ADAM9 in cancer: a comprehensive analysis of mutation, prognostic value, expression and TQFL12-induced inhibition
Abstract
Aim: A disintegrin and metalloproteinase domain 9 (ADAM9) is involved in various human diseases, including cone-rod dystrophy, Alzheimer’s disease, cancer, and viral infections. However, its comprehensive expression profile and therapeutic potential across cancers remain poorly understood.
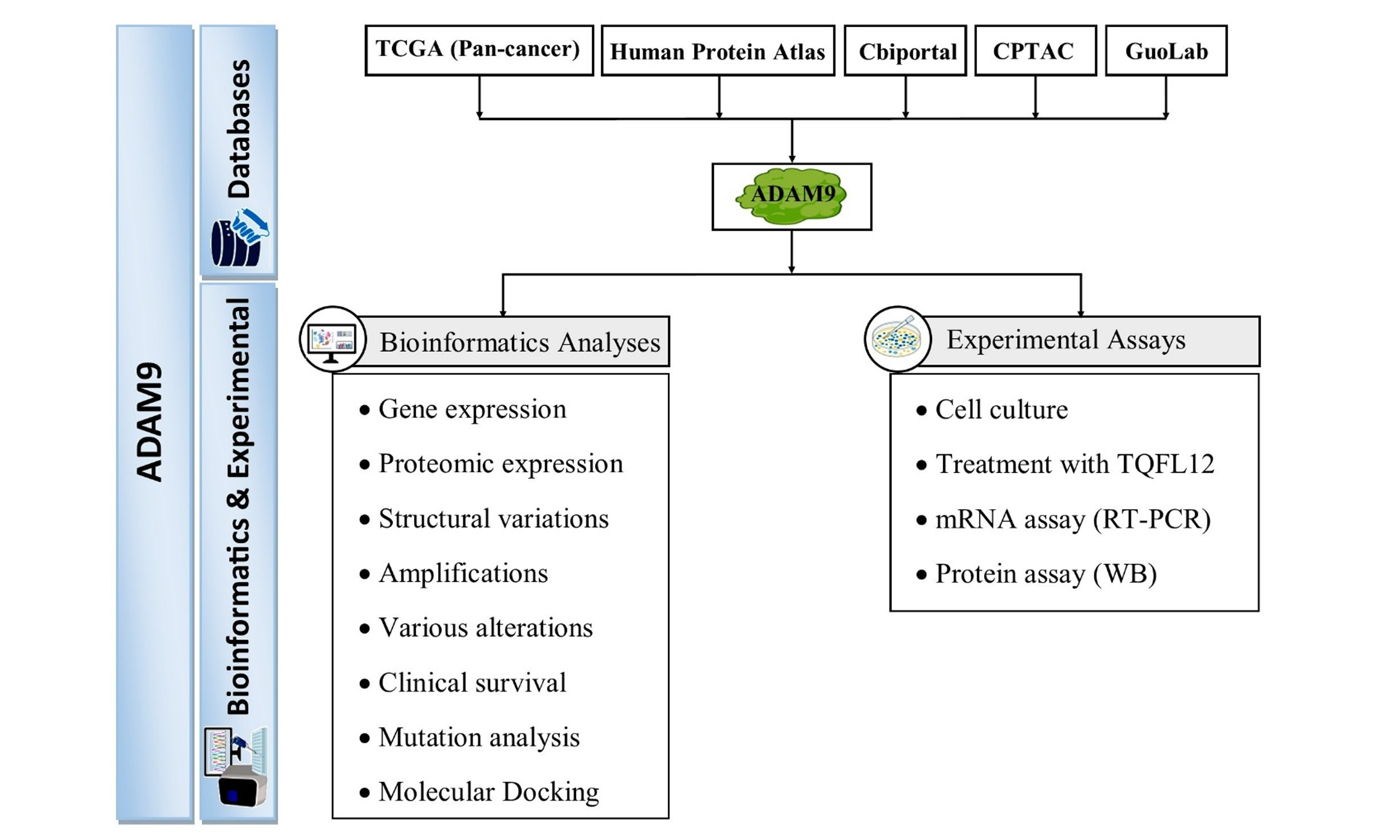
Methods: Pan-cancer analyses of ADAM9 expression, mutation status, and prognostic value were conducted using The Cancer Genome Atlas (TCGA) and cBioPortal datasets. The differences between cancer and normal tissues, as well as survival associations, were examined. Mutation landscapes were characterized, and in vitro assays were performed in prostate, breast, and lung cancer cell lines treated with increasing concentrations of Thymoquinone Derivative FL12 (TQFL12). ADAM9 messenger RNA (mRNA) and protein levels were determined by quantitative polymerase chain reaction and Western blotting, respectively.
