in Parkinson’s disease: an acoustic study. J Voice. 2019;33:526-35.
46. Bayestehtashk A, Asgari M, Shafran I, McNames J. Fully automated assessment of the severity of Parkinson’s disease from speech. Comput Speech Lang. 2015;29:172-85.
47. Berardi M, Brosch K, Pfarr JK, et al. Relative importance of speech and voice features in the classification of schizophrenia and depression. Transl Psychiatry. 2023;13:298.
49. Wang M, Wen Y, Mo S, et al. Distinctive acoustic changes in speech in Parkinson’s disease. Comput Speech Lang. 2022;75:101384.
50. Hason L, Krishnan S. Spontaneous speech feature analysis for Alzheimer’s disease screening using a random forest classifier. Front Digit Health. 2022;4:901419.
51. Karan B. Speech-based Parkinson’s disease prediction using XGBoost-Based features selection and the stacked ensemble of classifiers. J Inst Eng India Ser B. 2023;104:475-83.
52. Tunc HC, Sakar CO, Apaydin H, et al. Estimation of Parkinson’s disease severity using speech features and extreme gradient boosting. Med Biol Eng Comput. 2020;58:2757-73.
53. Rumelhart DE, Hinton GE, Williams RJ. Learning representations by back-propagating errors. Nature. 1986;323:533-536.
55. Cho K, van Merriënboer B, Gulcehre C, et al. Learning phrase representations using RNN encoder-decoder for statistical machine translation. In Moschitti A, Pang B, Daelemans W, Editors. EMNLP 2014: Proceedings of the 2014 conference on empirical methods in natural language processing (EMNLP); 2014 Oct 25-29; Doha, Qatar. Association for Computational Linguistics; 2014. pp.1724-34.
56. Chu Y, Xu J, Zhou X, et al. Qwen-audio: advancing universal audio understanding via unified large-scale audio-language models. ArXiv 2023;arXiv:2311.07919. Available online: https://doi.org/10.48550/arXiv.2311.07919.
57. Touvron H, Martin L, Stone K, et al. Llama 2: open foundation and fine-tuned chat models. ArXiv 2023;arXiv:2307.09288. Available online: https://doi.org/10.48550/arXiv.2307.09288.
58. Wang T, Zhou L, Zhang Z, et al. VioLA: Conditional language models for speech recognition, synthesis, and translation. IEEE/ACM Trans. Audio Speech Lang Process. 2024;32:3709-16.
59. Zhao L, Li J, Xu X, et al. A deep learning-based ocular structure segmentation for assisted myasthenia gravis diagnosis from facial images. Tsinghua Sci Technol. 2025;30:2592-605.
60. Steinmetz JD, Seeher KM, Schiess N, et al. Global, regional, and national burden of disorders affecting the nervous system, 1990-2021: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Neurol. 2024;23:344-81.
61. Moro-Velazquez L, Gomez-Garcia JA, Godino-Llorente JI, et al. Phonetic relevance and phonemic grouping of speech in the automatic detection of Parkinson’s disease. Sci Rep. 2019;9:19066.
62. Shastry KA. An ensemble nearest neighbor boosting technique for prediction of Parkinson’s disease. Healthc Anal. 2023;3:100181.
63. Mohammadi AG, Mehralian P, Naseri A, Sajedi H. Parkinson’s disease diagnosis: the effect of autoencoders on extracting features from vocal characteristics. Array. 2021;11:100079.
64. Govindu A, Palwe S. Early detection of Parkinson’s disease using machine learning. Procedia Comput Sci. 2023;218:249-61.
65. Mahesh TR, Bhardwaj R, Khan SB, et al. An artificial intelligence-based decision support system for early and accurate diagnosis of Parkinson’s disease. Decis Anal J. 2024;10:100381.
66. Mudawi NA. Developing a model for Parkinson’s disease detection using machine learning algorithms. CMC. 2024;79:4945-62.
67. Jain V, Singh R, Gupta A. Exploring binary classification models for Parkinson’s disease detection. Procedia Comput Sci. 2024;235:2332-41.
68. Wang Y, Li F, Zhang X, Wang P, Li Y, Zhang Y. Intra-subject enveloped multilayer fuzzy sample compression for speech diagnosis of Parkinson’s disease. Med Biol Eng Comput. 2024;62:371-88.
69. Deepa P, Khilar R. Parkinson’s disease detection and classification: leveraging voice features and ensemble methods with feature selection and ERT classifier. Procedia Computer Sci. 2024;235:1695-706.
70. Torghabeh F, Hosseini SA, Ahmadi Moghadam E. Enhancing Parkinson’s disease severity assessment through voice-based wavelet scattering, optimized model selection, and weighted majority voting. Med Nov Technol Devices. 2023;20:100266.
71. Laudis LL, A LF, Jambek AB. A nature inspired optimization algorithm for Parkinson’s disease classification through speech analysis. Procedia Comput Sci. 2024;235:840-51.
72. Yuan L, Liu Y, Feng H. Parkinson disease prediction using machine learning-based features from speech signal. SOCA. 2023;18:101-7.
73. Wrobel K. Diagnosing Parkinson’s disease by means of ensemble classification of patients’ voice samples. Procedia Comput Sci. 2021;192:3905-14.
74. Liu W, Liu J, Peng T, et al. Prediction of Parkinson’s disease based on artificial neural networks using speech datasets. J Ambient Intell Human Comput. 2022;14:13571-84.
