Genetic factors associated with dyslipidemia, T2DM and DKD: a cross-sectional study on FXR gene polymorphisms
Abstract
Aim: Genetic factors play a critical role in the development and progression of type 2 diabetes (T2DM) and diabetic kidney disease (DKD). Recent studies have reported associations between single nucleotide polymorphisms (SNPs) in the farnesoid X receptor (FXR) gene and the risk of T2DM and DKD. Dyslipidemia is increasingly recognized as a risk factor for DKD in patients with T2DM. This study aimed to investigate the potential genetic contribution of FXR gene SNPs rs11110390 and rs4764980 to susceptibility to T2DM and DKD in a Chinese population.
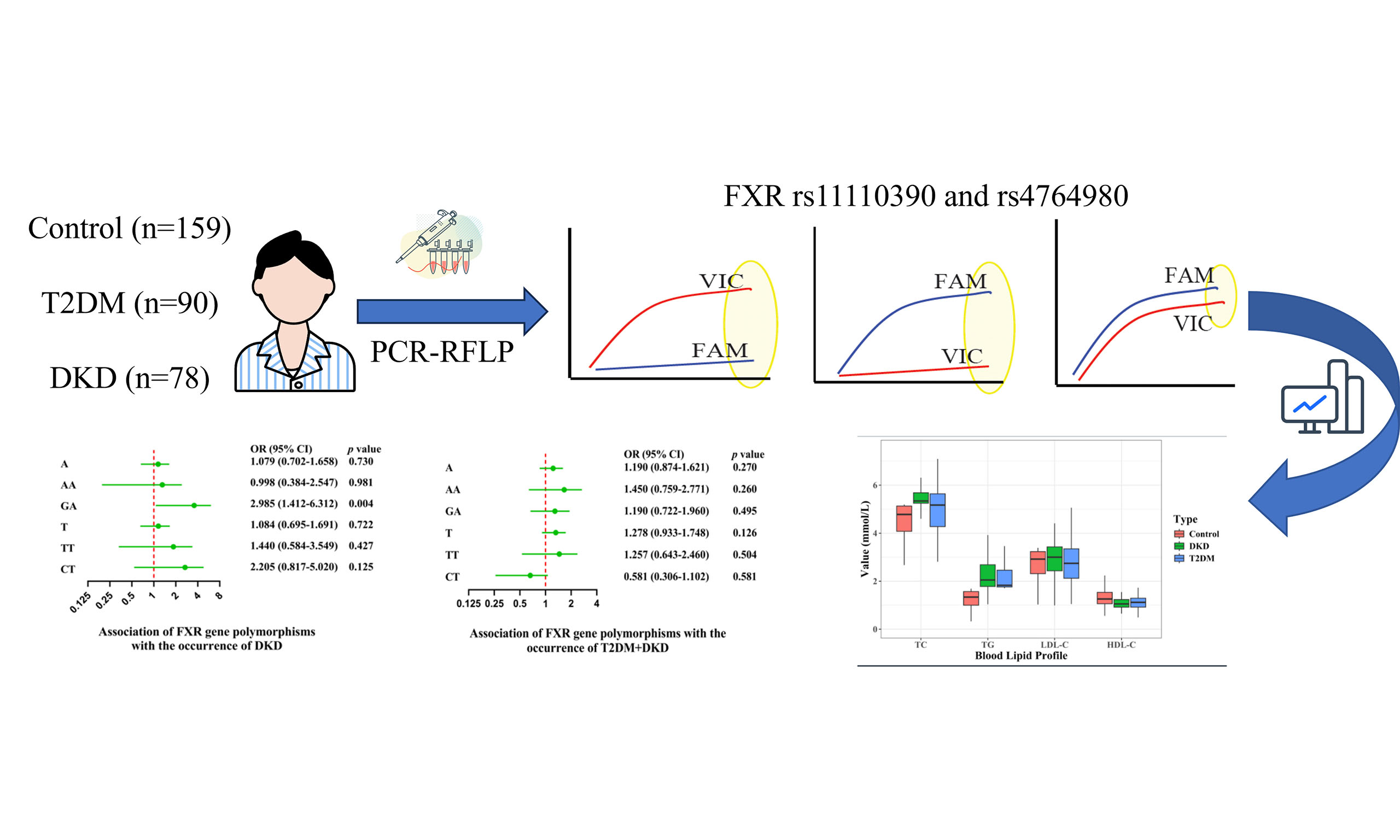
Methods: This cross-sectional study included 168 patients with dyslipidemia and diabetes, comprising 90 T2DM patients and 78 DKD patients, along with 159 healthy controls. Genotyping of FXR SNPs rs11110390 and rs4764980 was performed using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis. Associations between FXR polymorphisms and disease risk were evaluated using odds ratios (ORs) and 95% confidence intervals (CIs).
Results: Regression analysis revealed that neither the genotypes nor the alleles of FXR rs11110390 and rs4764980 were significantly associated with T2DM risk. In contrast, the rs4764980 GA genotype was significantly associated with an increased risk of DKD (OR = 2.985; 95%CI = 1.412-6.312; P = 0.004).