Results: ADAM9 expression was elevated in multiple cancers, but decreased in Kidney chromophobe (KICH) and Thyroid carcinoma (THCA). High ADAM9 expression correlated with longer overall survival (OS) in Colon adenocarcinoma (COAD), while predicting poorer OS in Breast Invasive Carcinoma (BRCA), Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma (CESC), KICH, Liver hepatocellular carcinoma (LIHC), Brain Lower Grade Glioma (LGG), Mesothelioma (MESO), Pancreatic Adenocarcinoma (PAAD), and Uveal Melanoma (UVM), suggesting its role as an unfavorable prognostic biomarker in cancers. ADAM9 also exhibited frequent mutations, with mutated cases showing improved progression-free and disease-specific survival, implying favorable prognostic relevance. Notably, TQFL12 - a novel compound synthesized in our laboratory - significantly suppressed ADAM9 protein expression in a dose-dependent manner in 22RV1, MDA-MB-231, and H1975 cells without altering mRNA levels, suggesting that the regulatory effect may occur at the post-translational or translational level.
Conclusions: These results highlight ADAM9 as a potential prognostic marker and therapeutic target, while identifying TQFL12 as a promising inhibitor across multiple cancers.
Keywords
INTRODUCTION
A disintegrin and metalloproteinase domain 9 (ADAM9) (OMIM: 602713), also known as MCMP (myeloma cell metalloproteinase), MDC9 (metalloprotease/disintegrin/cysteine-rich protein 9), CORD9 (cone-rod dystrophy 9), KIAA0021, or Mltng, is cytogenetically located at 8p11.22. It encodes a protein of 819 amino acids with a predicted molecular weight of 90,556 Da (NM_003816.3, NP_003807.1). ADAM9 (A Disintegrin and Metalloproteinase 9) is a family member of the ADAM (a disintegrin and metalloprotease domain) family[1,2]. It has been implicated in several human diseases, including cone-rod dystrophy (CRD), Alzheimer’s disease, and cancer[2]. ADAM9 mutations are associated with CRD, a genetic disorder characterized by progressive visual degeneration[3]. ADAM9 can cleave and release numerous substrates and proteins that play critical roles in tumorigenesis and angiogenesis. Dysregulation of ADAM9 expression is correlated with a variety of cancers, including lung cancer[4], pancreatic cancer[5,6], prostate cancer[7], oral squamous cell carcinoma[8], colorectal cancer[9], esophageal cancer[10], and breast cancer[11], demonstrating its role as a potential target for therapeutics and diagnostics. Moreover, ADAM9 is involved in drug resistance, such as resistance to cisplatin in gastric cancer cells, via the DNA damage response pathway[12]. Interestingly, an antibody targeting ADAM9 was recently reported to inhibit epithelial-mesenchymal transition (EMT) and migration in prostate cancer cell lines and in mice by suppressing ADAM9 expression[13]. A cyclic increase in the Histamine receptor H1-ADAM9-Snail/Slug axis has been reported to promote the progression of EMT[14]. ADAM9 may be involved in numerous cancer-related biological processes, such as cancer cell proliferation, apoptosis, invasion, migration, and drug resistance.