75. Quan C, Ren K, Luo Z, Chen Z, Ling Y. End-to-end deep learning approach for Parkinson’s disease detection from speech signals. Biocybern Biomed Eng. 2022;42:556-74.
76. Tayebi Arasich S, Rios-Urrego CD, Noeth E, et al. Federated learning for secure development of AI models for Parkinson’s disease detection using speech from different languages. ArXiv 2023;arXiv:2305.11284. Available online: https://doi.org/10.48550/arXiv.2305.11284.
77. Wang Y, Li F, Zhang X, Wang P, Li Y. Subject enveloped deep sample fuzzy ensemble learning algorithm of Parkinson’s speech data. ArXiv 2021;arXiv:2111.09014. Available online: https://doi.org/10.48550/arXiv.2111.09014.
78. Xu Z, Wang J, Zhang Y, He X. Voiceprint recognition of Parkinson patients based on deep learning. ArXiv 2018;arXiv:1812.06613. Available online: https://doi.org/10.48550/arXiv.1812.06613.
79. Pandey PVK, Sahu SS, Karan B, Mishra SK. Parkinson Disease Prediction Using CNN-LSTM Model from voice signal. SN COMPUT. SCI. 2024;5:381.
80. Mishra S, Jena L, Mishra N, Chang HT. PD-DETECTOR: a sustainable and computationally intelligent mobile application model for Parkinson’s disease severity assessment. Heliyon. 2024;10:e34593.
81. Jeancolas L, Petrovska-Delacrétaz D, Mangone G, et al. X-Vectors: new quantitative biomarkers for early Parkinson’s disease detection from speech. Front Neuroinform. 2021;15:578369.
82. Chronowski M, Klaczynski M, Dec-Cwiek M, Porebska K, et al. Parkinson’s disease diagnostics using AI and natural language knowledge transfer. ArXiv 2022;arXiv:2204.12559. Available online: https://doi.org/10.48550/arXiv.2204.12559.
83. Khaskhoussy R, Ben Ayed Y. A deep convolutional autoencoder-based approach for Parkinson’s disease diagnosis through speech signals. In: Chen W, Yao L, Cai T, Pan S, Shen T, Li X, Editors. Advanced Data Mining and Applications. Cham: Springer Nature Switzerland; 2022. pp. 15-26.
84. Hireš M, Gazda M, Drotár P, Pah ND, Motin MA, Kumar DK. Convolutional neural network ensemble for Parkinson’s disease detection from voice recordings. Comput Biol Med. 2022;141:105021.
85. Akila B, Nayahi JJV. Parkinson classification neural network with mass algorithm for processing speech signals. Neural Comput Applic. 2024;36:10165-81.
86. Palakayala AR, P K. Differentiating Parkinson’s disease from other neuro diseases and diagnosis using deep learning with nature inspired algorithms and ensemble learning. Procedia Comput Sci. 2024;235:588-97.
87. Skibińska J, Hosek J. Computerized analysis of hypomimia and hypokinetic dysarthria for improved diagnosis of Parkinson’s disease. Heliyon. 2023;9:e21175.
88. Tusar MTHK, Islam MT, Sakil AH. An experimental study for early diagnosing Parkinson’s disease using machine learning. ArXiv 2023;arXiv:2310.13654. Available online: https://doi.org/10.48550/arXiv.2310.13654.
89. Yousif NR, Balaha HM, Haikal AY, El-Gendy EM. A generic optimization and learning framework for Parkinson disease via speech and handwritten records. J Ambient Intell Humaniz Comput. ;2022:1-21.
90. Yuan J, Cai X, Church K. Pause-encoded language models for recognition of Alzheimer’s disease and emotion. In ICASSP 2021: Proceedings of 2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2021 Jun 6-11; Toronto, ON, Canada. IEEE; 2021. pp. 7293-7.
91. Koo J, Lee JH, Pyo J, Jo Y, Lee K. Exploiting multi-modal features from pre-trained networks for Alzheimer's dementia recognition. ArXiv 2020;arXiv:2009.04070. Available online: https://doi.org/10.48550/arXiv.2009.04070.
92. Li J, Yu J, Ye Z, et al. A comparative study of acoustic and linguistic features classification for Alzheimer’s disease detection. In ICASSP 2021: Proceedings of 2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2021 Jun 6-11; Toronto, ON, Canada. IEEE; 2021. pp. 6423-7.
93. Pappagari R, Cho J, Moro-velázquez L, Dehak N. Using state of the art speaker recognition and natural language processing technologies to detect Alzheimer’s disease and assess its severity. Interspeech. ;2020:2177-81.
94. Cummins N, Pan Y, Ren Z, et al. A comparison of acoustic and linguistics methodologies for Alzheimer’s Dementia recognition. Interspeech. ;2020:2182-6.
95. Khodabakhsh A, Yesil F, Guner E, Demiroglu C. Evaluation of linguistic and prosodic features for detection of Alzheimer’s disease in Turkish conversational speech. J AUDIO SPEECH MUSIC PROC. 2015;2015:9.
96. Konig A, Satt A, Sorin A, et al. Use of speech analyses within a mobile application for the assessment of cognitive impairment in elderly people. Curr Alzheimer Res. 2018;15:120-9.
97. Li B, Hsu YT, Rudzicz F. Detecting dementia in mandarin Chinese using transfer learning from a parallel corpus. ArXiv 2019;arXiv:1903.00933. Available online: https://doi.org/10.48550/arXiv.1903.00933.