Conclusion: The FXR rs4764980 GA genotype may confer increased susceptibility to DKD in the Chinese population. However, no significant associations were observed between rs11110390 or rs4764980 and T2DM risk or overall DKD risk.
Keywords
INTRODUCTION
Diabetes has become a major chronic metabolic disease worldwide, with over 90% of cases attributed to type 2 diabetes mellitus (T2DM)[1]. T2DM is a complex metabolic disorder with high genetic susceptibility and represents a significant threat to global health and longevity. Studies have shown that having one parent with diabetes increases an individual's risk of developing T2DM by 30%-40%, whereas having both parents affected raises the risk by 70%[2]. Among the complications of diabetes, diabetic kidney disease (DKD) is clinically significant, being one of the most frequent microvascular complications and a leading contributor to end-stage renal disease (ESRD)[3,4]. Current treatments for DKD primarily focus on correcting metabolic imbalances and hemodynamic abnormalities[5]. Notably, dyslipidemia has been widely recognized as a risk factor for diabetic nephropathy[6]. Therefore, identifying new therapeutic targets for the prevention and treatment of dyslipidemia in T2DM and DKD (hereafter abbreviated as T2DM, DKD) remains a major challenge.
As global mortality from kidney disease continues to rise, accumulating evidence suggests that ligands for nuclear receptors (NRs), a superfamily of ligand-gated transcription factors, could provide protection against renal ischemia/reperfusion (I/R) injury, drug-induced acute kidney injury (AKI), DKD, renal fibrosis, and renal cancer[7]. Farnesoid X receptor (FXR), a member of the nuclear receptor superfamily, is activated by bile acids and plays a crucial role in both physiological and pathological processes. FXR regulates multiple functions, including bile acid metabolism, glycolipid metabolism, and inflammatory responses. Bile acids themselves are key signaling molecules that interact with several nuclear receptors, such as FXR, the vitamin D receptor, pregnane X receptor, and G-protein-coupled receptors, and influence cellular signaling pathways including c-Jun N-terminal kinases (JNKs) and extracellular signal-regulated kinases (ERKs) to modulate metabolic processes[8]. Experimental studies support the therapeutic potential of FXR activation. For example, FXR agonists prevent high-fat diet-induced upregulation of renal SREBP-1 in C57BL/6J mice and reduce proteinuria and glomerulosclerosis in db/db mice, thereby improving renal function[9]. Our previous work demonstrated that FXR gene knockout exacerbates renal injury in the streptozotocin (STZ)-induced diabetic nephropathy model, whereas FXR agonist treatment reduces proteinuria, suppresses renal fibrosis, and protects podocytes in db/db mice by decreasing the expression of proinflammatory and pro-fibrotic factors[10]. These findings highlight the potential of FXR agonists as a therapeutic strategy for DKD.
Genetic factors also play a critical role in susceptibility to obesity and T2DM. Single nucleotide polymorphisms (SNPs) have been consistently associated with increased T2DM risk[11,12]. Among the identified susceptibility genes, TCF7L2 variants rs7903146 and rs12255372 show strong associations with T2DM across multiple populations. These risk alleles may impair pancreatic β-cell function by modulating Wnt signaling and glucagon-like peptide-1 (GLP-1) secretion[13-17]. The PPAR-γ gene rs1801282 (Pro12Ala) polymorphism influences adipocyte differentiation and insulin sensitivity, with the Ala allele generally conferring protective effects[18]. Similarly, the FTO gene rs9939609 variant, which affects energy metabolism, can increase body mass index (BMI) and indirectly elevate T2DM risk[19]. FXR activation modulates the expression of genes involved in bile acid, lipid, and glucose metabolism and transport, highlighting its clinical potential for treating diabetes and metabolic disorders[20]. In addition, FXR has demonstrated renoprotective effects in diabetes- and obesity-related nephropathy[21]. For example, a 2013 study by
METHODS
This case-control study included 168 patients with DM and 159 healthy control subjects. Participants were consecutively recruited between October 2022 and November 2024 from the outpatient and inpatient departments of Shunyi Hospital, Beijing Hospital of Traditional Chinese Medicine. The study protocol was approved by the Medical Ethics Committee of Shunyi Hospital, Beijing Hospital of Traditional Chinese Medicine (Approval No. 2022SYKY007-01), and all participants provided written informed consent. The study was registered with the China Clinical Trial Registry (ChiCTR2200064703).
In accordance with international guidelines and previous studies, the diagnostic criteria for DM were defined as the use of antidiabetic medication, fasting blood glucose ≥ 7.0 mmol/L, or glycated hemoglobin (HbA1c) ≥ 6.5%[24]. Based on clinical characteristics and sample information, patients aged 18-75 years with DM were screened for DKD. For this study, diagnostic criteria for DKD were applied independently, and participants were classified as having DKD if they met any one of the following conditions: estimated glomerular filtration rate (eGFR) < 90 mL/min, urinary microalbumin > 30 mg/L, or blood urea nitrogen (BUN) > 7.14 mmol/L. This approach was chosen to identify both early and advanced stages of DKD, since some patients may present with reduced eGFR in the absence of overt proteinuria, while others may exhibit elevated urinary microalbumin despite preserved eGFR. All laboratory measurements were performed using standardized clinical procedures.