Thymoquinone (TQ) is a small molecule and the active ingredient extracted from seeds of the natural product Nigella sativa, which has been reported to be effective against a wide range of diseases, including inflammatory and immunological diseases, cancer, and others[15,16]. Thymoquinone Derivative FL12 (TQFL12) is a novel synthetic derivative of TQ in our laboratory, which suppresses triple-negative breast cancer (TNBC) metastasis and invasion both in vitro and in vivo, with lower toxicity to normal tissues compared with TQ[17]. TQFL12 has also been shown to inhibit the protein expression of TMPRSS2 (Transmembrane protease, serine 2), a receptor for SARS-CoV-2 (severe acute respiratory syndrome-like coronavirus) and biomarker for prostate cancer, in 22RV1 prostate cancer cells[18]. However, it remains unclear whether ADAM9 is a potential oncotherapeutic target inhibited by TQFL12.
Nevertheless, the regulation of ADAM9 expression by small-molecule medicines may influence tumorigenesis and cancer progression. Therefore, evaluating ADAM9 expression across different cancer tissues and the inhibitory effects of small molecules is critical. In this study, we comprehensively analyzed ADAM9 expression in various tumor tissues and explored a potential therapeutic strategy targeting ADAM9 using the small-molecule TQFL12, which may be useful for the treatment of multiple cancers.
MATERIALS AND METHODS
Databases for bioinformatics
The Human Protein Atlas (HPA) was used to obtain messenger RNA (mRNA) and protein expression data for ADAM9 in normal and tumor tissues[19,20]. The Cancer Genome Atlas (TCGA) plot R package (Ver 8.0.0) was used to analyze ADAM9 expression, overall survival (OS), and domain structures across pan-cancer datasets and matched normal tissues[21]. The Cbioportal for Cancer Genomics website was used to analyze somatic mutations, structural variations, amplifications, various alterations, and clinical survival data of ADAM9 across different malignancies. Mutation analysis was also performed using the GuoLab website (https://guolab.wchscu.cn/GuoLab/). The proteomic expression profile of ADAM9 was analyzed using the Clinical Proteomic Tumor Analysis Consortium (CPTAC) data in the University of Alabama at Birmingham Cancer Data Analysis Portal (https://ualcan.path.uab.edu/index.html)[22].
Reagents
TQFL12, a novel synthetic derivative of TQ inhibiting TNBC metastasis and invasion, was synthesized in our laboratory[17]. The RNA extraction kit was purchased from TRANGEN (cat no.: DP419, Beijing, China), and the reverse transcription polymerase chain reaction (RT-PCR) kit was purchased from Toyobo Biotech Co., Ltd (code number: FSQ-201, Shanghai, China. RPMI 1640 medium was purchased in Sigma (cat. no.: C3010-0500, St. Louis, MO, USA), and DMEM (Dulbecco’s modified Eagle’s medium) was purchased from VivaCell Biosciences Technology Co., Ltd. (cat. no.: C3113-0500, Shanghai, China). FBS (Fetal bovine serum) was purchased from Invitrogen, Irvine (cat.no.: A6907, CA, USA). The anti-ADAM9 antibody for western blotting (WB) and immunohistochemistry (IHC) was purchased in Invitrogen, Irvine (cat no.: PA5-47970, CA, USA).
IHC assay
The IHC assay was carried out on cancer tissues from Chinese patients using the ADAM9 antibody. All tissue samples were collected from the Affiliated Huaian No. 1 People’s Hospital of Nanjing Medical University (NO. YX-2021-090-01, November 19, 2021) with informed consent. For antigen retrieval, de-paraffinized and re-hydrated tissue sections were treated with sodium citrate buffer (pH 6.0, 10 μM) at
Cell culture
The human cancer cell lines 22RV1 (prostate cancer), MDA-MB-231 (triple‑negative breast cancer), and H1975 (lung adenocarcinoma) were used in this study. The 22RV1 cell line was cultured in RPMI 1640 medium supplemented with 10% (wt/vol) FBS with 1% antibiotics (penicillin-streptomycin), while MDA-MB-231 cells were cultured in DMEM medium with the same supplements. All cells were cultured in 12-well plates in an incubator at 37 °C under 5% CO2. Each group received varying amounts of TQFL12 (0, 10, 20, and 40 μM) for 24 h when the cell densities reached 50%~70%[23]. Then proteins were extracted for WB and total RNA for semi-quantitative RT-PCR after trypsinization of the cells. All cells were from American Type Culture Collection (ATCC).