98. Ammar RB, Ayed YB. Language-related features for early detection of Alzheimer disease. Procedia Comput Sci. 2020;176:763-70.
99. Nasrolahzadeh M, Mohammadpoori Z, Haddadnia J. Analysis of mean square error surface and its corresponding contour plots of spontaneous speech signals in Alzheimer’s disease with adaptive wiener filter. Comput Hum Behav. 2016;61:364-71.
100. König A, Satt A, Sorin A, et al. Automatic speech analysis for the assessment of patients with predementia and Alzheimer’s disease. Alzheimers Dement (Amst). 2015;1:112-24.
101. López-de-ipiña K, Alonso JB, Solé-casals J, et al. On automatic diagnosis of Alzheimer’s disease based on spontaneous speech analysis and emotional temperature. Cogn Comput. 2013;7:44-55.
102. García-Gutiérrez F, Alegret M, Marquié M, et al. Unveiling the sound of the cognitive status: Machine Learning-based speech analysis in the Alzheimer’s disease spectrum. Alzheimers Res Ther. 2024;16:26.
103. Kim TM, Son J, Chun JW, et al. Comparison of AI with and without hand-crafted features to classify Alzheimer’s disease in different languages. Comput Biol Med. 2024;180:108950.
104. Chien YW, Hong SY, Cheah WT, Yao LH, Chang YL, Fu LC. An automatic assessment system for Alzheimer’s disease based on speech using feature sequence generator and recurrent neural network. Sci Rep. 2019;9:19597.
105. Roshanzamir A, Aghajan H, Soleymani Baghshah M. Transformer-based deep neural network language models for Alzheimer’s disease risk assessment from targeted speech. BMC Med Inform Decis Mak. 2021;21:92.
106. Dong Z, Zhang Z, Xu W, Han J, Ou J, Schuller BW. HAFFormer: A hierarchical attention-free framework for Alzheimer’s disease detection from spontaneous speech. In ICASSP 2024: Proceedings of 2024 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2024 Apr 14-19; Seoul, Korea, Republic of. IEEE; 2024. pp. 11246-50.
107. Liu N, Yuan Z, Tang Q. Improving Alzheimer’s disease detection for speech based on feature purification network. Front Public Health. 2021;9:835960.
108. Liu Z, Guo Z, Ling Z, Li Y. Detecting Alzheimer’s disease from speech using neural networks with bottleneck features and data augmentation. In ICASSP 2021: Proceedings of 2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2021 Jun 6-11; Toronto, ON, Canada. IEEE; 2021. pp. 7323-7.
109. Ahn K, Cho M, Kim SW, et al. Deep learning of speech data for early detection of Alzheimer’s disease in the elderly. Bioengineering (Basel). 2023;10:1093.
110. Farazi S, Shekofteh Y. Voice pathology detection on spontaneous speech data using deep learning models. Int J Speech Technol. 2024;27:739-51.
111. Mittal A, Sahoo S, Datar A, Kadiwala J, Shalu H, Mathew J. Multi-modal detection of Alzheimer’s disease from speech and text[J]. ArXiv 2021 arXiv:2012.00096. Available online: https://doi.org/10.48550/arXiv.2012.00096.
112. Haulcy R, Glass J. Classifying Alzheimer’s disease using audio and text-based representations of speech. Front Psychol. 2020;11:624137.
113. Li H, Zeng W, Dai Y, Chen C, Hu L, Yin J. Judgment of Alzheemer’s desease based on multi-feature mixed model. In PRAI 2022: Proceedings of 2022 5th International Conference on Pattern Recognition and Artificial Intelligence (PRAI). 2022 Aug 19-21; Chengdu, China. IEEE, 2022. pp. 1239-44.
114. Jang H, Soroski T, Rizzo M, et al. Classification of Alzheimer’s disease leveraging multi-task machine learning analysis of speech and eye-movement data. Front Hum Neurosci. 2021;15:716670.
115. Ablimit A, Scholz K, Schultz T. Deep learning approaches for detecting Alzheimer’s Dementia from conversational speech of ILSE study. Interspeech. ;2022:3348-52.
116. Martinc M, Haider F, Pollak S, Luz S. Temporal integration of text transcripts and acoustic features for Alzheimer’s diagnosis based on spontaneous speech. Front Aging Neurosci. 2021;13:642647.
117. Mahajan P, Baths V. Acoustic and language based deep learning approaches for Alzheimer’s Dementia detection from spontaneous speech. Front Aging Neurosci. 2021;13:623607.
118. Zhang X, Fu W, Liang M. Soft-weighted CrossEntropy loss for continous Alzheimer’s disease detection. ArXiv 2024;arXiv:2402.11931. Available online: https://doi.org/10.48550/arXiv.2402.11931.
119. Li J, Zhang W. Whisper-based transfer learning for Alzheimer disease classification: leveraging speech segments with full transcripts as prompts. ICASSP 2024 - 2024 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2024 Apr 14-19; Seoul, Korea, Republic of. IEEE; 2024. pp. 11211-5.
120. Bang JU, Han SH, Kang BO. Alzheimer’s disease recognition from spontaneous speech using large language models. ETRI J. 2024;46:96-105.