Dyslipidemia was defined according to established criteria[25]:
• Hypercholesterolemia: elevated total cholesterol (TC ≥ 5.2 mmol/L, triglycerides (TG) < 1.7 mmol/L);
• Hypertriglyceridemia: elevated TG (TG ≥ 1. 7 mmol/L, TC < 5.2 mmol/L);
• Mixed hyperlipidemia: elevated TC and TG (TC ≥ 5.2 mmol/L, TG ≥ 1.7 mmol/L);
• Hypo-HDL cholesterolemia: low HDL-C (HDL-C < 1.0 mmol/L).
Accordingly, dyslipidemia was defined in this study as TC ≥ 5.2 mmol/L and/or TG ≥ 1.7 mmol/L and/or HDL-C < 1.0 mmol/L, and was used as an inclusion criterion for patients with T2DM and DKD (shown in Figure 1).
The diabetes cohort was further classified into two subgroups: the T2DM group (without nephropathy) and the DKD group, based on annual urine microalbumin test results obtained from medical records.
The control group comprised 159 age- and sex-matched healthy individuals recruited from the same hospital. Individuals with obesity, hypertension, or known risk factors for diabetes were excluded. Additional exclusion criteria were congenital or acquired renal disease, recent infection, or current/past use of nephrotoxic medications.
Sample size estimation
The sample size was calculated based on the difference in minor allele frequency between the case and control groups. A total of 100 participants per group (n = 100) was required to achieve 85% power at a 5% significance level, as determined using G*Power online sample size estimation software.
Clinical evaluation
All participants underwent a comprehensive medical history review and physical examination. Data on sex, age, HbA1c, serum creatinine, BUN, and urinary microalbumin from the previous year were collected from medical records. Height and weight were measured following standardized procedures, and BMI was calculated as weight (kg) divided by height squared (m2). BMI categories were defined using the WHO 2007 criteria: underweight (BMI < 20), normal weight (BMI 20-24.9), overweight (BMI 25-29.9), and obese (BMI ≥ 30)[26].
Systolic and diastolic blood pressure (SBP and DBP) were measured using a standardized method. Three readings were obtained with participants in a seated position using a conventional sphygmomanometer. Hypertension was defined as SBP ≥ 140 mmHg and/or DBP ≥ 90 mmHg, based on the average of the second and third readings[27].
Biochemical evaluation
The average serum creatinine and BUN levels over the past year were extracted from patients' medical records. eGFR was calculated using the 2021 Chronic Kidney Disease Epidemiology Collaboration (CKD-EPI) equation, as recommended by the American Society of Nephrology and the National Kidney Foundation, to define, classify, screen, and monitor CKD. The CKD-EPI formula, based on serum creatinine, is applicable to most adult patients with CKD[28,29].
FXR gene polymorphism analysis
Sample collection and DNA extraction
Two milliliters of venous blood were collected into EDTA tubes and stored at -20 °C until DNA extraction. Genomic DNA was extracted using the QIAamp DNA Mini Kit (QIAGEN, Germany).
PCR amplification and genotyping
PCR-RFLP analysis was used to detect polymorphisms in the promoter region of the FXR gene. Genotyping of SNPs rs11110390 and rs4764980 was performed using a PCR kit (Thermo Fisher Scientific, USA). To confirm accuracy, randomly selected PCR products were sequenced by TsingKe Biological Technology (Beijing, China). PCR reactions were performed in a final volume of 25 μL containing 2 μL of extracted DNA, 1 μL each of forward and reverse primers, 13.5 μL of deionized water, and 8.5 μL of master mix.
The primer sequences were as follows:
1. SNP rs11110390: Forward 5′-AGACCAGCTCAGGTCCAAGGGCTAG-3′; Reverse 5′-GAAACAGACTTTGTGCATTGATGAG-3′.
2. SNP rs4764980: Forward 5′-TCTGCTGCCCAGCCCTCTTGGGTTC-3′; Reverse 5′-GAGTTCCACTGTGATATATGGCATA-3′.
Primer sequences were validated using the NCBI Primer-BLAST tool. The PCR amplification protocol included an initial denaturation at 95 °C for 10 min, followed by 40 cycles of 92 °C for 15 s and 60 °C for
Statistical analysis
Data analysis was conducted using IBM SPSS Statistics for Windows, version 27.0. Categorical variables were summarized as frequencies and percentages, and comparisons were performed using the chi-squared test. Continuous variables with skewed distributions were expressed as medians (minimum-maximum values). Normality was tested using the Kolmogorov-Smirnov test. Normally distributed continuous variables were expressed as mean ± standard deviation (SD). Comparisons between two groups of normally distributed continuous variables were conducted using the t-test, while comparisons among more than two groups were performed using one-way analysis of variance (ANOVA), followed by Bonferroni post hoc tests for pairwise comparisons. For non-normally distributed quantitative variables, the Mann-Whitney test was used for two-group comparisons, and the Kruskal-Wallis test for more than two groups, with pairwise comparisons performed when appropriate. Logistic regression analysis was used to assess the association between FXR polymorphisms and the risk of T2DM and DKD. Hardy-Weinberg equilibrium was tested using the chi-squared test. Genotype and allele frequencies of SNPs were also compared using the chi-squared test.