Western blotting assay
WB was conducted as described previously[24]. For details, the cells were lysed with EBC buffer containing a cocktail of protease inhibitors and then incubated at 4 °C for 15 min. The cellular lysates were cleared by centrifuging at 13,000 revolutions per minute (RPM) for 15 min to remove debris. Proteins were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) (10%) electrophoresis and transferred to a polyvinylidene fluoride (PVDF) membrane (Millipore ISEQ00010) for 100 min at 100 V (300 mA maximum). Then the membrane was blocked with 3% BSA or 5% nonfat dry milk in Tris-buffered saline (TBS) with 0.1% Tween 20 (TBST) for at least 1 h and incubated with the primary goat anti-ADAM9 antibody (1:5000 dilution) overnight at 4 °C with gentle rocking/shaking. The secondary HRP-conjugated antibody was diluted in TBST with 3% BSA or 5% nonfat dry milk (1:5000 dilutions), added to the membrane, and incubated for approximately 3 h at room temperature with shaking. Membranes were rinsed three times with TBST and visualized using enhanced chemiluminescence (ECL) (Thermo ScientificTM 34580), with exposure performed as needed by a gel imaging system (Bio-Rad Laboratories, Inc., USA). β-actin served as an internal control to determine protein loading.
Quantitative RT-PCR assay
TQFL12-treated cells were used to extract total RNA for semi-quantitative RT-PCR. This RT-PCR assay was carried out to monitor ADAM9 mRNA expression. The primers’ sequences targeting ADAM9 (NM_003816.3) were RT-ADAM9-L: 5’- gcatttgtgggaacagtgtg-3’ (left primer), and RT-ADAM9-R: 5’- ccagcgtccaccaacttatt-3’ (right primer). The amplified product size was 329bp. The primer sequences targeting Actin Beta (ACTB) were RT-ACTB-5: 5’-CTCTTCCAGCCTTCCTTCCT-3’ (forward primer) and RT-ACTB-3: 5’-CACCTTCACCGTTCCAGTTT-3’ (reverse primer). The amplified product size for ACTB was 510 bp. The volume of each polymerase chain reaction was 10 µL. RT-PCR was performed semi-quantitatively as previously mentioned. The RT-PCR condition and primers for ADAM9 were described previously[24]. All experiments were repeated three times.
Molecular docking assessment
The three-dimensional structure of human full-length ADAM9 (ID: Q13443) was retrieved from the UniProt (https://www.uniprot.org/) database and prepared for docking with Chimera 1.18 software[25]. The three-dimensional structure of TQFL12 was generated by Chemaxon (https://chemaxon.com/) and converted to simulation description format (SDF) format by Open Babel software[26]. Molecular docking was performed to determine the binding free energy between TQFL12 and ADAM9 using Chimera 1.18 software and AutoDock Vina. Finally, visualization of the molecular interaction between them was generated by the Discovery Studio 2016 Client software.
Statistical analysis
Data were collected from more than three repeat experiments and analyzed using SPSS Statistics 25 (SPSS Inc., Chicago, USA). Differences among groups were analyzed using one-way analysis of variance (ANOVA) for data that fit a normal distribution. Student’s t-test was used to compare two groups. Survival analysis was performed using the Kaplan-Meier method to estimate survival probabilities, and differences between high and low ADAM9 expression groups were assessed using the log-rank test. Hazard ratios (HRs) with 95% confidence intervals (CIs) were calculated using univariate Cox proportional hazards regression analysis. A P-value of < 0.05 was considered statistically significant.