121. Cui Z, Wu W, Zhang W, Wu J, Zhang C. Transferring speech-generic and depression-specific knowledge for Alzheimer’s disease detection. 2023 IEEE Automatic Speech Recognition and Understanding Workshop (ASRU); 2023 Dec 16-20; Taipei, Taiwan. IEEE; 2023. pp. 1-8.
122. Fan Y, Fan A, Yang Z, Fan D. Global burden of mental disorders in 204 countries and territories, 1990-2021: results from the global burden of disease study 2021. BMC Psychiatry. 2025;25:486.
123. Scibelli F, Roffo G, Tayarani M, et al. Depression speaks: automatic discrimination between depressed and non-depressed speakers based on nonverbal speech features. In ICASSP 2018: Proceedings of 2018 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). 2018 Apr 15-20; Calgary, Canada. IEEE, 2018. pp. 6842-46.
124. Jiang H, Hu B, Liu Z, et al. Detecting depression using an ensemble logistic regression model based on multiple speech features. Comput Math Methods Med. 2018;2018:6508319.
125. Xu S, Yang Z, Chakraborty D, et al. Identifying psychiatric manifestations in schizophrenia and depression from audio-visual behavioural indicators through a machine-learning approach. Schizophrenia (Heidelb). 2022;8:92.
126. Shankayi R, Vali M, Salimi M, Malekshahi M. Identifying depressed from healthy cases using speech processing. In ICBME 2012: Proceedings of 2012 19th Iranian Conference of Biomedical Engineering (ICBME). 2012 Dec 20-21; Tehran, Iran. IEEE, 2012: pp. 242-5.
127. Zulfiker MS, Kabir N, Biswas AA, Nazneen T, Uddin MS. An in-depth analysis of machine learning approaches to predict depression. Curr Res Behav Sci. 2021;2:100044.
128. König A, Tröger J, Mallick E, et al. Detecting subtle signs of depression with automated speech analysis in a non-clinical sample. BMC Psychiatry. 2022;22:830.
129. Kim K, Ryu JI, Lee BJ, et al. A machine-learning-algorithm-based prediction model for psychotic symptoms in patients with depressive disorder. J Pers Med. 2022;12:1218.
130. He L, Cao C. Automated depression analysis using convolutional neural networks from speech. J Biomed Inform. 2018;83:103-11.
131. Chlasta K, Wołk K, Krejtz I. Automated speech-based screening of depression using deep convolutional neural networks. Procedia Comput Sci. 2019;164:618-28.
132. Muzammel M, Salam H, Hoffmann Y, Chetouani M, Othmani A. AudVowelConsNet: a phoneme-level based deep CNN architecture for clinical depression diagnosis. Mach Learn Appl. 2020;2:100005.
133. Srimadhur N, Lalitha S. An end-to-end model for detection and assessment of depression levels using speech. Procedia Comput Sci. 2020;171:12-21.
134. Kim AY, Jang EH, Lee SH, Choi KY, Park JG, Shin HC. Automatic depression detection using smartphone-based text-dependent speech signals: deep convolutional neural network approach. J Med Internet Res. 2023;25:e34474.
135. Ishimaru M, Okada Y, Uchiyama R, Horiguchi R, Toyoshima I. Classification of depression and its severity based on multiple audio features using a graphical convolutional neural network. Int J Environ Res Public Health. 2023;20:1588.
136. Das AK, Naskar R. A deep learning model for depression detection based on MFCC and CNN generated spectrogram features. Biomed Signal Process Control. 2024;90:105898.
137. Zhang X, Zhang X, Chen W, Li C, Yu C. Improving speech depression detection using transfer learning with wav2vec 2.0 in low-resource environments. Sci Rep. 2024;14:9543.
138. Gupta S, Agarwal G, Agarwal S, Pandey D. Depression detection using cascaded attention based deep learning framework using speech data. Multimed Tools Appl. 2024;83:66135-73.
139. Lin Y, Liyanage BN, Sun Y, et al. A deep learning-based model for detecting depression in senior population. Front Psychiatry. 2022;13:1016676.
140. Pandey SK, Shekhawat HS, Prasanna SRM, Bhasin S, Jasuja R. A deep tensor-based approach for automatic depression recognition from speech utterances. PLoS ONE. 2022;17:e0272659.
141. Huang X, Wang F, Gao Y, et al. Depression recognition using voice-based pre-training model. Sci Rep. 2024;14:12734.
142. Wang J, Ravi V, Flint J, Alwan A. Speechformer-CTC: Sequential modeling of depression detection with speech temporal classification. Speech Commun. 2024:163.
143. Tian H, Zhu Z, Jing X. Deep learning for depression recognition from speech. Mobile Netw Appl. 2023;29:1212-27.
144. Harati S, Crowell A, Mayberg H, Nemati S. Depression severity classification from speech emotion. Annu Int Conf IEEE Eng Med Biol Soc. 2018;2018:5763-6.
145. He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2016 Jun 27-30; Las Vegas, NV, USA. IEEE; 2016. pp. 770-8.
146. Liu Z, Yu H, Li G, et al. Ensemble learning with speaker embeddings in multiple speech task stimuli for depression detection. Front Neurosci. 2023;17:1141621.
147. Ravi V, Wang J, Flint J, Alwan A. Enhancing accuracy and privacy in speech-based depression detection through speaker disentanglement. Comput Speech Lang. 2024:86.
148. Wang H, Ye J, Yu Y, Lu L, Yuan L, Wang Q. MFE-Former: disentangling emotion-identity dynamics via self-supervised learning for enhancing speech-driven depression detection. IEEE J Biomed Health Inform. 2025:1-12.