RESULTS
Two groups were included in this cross-sectional case-control study: the DM group (n = 168; male: 63.1%) and the control group (n = 159; male: 65.4%). The DM group was further divided into two subgroups based on annual urinary microalbumin levels: T2DM without diabetic nephropathy (T2DM; n = 90; male: 56.7%) and T2DM with diabetic nephropathy (DKD; n = 78; male: 70.5%). The clinical characteristics of the study participants are shown in Table 1, with data sources provided in the Supplementary Materials.
Clinical and biochemical characteristics of the study population
| Variables | Total (n = 327) | Control (n = 159) | T2DM (n = 90) | DKD (n = 78) | H/χ2 | P |
| Age (years) | 53.00 (44.00, 61.00) | 49.00 (38.00, 56.00) | 59.00 (50.00, 64.25)a | 58.00 (49.50, 63.00)a | H: 50.140 | < 0.01c |
| BMI (kg/m2) | 23.20 (22.40, 24.80) | 22.80 (22.10, 23.20) | 25.10 (23.00, 27.62)a | 24.50 (23.10, 25.55)a | H: 84.613 | < 0.01c |
| SBP (mmHg) | 131.00 (126.00, 136.00) | 129.00 (125.00, 133.00) | 132.50 (125.00, 136.00)a | 135.50 (129.00, 137.00)a | H: 28.898 | < 0.01c |
| DBP (mmHg) | 80.00 (74.00, 84.00) | 78.00 (73.00, 82.00) | 82.50 (76.00, 85.00)a | 83.00 (78.00, 86.00)a | H: 31.218 | < 0.01c |
| HbA1c (%) | 7.90 (6.90, 9.70) | - | 7.50 (6.67, 9.20) | 8.40 (7.30, 10.10) | H: 7.484 | < 0.01c |
| Egfr (mL/min/1.73m2) | 98.50 (89.15, 107.60) | 98.00 (87.50, 106.10) | 98.45 (91.67, 106.72) | 100.10 (88.47, 109.62) | H: 0.891 | 0.640 |
| ALB (mg/L) | 30.81 (15.50, 66.17) | - | 16.85 (10.90, 23.50) | 66.80 (36.90, 151.00) | H: 111.840 | < 0.01c |
| BUN (mmol/L) | 5.20 (4.30, 6.45) | 4.72 (4.10, 5.70) | 5.56 (4.44, 6.97)a | 5.87 (5.14, 7.39)a | H: 38.985 | < 0.01c |
| Gender, n (%) | ||||||
| Male | 210 (64.2%) | 104 (65.4%) | 51 (56.7%) | 55 (70.5%) | χ2: 3.677 | 0.159 |
| Female | 117 (35.8%) | 55 (34.6%) | 39 (43.3%) | 23 (29.5%) | ||
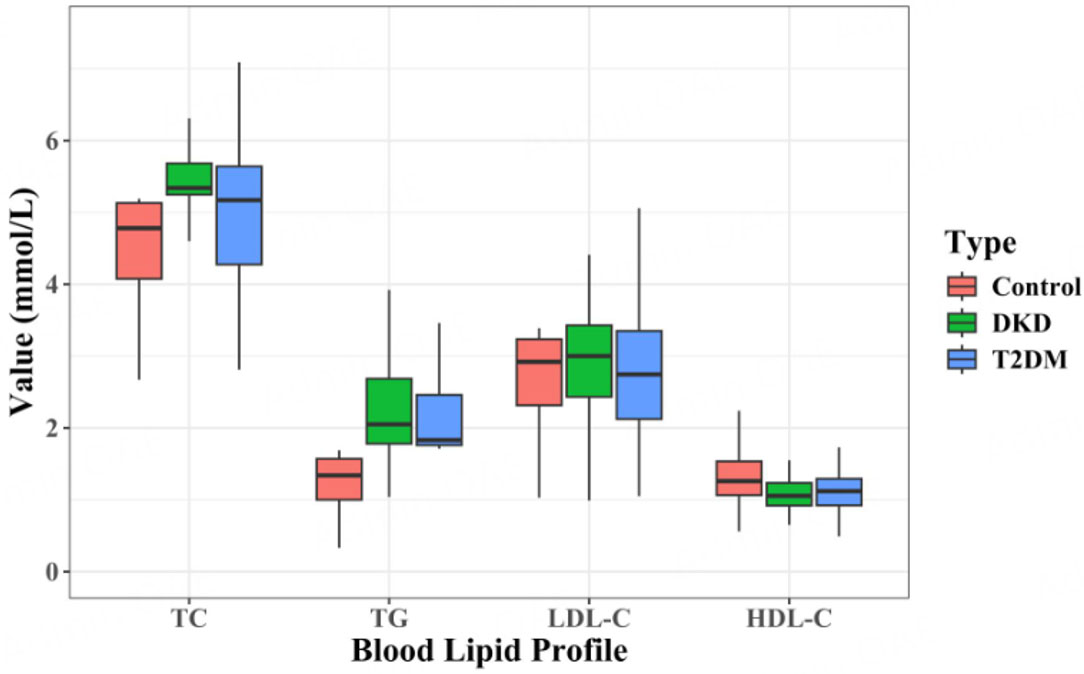
Lipid data from the T2DM, DKD, and healthy groups were expressed as mean ± SD, and group characteristics are presented in Table 2 (data sources Lipid data in the Supplementary Materials).