RESULTS
Expression of ADAM9 in different types of cancer
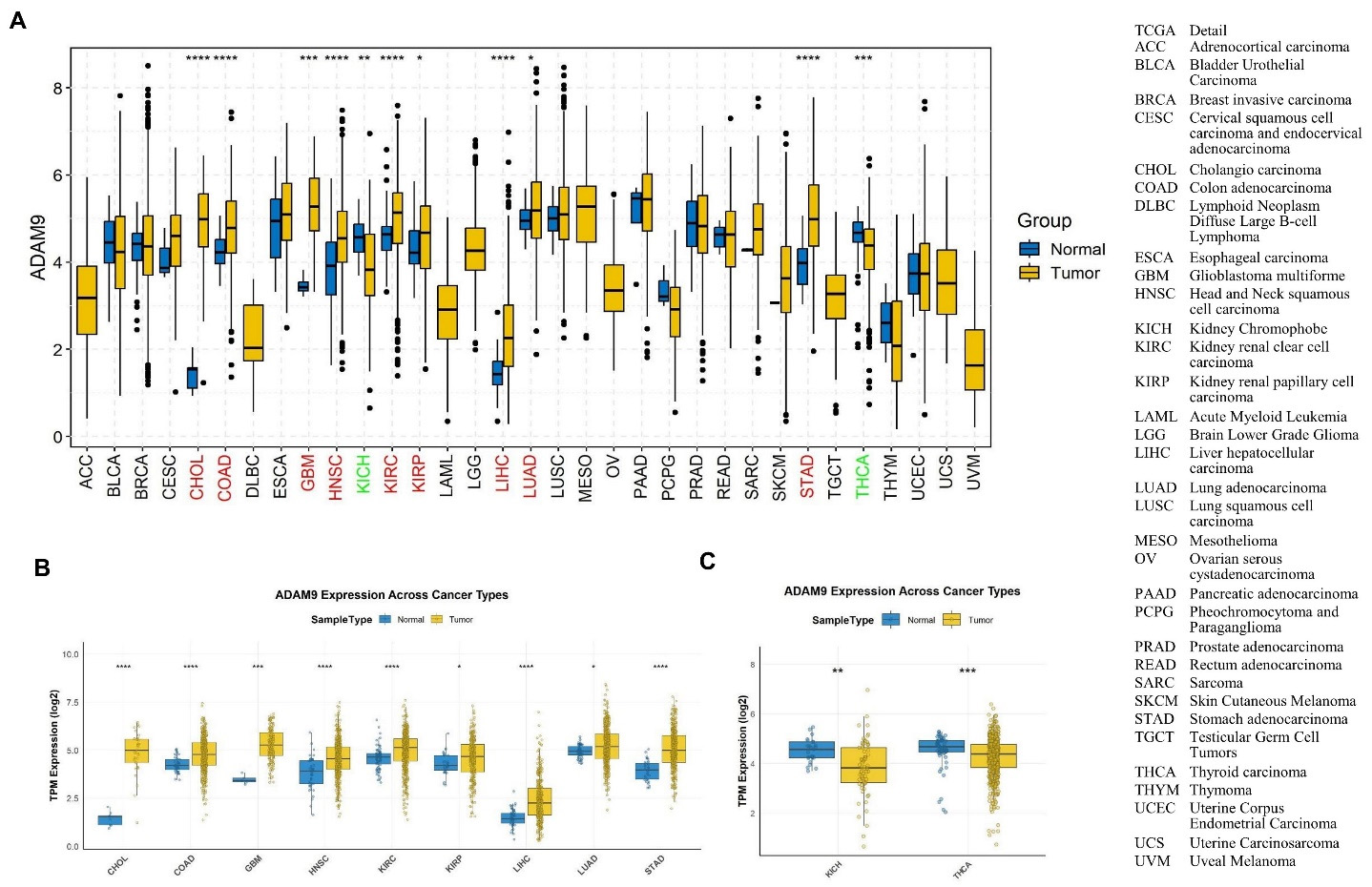
We conducted a comparison of ADAM9 expression using the TCGA plot R package, and the results are shown in Figure 1A-C. We applied multiple testing correction across all cancer-type comparisons using the Benjamini-Hochberg false discovery rate (FDR) method. The expression of ADAM9 mRNA showed a notable increase in nine types of cancer, including Cholangiocarcinoma (CHOL), Colon adenocarcinoma (COAD), Glioblastoma multiforme (GBM), Head and Neck squamous cell carcinoma (HNSC), Kidney renal clear cell carcinoma (KIRC), Kidney renal papillary cell carcinoma (KIRP), Liver hepatocellular carcinoma (LIHC), Lung adenocarcinoma (LUAD), Stomach adenocarcinoma (STAD) [Figure 1A and B], but was remarkably downregulated in Kidney chromophobe (KICH) and Thyroid carcinoma (THCA) tissues [Figure 1C] compared with normal tissues.