149. Yang S, Cui L, Wang L, Wang T, You J. Enhancing multimodal depression diagnosis through representation learning and knowledge transfer. Heliyon. 2024;10:e25959.
150. Rejaibi E, Komaty A, Meriaudeau F, Agrebi S, Othmani A. MFCC-based recurrent neural network for automatic clinical depression recognition and assessment from speech. Biomed Signal Process Control. 2022;71:103107.
151. Zhang X, Liu H, Xu K, et al. When LLMs meets acoustic landmarks: an efficient approach to integrate speech into large language models for depression detection. arXiv2024;arXiv:2402.13276. Available online: https://doi.org/10.48550/arXiv.2402.13276.
152. Tank C, Pol S, Katoch V, Mehta S, Anand A. Depression detection and analysis using large language models on textual and audio-visual modalities. ArXiv 2024;arXiv:2407.06125. Available online: https://doi.org/10.48550/arXiv.2407.06125.
153. Patapati, SV. Integrating large language models into a tri-modal architecture for automated depression classification on the DAIC-WOZ. ArXiv 2024;arXiv:2407.19340. Available online: https://doi.org/10.48550/arXiv.2407.19340.
154. Parola A, Simonsen A, Bliksted V, Fusaroli R. Voice patterns in schizophrenia: A systematic review and Bayesian meta-analysis. Schizophr Res. 2020;216:24-40.
155. He L, Fu J, Li Y, Xiong X, Zhang J. WNSA-Net: An axial-attention-based network for schizophrenia detection using wideband and narrowband spectrograms. IEEE/ACM Trans Audio Speech Lang Process. 2023;31:721-33.
156. He F, He L, Zhang J, Li Y, Xiong X. Automatic Detection of Affective Flattening in Schizophrenia: Acoustic Correlates to Sound Waves and Auditory Perception. IEEE/ACM Trans Audio Speech Lang Process. 2021;29:3321-34.
157. Chakraborty D, Yang Z, Tahir Y, Maszczyk T, Dauwels J, Thalmann N. Prediction of negative symptoms of schizophrenia from emotion related low-level speech signals. In: 2018 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2018, pp. 6024-8.
158. Premanamin G, Espy-Wilson C. Self-supervised multimodal speech representations for the assessment of Schizophrenia symptoms. In: ICASSP 2025-2025 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2025, pp. 1-5.
159. Tong Xia et al. COVID-19 sounds: a large-scale audio dataset for digital respiratory screening. Thirty-fifth conference on neural information processing systems datasets and benchmarks track (round 2). 2021. Available from https://openreview.net/forum?id=9KArJb4r5ZQ [Last accessed on 13 February 2026].
160. Dash TK, Chakraborty C, Mahapatra S, Panda G. Gradient boosting machine and efficient combination of features for speech-based detection of COVID-19. IEEE J Biomed Health Inform. 2022;26:5364-71.
161. Zhu Y, Falk TH. Fusion of modulation spectral and spectral features with symptom metadata for improved speech-based COVID-19 detection. In: ICASSP 2022-2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2022, pp. 8997-9001.
162. Xia T, Han J, Qendro L, Dang T, Mascolo C. Uncertainty-aware covid-19 detection from imbalanced sound data. ArXiv 2021;arXiv:2104.02005. Available online: https://doi.org/10.48550/arXiv.2104.02005.
163. Zhang X, Shen J, Zhou J. et al. Robust cough feature extraction and classification method for COVID-19 cough detection based on vocalization characteristics. Interspeech. ;2022:2168-72.
164. Cai C, Liu B, Tao J, Tian Z, Lu J, Wang K. End-to-end network based on transformer for automatic detection of COVID-19. In: ICASSP 2022-2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2022, pp. 9082-6.
165. Liu S, Mallol-Ragolta A, Schuller BW. COVID-19 detection from speech in noisy conditions. In: ICASSP 2023-2023 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2023, pp. 1-5.
166. Reiter M, Pernkopf F. Acoustic COVID-19 detection using multiple instance learning. IEEE J Biomed Health Inform. 2025;29:620-30.
167. Chen XY, Zhu QS, Zhang J, Dai LR. Supervised and self-supervised pretraining based COVID-19 detection using acoustic breathing/cough/speech signals. In: ICASSP 2022-2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2022, pp. 561-5.
168. Dutta D, Bhattacharya D, Ganapathy S, Poorjam AH, Mittal D, Singh M. Interpretable acoustic representation learning on breathing and speech signals for COVID-19 detection. ArXiv 2022;arXiv:2206.13365. Available online: https://doi.org/10.48550/arXiv.2206.13365 [accessed 13 February 2026].
169. Srikanth Nallanthighal V, Härmä A, Strik H. Detection of COPD exacerbation from speech: comparison of acoustic features and deep learning based speech breathing models. In: ICASSP 2022-2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2022, pp. 9097-101.
170. Claxton S, Porter P, Brisbane J, et al. Identifying acute exacerbations of chronic obstructive pulmonary disease using patient-reported symptoms and cough feature analysis. NPJ Digit Med. 2021;4:107.
171. Roy A, Satija U. AsthmaSCELNet: a lightweight supervised contrastive embedding learning framework for asthma classification using lung sounds. entropy. ;2023:1282:100.