Lipid profile across study groups
| Variables | Total (n = 327) | Control (n = 159) | T2DM (n = 90) | DKD (n = 78) | P |
| TC (mmol/L) | 4.98 ± 1.06 | 4.56 ± 0.68 | 5.17 ± 1.23 | 5.77 ± 1.09 | < 0.01c |
| TG (mmol/L) | 1.89 ± 1.56 | 1.24 ± 0.35 | 2.49 ± 2.01 | 2.77 ± 2.01 | < 0.01c |
| LDL-C (mmol/L) | 2.84 ± 0.78 | 2.74 ± 0.59 | 2.90 ± 0.95 | 2.99 ± 0.94 | 0.177 |
| HDL-C (mmol/L) | 1.31 ± 0.36 | 1.42 ± 0.37 | 1.22 ± 0.34 | 1.16 ± 0.27 | < 0.01c |
All three groups were comparable in terms of age, BMI, SBP, DBP, ALB, and BUN (all P < 0.05). SBP and DBP values were within the normal range for all participants. eGFR was measured using standardized serum creatinine assays and calculated with the CKD-EPI creatinine formula. After extensive validation, no statistically significant differences in eGFR were found among the three groups (P = 0.640).
Hardy-weinberg equilibrium (HWE) testing
In the control group, the genotype distribution of FXR gene SNPs rs11110390 and rs4764980 conformed to Hardy-Weinberg equilibrium (P > 0.05), with P values of 0.936 and 0.634, indicating representativeness of the control population.
FXR gene SNP genotype and allele distributions
Detailed genotypic and allelic distributions of rs11110390 and rs4764980 in the FXR gene are shown in Table 3, with data sources available in the Supplementary Materials.
Genotype and allele distribution of FXR gene polymorphismsrs11110390 (C>T) and rs4764980 (A>G) in study groups
| Variables | Control (n = 159) | T2DM (n = 90) | DKD (n = 78) | P |
| rs11110390 (C>T) | ||||
| CC | 22 (13.8) | 18 (20) | 10 (12.8) | < 0.01c |
| CT | 92 (57.9) | 32 (35.6) | 36 (46.2) | < 0.01c |
| TT | 45 (28.3) | 40 (44.4) | 32 (41.0) | < 0.01c |
| C allele | 136 (42.8) | 68 (37.8) | 56 (35.9) | - |
| T allele | 182 (57.2) | 112 (62.2) | 100 (64.1) | - |
| rs4764980 (A>G) | ||||
| GG | 51 (32.1) | 31 (34.4) | 15 (19.2) | < 0.01c |
| GA | 82 (51.6) | 36 (40) | 52 (66.7) | < 0.01c |
| AA | 26 (16.3) | 23 (25.6) | 11 (14.1) | < 0.01c |
| G allele | 184 (57.9) | 98 (54.4) | 82 (52.6) | - |
| A allele | 134 (42.1) | 82 (45.6) | 74 (47.4) | - |
For rs11110390, genotype frequencies were generally higher in the healthy group compared to the T2DM and DKD groups. The CC genotype frequency was similar between the healthy and T2DM groups but was markedly lower in the DKD group. A similar pattern was observed for the TT genotype. For the CT genotype, frequencies were higher in the healthy group compared to both the T2DM and DKD groups, while the latter two showed no substantial difference. In addition, the frequency of the C allele was lower than that of the T allele in all groups. However, the T allele frequency was higher in the healthy group than in the T2DM and DKD groups.
For rs4764980, genotype frequencies were also higher in the healthy group compared to the T2DM and DKD groups. The GG genotype was more frequent in the healthy group, whereas the DKD group showed a significantly lower frequency. The GA genotype was less common in the DKD group compared with the healthy and T2DM groups. Similarly, the AA genotype was markedly less frequent in the DKD group compared to the other two groups. Across all study groups, the G allele was more common than the A allele. However, both G and A allele frequencies were lower in the DKD group compared to the T2DM and healthy groups.
Association between FXR gene SNPs and T2DM + DKD risk
Logistic regression analysis was used to evaluate the association between FXR gene SNPs and the combined risk of T2DM and DKD (Table 4; data sources available in the Supplementary Materials).