Figure 1. The expression of ADAM9 mRNA in pan-cancers and the corresponding normal tissues. (A) The expression of ADAM9 in the different cancers and the matched normal tissues; (B) The expression of ADAM9 is significantly higher in the different cancer tissues compared with the matched normal tissues; (C) The expression of ADAM9 is remarkably lower in the cancer tissues KICH and THCA compared with the matched normal tissues. P < 0.05 (*), P < 0.01 (**), P < 0.001 (***) and P < 0.0001 (****).
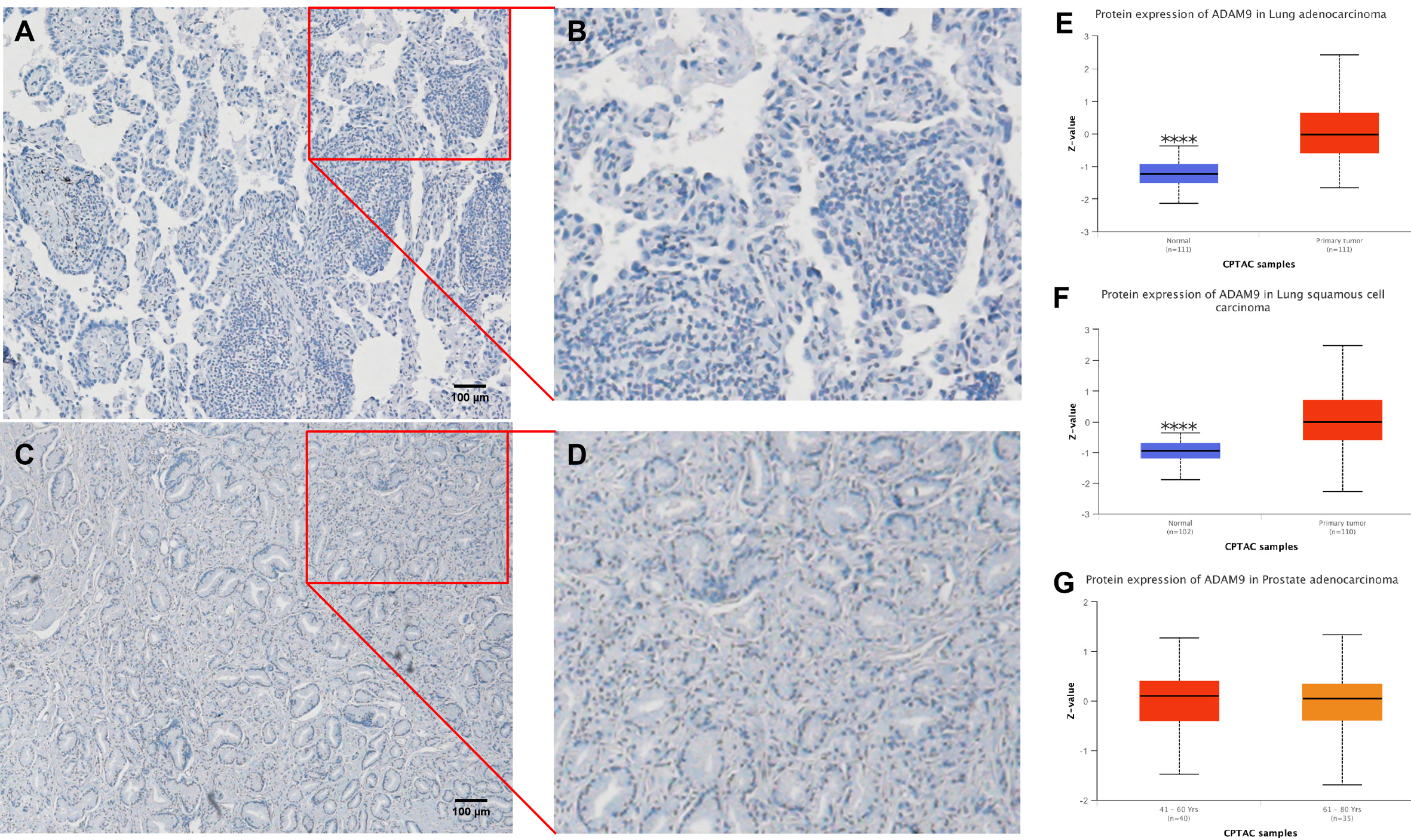
Additionally, we performed IHC on tissues from lung and prostate cancer, as well as their corresponding normal tissues. The representative results, displayed in Figure 2, show that ADAM9 mainly localizes to the vesicles, and additionally localizes to the endoplasmic reticulum in both lung [Figure 2A and B] and prostate tumor tissues [Figure 2C and D]. A significant increase in ADAM9 was observed in LUAD (Figure 2E, P = 1.34842479651913e-24) and lung squamous cell carcinoma (LUSC) (Figure 2F, P = 1.86754380592341e-18) tissues, whereas no change was observed in prostate adenocarcinoma (PRAD) tissues [Figure 2G] compared with normal tissues, according to CPTAC.