172. Frost G, Theron G, Niesler T. TB or not TB? Acoustic cough analysis for tuberculosis classification. ArXiv 2022;arXiv:2209.00934. Available online: https://doi.org/10.48550/arXiv.2209.00934.
173. Guan Y, Wen P, Li J, et al. Deep learning blockchain integration framework for ureteropelvic junction obstruction diagnosis using ultrasound images. Tsinghua Sci Technol. 2023;29:1-12.
174. Shen M, Mortezaagha P, Rahgozar A. Explainable artificial intelligence to diagnose early Parkinson’s disease via voice analysis. Sci Rep. 2025;15:11687.
175. Fang M, Peng S, Liang Y, Hung C, Liu S. A multimodal fusion model with multi-level attention mechanism for depression detection. Biomed Signal Process Control. 2023;82:104561.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data






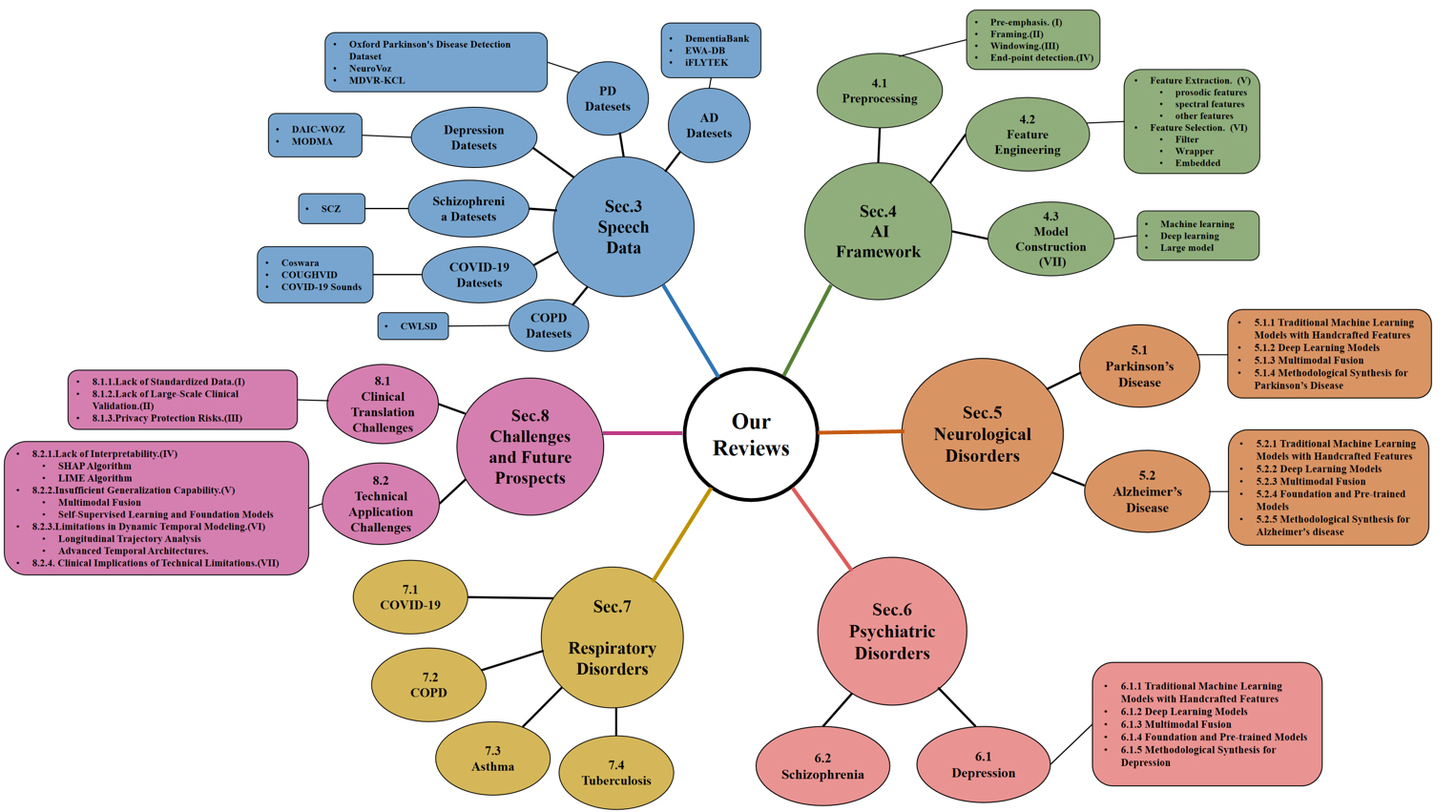
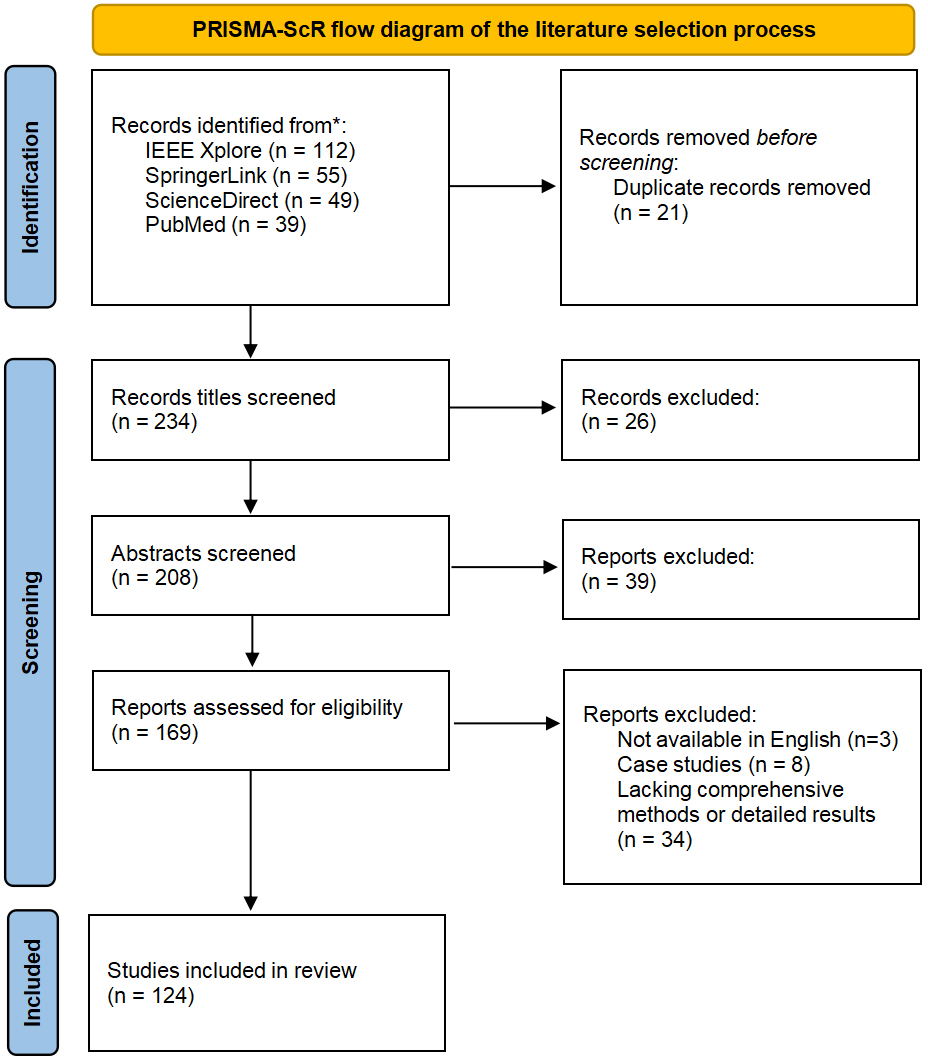
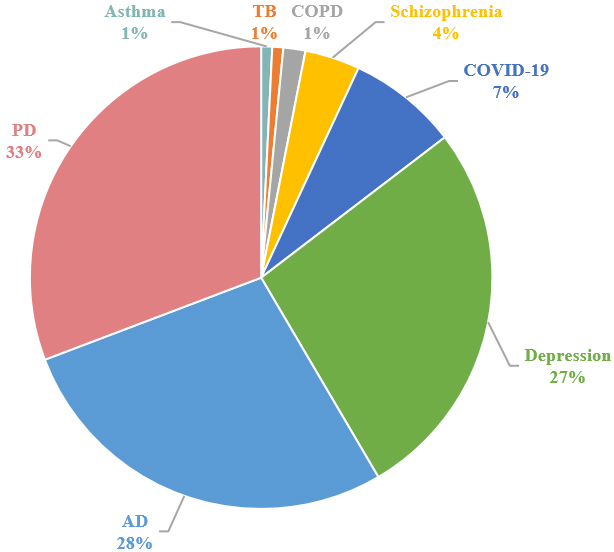
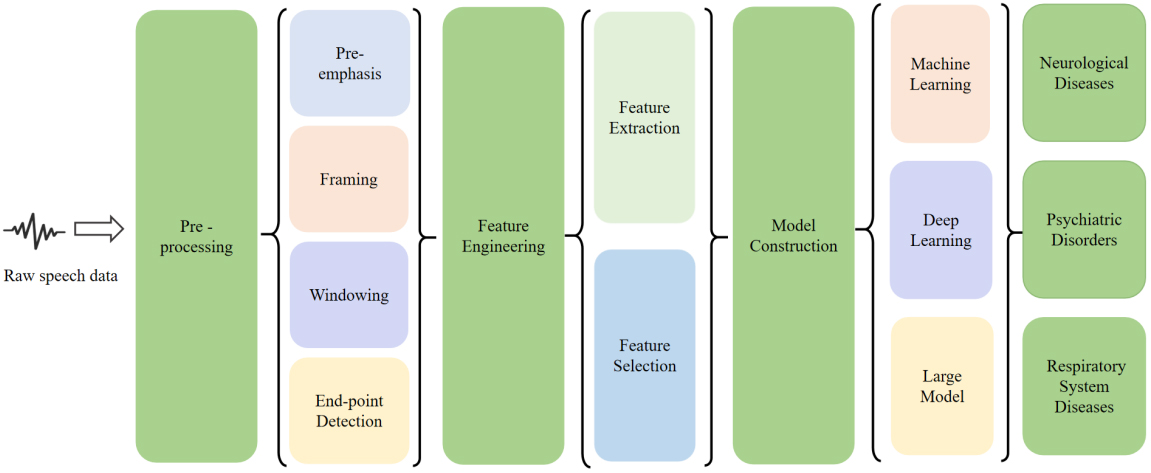
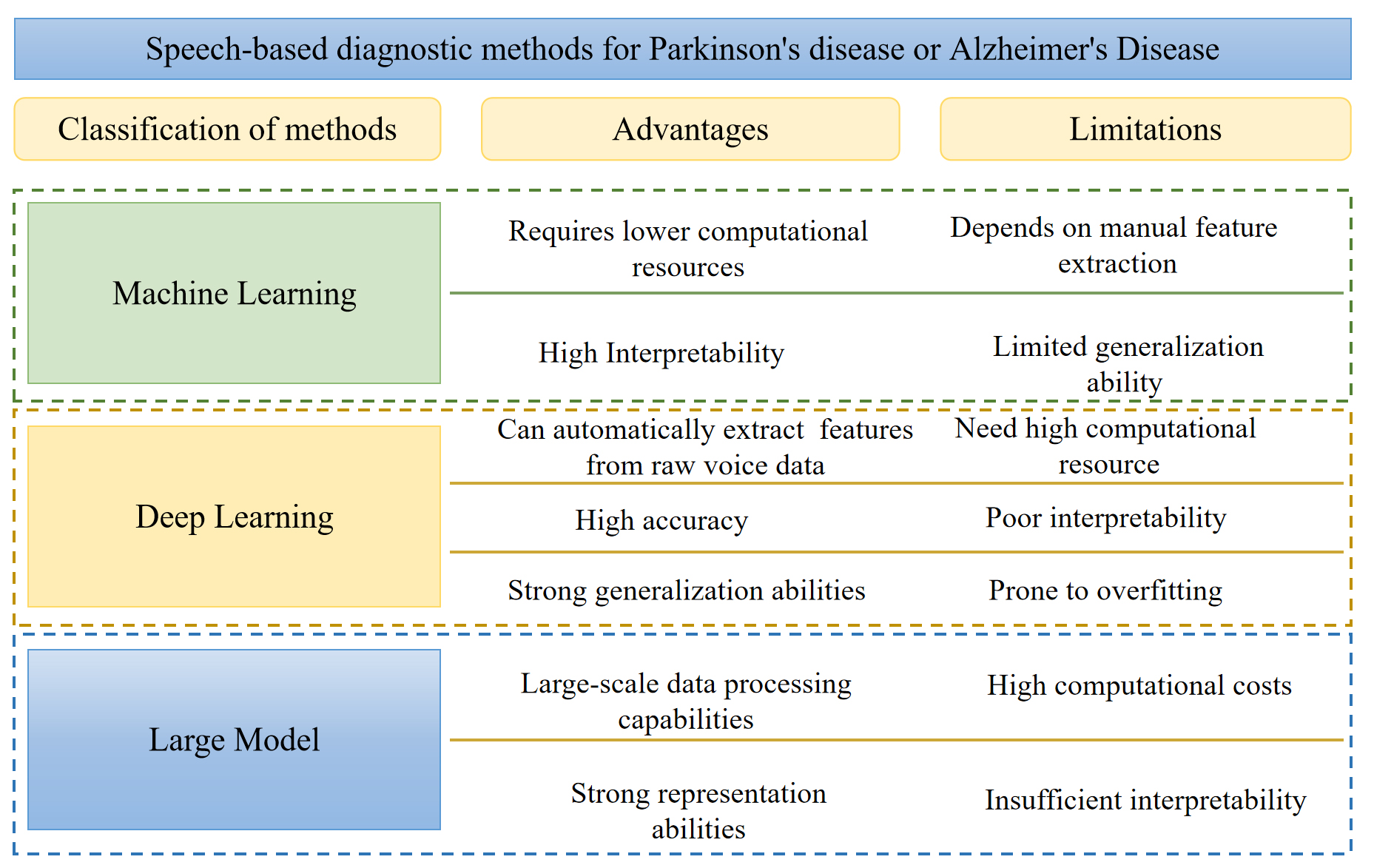
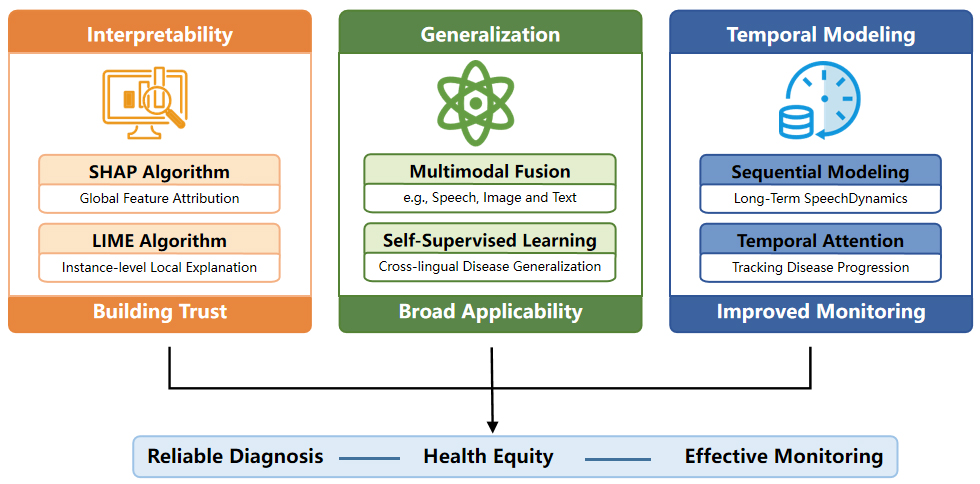











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].