Correlation analysis of FXR gene 390 (C>T) and 980 (A>G) polymorphisms with T2DM and DKD in each study group
| Variables | Total T2DM + DKD (n = 168) | Control (n = 159) | T2DM (n = 90) | DKD (n = 78) | Total T2DM + DKD vs. Control | DKD vs. T2DM |
| FXR Gene 390 n (%) | ||||||
| CC | 28 (16.67) | 22 (13.84) | 18 (20.00) | 10 (12.82) | Ref. | Ref. |
| CT | 68 (40.48) | 92 (57.86) | 32 (35.56) | 36 (46.15) | X2 = 2.798 P = 0.096 OR = 0.581 (0.306-1.102) | X2 = 2.358 P = 0.125 OR = 2.205 (0.817-5.020) |
| TT | 72 (42.86) | 45 (28.30) | 40 (44.44) | 32 (41.03) | X2 = 0.447 P = 0.504 OR = 1.257 (0.643-2.460) | X2 = 0.631 P = 0.427 OR = 1.440 (0.584-3.549) |
| C allele | 124 (36.90) | 136 (42.77) | 68 (37.78) | 56 (35.90) | Ref. | Ref. |
| T allele | 212 (63.10) | 182 (57.23) | 112 (62.22) | 100 (64.10) | X2 = 2.344 P = 0.126 OR = 1.278 (0.933-1.748) | X2 = 0.127 P = 0.722 OR = 1.084 (0.695-1.691) |
| FXR Gene 980 n (%) | ||||||
| GG | 46 (27.38) | 51 (32.08) | 31 (34.44) | 15 (19.23) | Ref. | Ref. |
| GA | 88 (52.38) | 82 (51.57) | 36 (40.00) | 52 (66.67) | X2 = 0.466 P = 0.495 OR = 1.190 (0.722-1.960) | X2 = 8.474 P = 0.004c OR = 2.985 (1.412-6.312) |
| AA | 34 (20.24) | 26 (16.35) | 23 (25.56) | 11 (14.10) | X2 = 1.268 P = 0.260 OR = 1.450 (0.759-2.771) | X2 = 0.001 P = 0.981 OR = 0.998 (0.384-2.547) |
| G allele | 180 (53.57) | 184 (57.86) | 98 (54.44) | 82 (52.56) | Ref. | Ref. |
| A allele | 156 (46.43) | 134 (42.14) | 82 (45.56) | 74 (47.44) | X2 = 1.218 P = 0.270 OR = 1.190 (0.874-1.621) | X2 = 0.119 P = 0.730 OR = 1.079 (0.702-1.658) |
For rs11110390:
• CT genotype: OR = 0.581; 95%CI = 0.306-1.102; P = 0.096.
• TT genotype: OR = 1.257; 95%CI = 0.643-2.460; P = 0.504.
• T allele: OR = 1.278; 95%CI = 0.933-1.748; P = 0.126.
For rs4764980:
• GA genotype: OR = 1.190; 95%CI = 0.722-1.960; P = 0.495.
• AA genotype: OR = 1.450; 95%CI = 0.759-2.771; P = 0.260.
• A allele: OR = 1.190 (vs. G allele); 95%CI = 0.874-1.621; P = 0.270.
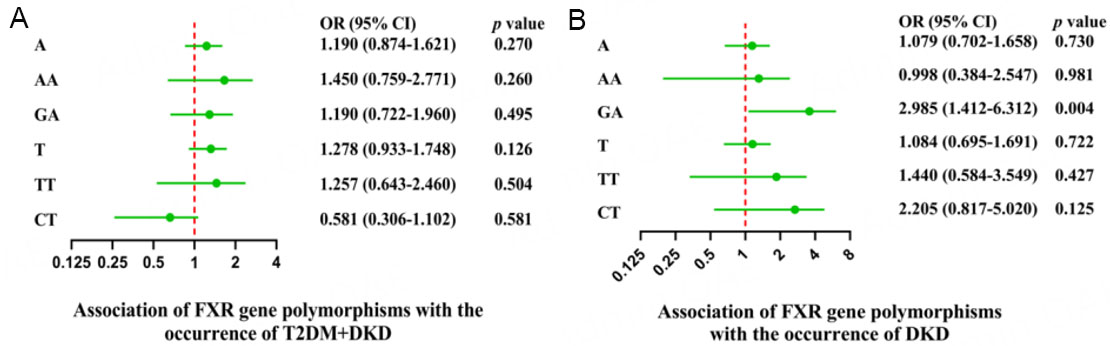
These results suggest that neither the genotypes nor alleles of rs11110390 and rs4764980 are significantly linked to increased risk of T2DM [Figure 2A].
Association between FXR gene SNPs and DKD risk
Logistic regression analysis was also performed to evaluate the association between FXR SNPs and the risk of DKD (Table 4; data sources available in the Supplementary Materials).
Compared with patients with T2DM, regression analysis of rs11110390 revealed no significant association of either genotype or allele with DKD susceptibility. However, for rs4764980, the GA genotype was significantly associated with an increased risk of DKD (OR = 2.985; 95%CI = 1.412-6.312; P = 0.004).
These results suggest a potential role of the GA genotype of rs4764980 in increasing the risk of DKD [Figure 2B].