Figure 2. ADAM9 location and protein levels in cancer tissues from the lung and prostate. (A) Representative staining by IHC (immunohistochemistry) of Chinese lung adenocarcinoma (LUAD) tissues; (B) Enlarged picture of A; (C) Representative staining by IHC from a Chinese prostate cancer patient; (D) Magnified image of C; (E) Comparison of ADAM9 protein levels between lung adenocarcinoma (LUAD) and the corresponding matched normal tissues. P < 0.0001 (****); (F) Comparison of ADAM9 protein levels between lung squamous cell carcinoma (LUSC) and the corresponding matched normal tissues. P < 0.0001 (****); (G) Comparison of ADAM9 protein levels between prostate adenocarcinoma (PRAD) and the corresponding matched normal tissues. Images of A, B, C, and D magnified 100X.
The prognostic values for ADAM9 expression in pan-cancers
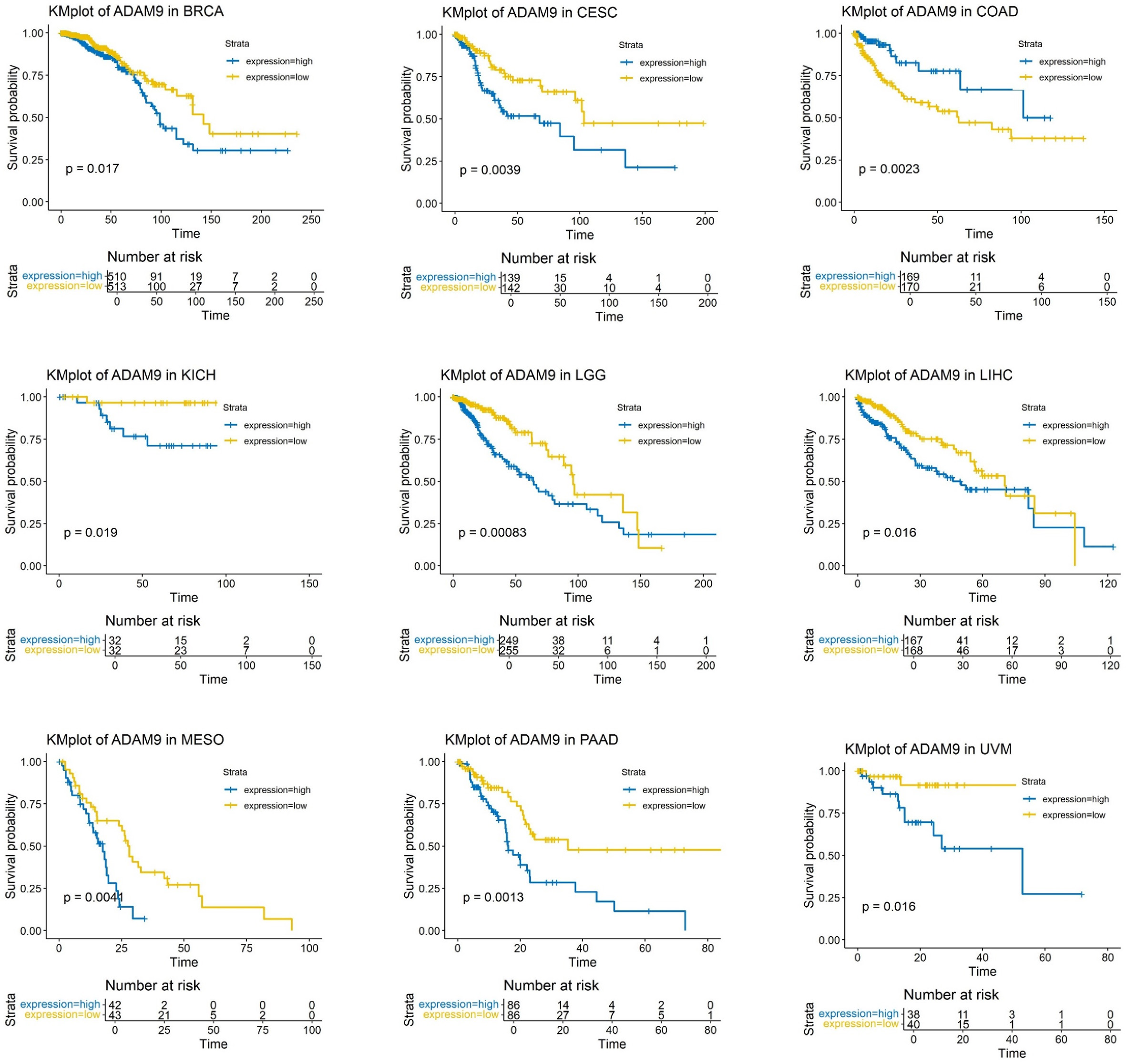