DISCUSSION
DKD is a leading cause of ESRD, affecting more than 40% of patients with T2DM. The natural course of DKD is characterized by a prolonged clinically silent phase, during which substantial renal damage may already be occurring. Among the many influencing factors, genetic predisposition plays a critical role in susceptibility to T2DM and DKD. From a molecular genetics perspective, specific gene variants and combinations can increase an individual's risk of developing these conditions. Such genetic influences may affect not only disease onset but also its progression, severity, and therapeutic response. Dyslipidemia is a key factor in the progression from T2DM to DKD, and most patients with T2DM and DKD present with abnormal lipid profiles. FXR, a bile acid receptor highly expressed in renal proximal tubules, regulates glucose and lipid metabolism as well as inflammatory pathways. Previous studies have shown that rs11110390 and rs4764980 are significantly associated with FFA levels and fasting plasma glucose, respectively, suggesting their potential role in metabolic regulation and, by extension, DKD susceptibility. Based on this rationale, our study investigated whether FXR variants contribute to the risk of T2DM and DKD in patients with dyslipidemia. Our findings confirmed that patients with T2DM and DKD had significantly elevated lipid levels compared with healthy controls. Although neither FXR rs11110390 nor rs4764980 genotypes or alleles were associated with increased T2DM risk, the GA genotype of FXR rs4764980 was significantly related to DKD. This suggests that FXR rs4764980 may serve as an independent genetic marker for DKD risk. Its pathogenic role may not involve increasing diabetes incidence per se, but rather modulating renal susceptibility to metabolic stress in the diabetic state. Incorporating FXR rs4764980 genotyping into DKD risk prediction models may facilitate early identification and intervention in high-risk individuals, while also providing a rationale for developing FXR-targeted therapies. Until stronger evidence is available, a cautious clinical approach is warranted for GA carriers. This includes prioritizing guideline-based renoprotective therapies (e.g., SGLT2 inhibitors, RAAS blockade) and closer monitoring of renal function and urinary albumin for early signs of progression. Regarding targeted interventions, current evidence does not support routine preventive therapy. Instead, GA carriers may be appropriate candidates for enriched or stratified cohorts in early-phase randomized trials designed to evaluate the safety and renoprotective potential of novel pharmacologic interventions. Collectively, our findings support the proposed pathway of “FXR genetic variation - metabolic phenotype - tubular susceptibility - DKD risk”. However, validation in larger cohorts, supplemented with tissue-specific eQTL and colocalization analyses, remains necessary.
Although FXR rs11110390 and rs4764980 have not reached genome-wide significance in existing GWAS for T2DM or DKD, our results indicate that they may nonetheless contribute to disease susceptibility in specific populations or under certain metabolic conditions. In our study, FXR rs11110390 showed no significant association with either T2DM or DKD risk, whereas FXR rs4764980 GA was clearly associated with DKD development. This association may represent a modest genetic effect or one confined to subpopulations. Replication in cohorts of diverse ethnic origins will therefore be essential for validation and generalization.
It is also important to acknowledge the limitations of our study. First, sample size significantly influences the precision and reliability of genetic association studies. Insufficient sample size may obscure true genetic effects and limit generalizability. Second, the sampling technique directly affects standardization, accuracy, and representativeness, thereby impacting the robustness of subsequent analyses. Therefore, future research should adopt larger and more rigorously sampled cohorts. Third, our study lacked information on potentially important clinical covariates, including diabetes duration, glycemic control (e.g., HbA1c), and medication use (e.g., statins, RAAS inhibitors). Because the original design focused primarily on genetic polymorphisms and basic clinical characteristics, these variables were not systematically collected. The absence of such adjustments may have introduced residual confounding, and thus, the associations observed should be interpreted with caution. Nonetheless, the robustness of our findings is supported by the relatively large effect size, plausible biological mechanism, and consistency with previous reports. Future studies should aim to include larger sample sizes, standardized data collection, broader covariate adjustment, and diverse populations to confirm and extend these observations, thereby clarifying the contribution of FXR variants to T2DM and DKD susceptibility.
DECLARATIONS
Acknowledgments
The authors thank the patients who participated in this study and the nephrology outpatient clinic of Shunyi Hospital, Beijing Hospital of Traditional Chinese Medicine, for their assistance in patient recruitment and support during the study period.
Authors' contributions
Made substantial contributions to this work and supported the publication of the manuscript: Lan X, Wu C, Shen Z, Zhang R, Yin C, Qi J, Wang L, Liu P, Li P
Conceptualization: Wang L, Liu P, Li P
Writing - original draft preparation: Lan X, Wu C, Shen Z, Zhang R, Yin C, Qi J
Editing and revision, supervision: Liu P, Li P
Contributed equally to this work: Lan X, Wu C, Shen Z
Availability of data and materials
All data relevant to this study are included in the article or provided as Supplementary Materials.
Financial support and sponsorship
This study was supported by the Capital’s Funds for Health Improvement and Research (No. 2022-4-7104), the National Natural Science Foundation of China (Nos. 82274489, 82174296, U23A20504), the Fifth Batch of National Training Program for Excellent Clinical Talents of Traditional Chinese Medicine [No.1 [2022] of People’s Education of Traditional Chinese Medicine], and the Beijing Hospital Research Incubation Program (PZ2023025).
Conflicts of interest
All authors declared that there are no conflicts of interest.
Ethics approval and consent to participate
The study protocol was approved by the Medical Ethics Committee of Shunyi Hospital, Beijing Hospital of Traditional Chinese Medicine (Approval No. 2022SYKY007-01). All participants provided written informed consent.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2025.
Supplementary Materials
REFERENCES
1. Zeng Q, Zou D, Zeng Q, Chen X, Wei Y, Guo R. Association between insulin-like growth factor-1 rs35767 polymorphism and type 2 diabetes mellitus susceptibility: a meta-analysis. Front Genet. 2021;12:774489.
2. Kwak SH, Park KS. Recent progress in genetic and epigenetic research on type 2 diabetes. Exp Mol Med. 2016;48:e220.
3. Wang J, Zhang R, Wu C, Wang L, Liu P, Li P. Exploring potential targets for natural product therapy of DN: the role of SUMOylation. Front Pharmacol. 2024;15:1432724.
4. Zhou Y, Wu Q, Wang X, Li W, Liu Q, Gao K. Insights into the functional mechanism of diabetic kidney disease treatment with sinensetin based on network pharmacology and molecular docking. Integr Med Nephrol Androl. 2023;10:e00033.
5. Zhou S, Su L, Xu R, et al. Statin initiation and risk of incident kidney disease in patients with diabetes. Can Med Assoc J. 2023;195:E729-38.
6. Deng Y, Zhu H, Xing J, et al. The role of natural products in improving lipid metabolism disorder-induced mitochondrial dysfunction of diabetic kidney disease. Front Physiol. 2025;16:1624077.
7. Luan ZL, Zhang C, Ming WH, Huang YZ, Guan YF, Zhang XY. Nuclear receptors in renal health and disease. EBioMedicine. 2022;76:103855.
9. Jiang T, Wang XX, Scherzer P, et al. Farnesoid X receptor modulates renal lipid metabolism, fibrosis, and diabetic nephropathy. Diabetes. 2007;56:2485-93.
10. Liu P, Shen Z, Wang C, Wang J, Qiu X, Li P. Tangshen formula ameliorates renal fibrosis of diabetic kidney disease through regulating the FXR/vimentin/α-SMA pathway. J Beijing University Traditional Chinese Med. 2022;45:1196-204.
11. Ma L, Yan M, Kong X, et al. Association of EPHX2 R287Q polymorphism with diabetic nephropathy in Chinese type 2 diabetic patients. J Diabetes Res. 2018:2786470.
12. Zhao H, Ma L, Yan M, et al. Association between MYH9 and APOL1 gene polymorphisms and the risk of diabetic kidney disease in patients with type 2 diabetes in a Chinese Han population. J Diabetes Res. 2018:5068578.
13. Grant SF, Thorleifsson G, Reynisdottir I, et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet. 2006;38:320-3.
14. Imamura M, Maeda S. Genetics of type 2 diabetes: the GWAS era and future perspectives [Review]. Endocr J. 2011;58:723-39.
15. Lyssenko V, Lupi R, Marchetti P, et al. Mechanisms by which common variants in the TCF7L2 gene increase risk of type 2 diabetes. J Clin Invest. 2007;117:2155-63.
16. Mahajan A, Wessel J, Willems SM, et al. Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat Genet. 2018;50:559-71.
17. Zeggini E, Scott LJ, Saxena R, et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat Genet. 2008;40:638-45.
18. Altshuler D, Hirschhorn JN, Klannemark M, et al. The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet. 2000;26:76-80.
19. Frayling TM, Timpson NJ, Weedon MN, et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science. 2007;316:889-94.
20. Hu M, Lui SS, Tam LS, Li EK, Tomlinson B. The farnesoid X receptor -1G>T polymorphism influences the lipid response to rosuvastatin. J Lipid Res. 2012;53:1384-9.
21. Wang XX, Wang D, Luo Y, et al. FXR/TGR5 dual agonist prevents progression of nephropathy in diabetes and obesity. J Am Soc Nephrol. 2018;29:118-37.
22. Heni M, Wagner R, Ketterer C, et al. Genetic variation in NR1H4 encoding the bile acid receptor FXR determines fasting glucose and free fatty acid levels in humans. J Clin Endocrinol Metab. 2013;98:E1224-9.
23. Fang Y, Qin M, Zheng Q, et al. Role of bile acid receptors in the development and function of diabetic nephropathy. Kidney Int Rep. 2024;9:3116-33.
24. Wang J, Liu F, Kong R, Han X. Association between globulin and diabetic nephropathy in Type2 diabetes mellitus patients: a cross-sectional study. Front Endocrinol. 2022;13:890273.
25. Hu Y, Du X. [Blood lipid indicators and different clinical classifications of dyslipidemia and diabetic kidney disease: correlation and predictive value]. Sichuan Da Xue Xue Bao Yi Xue Ban. 2023;54:1013-8.
26. Tralongo AC, Caspani F, Proserpio I, Volpi L, Campiotti L. Body mass index (BMI) influence on Cetuximab-induced antibody-dependent cellular cytotoxicity in advanced colon cancer. Intern Emerg Med. 2023;18:297-303.
27. Beaney T, Kerr GK, Kiru G, et al. May measurement month 2022: results from the global blood pressure screening campaign. BMJ Glob Health. 2024;9:e016557.
28. Delanaye P, Cavalier E, Pottel H, Stehlé T. New and old GFR equations: a European perspective. Clin Kidney J. 2023;16:1375-83.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Copyright
Data & Comments
Data

















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].