To explore the prognostic values, the correlation between ADAM9 expression and prognosis was analyzed by TCGA plot R package, and it was discovered that higher expression of ADAM9 is correlated with shorter OS in Breast Invasive Carcinoma (BRCA), Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma (CESC), Kidney Chromophobe (KICH), Brain Lower Grade Glioma (LGG), LIHC, Mesothelioma (MESO), Pancreatic Adenocarcinoma (PAAD), Uveal Melanoma (UVM) [Figure 3], but with longer OS only in COAD [Figure 3]. Thus, high expression of ADAM9 may be a marker of unfavorable outcomes in patients with BRCA, CESC, KICH, LGG, LIHC, MESO, PAAD, and UVM, and a favorable marker for COAD.
Figure 3. Overall survival for ADAM9 expression in pan-cancers. Overall survival for ADAM9 expression in BRCA, CESC, COAD, KICH, LGG, LIHC, MESO, PAAD, and UVM, respectively. The P < 0.05 was considered statistically significant. BRCA: Breast invasive carcinoma; CESC: cervical squamous cell carcinoma; COAD: colon adenocarcinoma; KICH: kidney chromophobe; LGG: brain lower grade glioma; LIHC: liver hepatocellular carcinoma; MESO: mesothelioma; PAAD: pancreatic adenocarcinoma; UVM: uveal melanoma.
ADAM9 mutation and prognosis in tumor tissues and the matched normal tissues
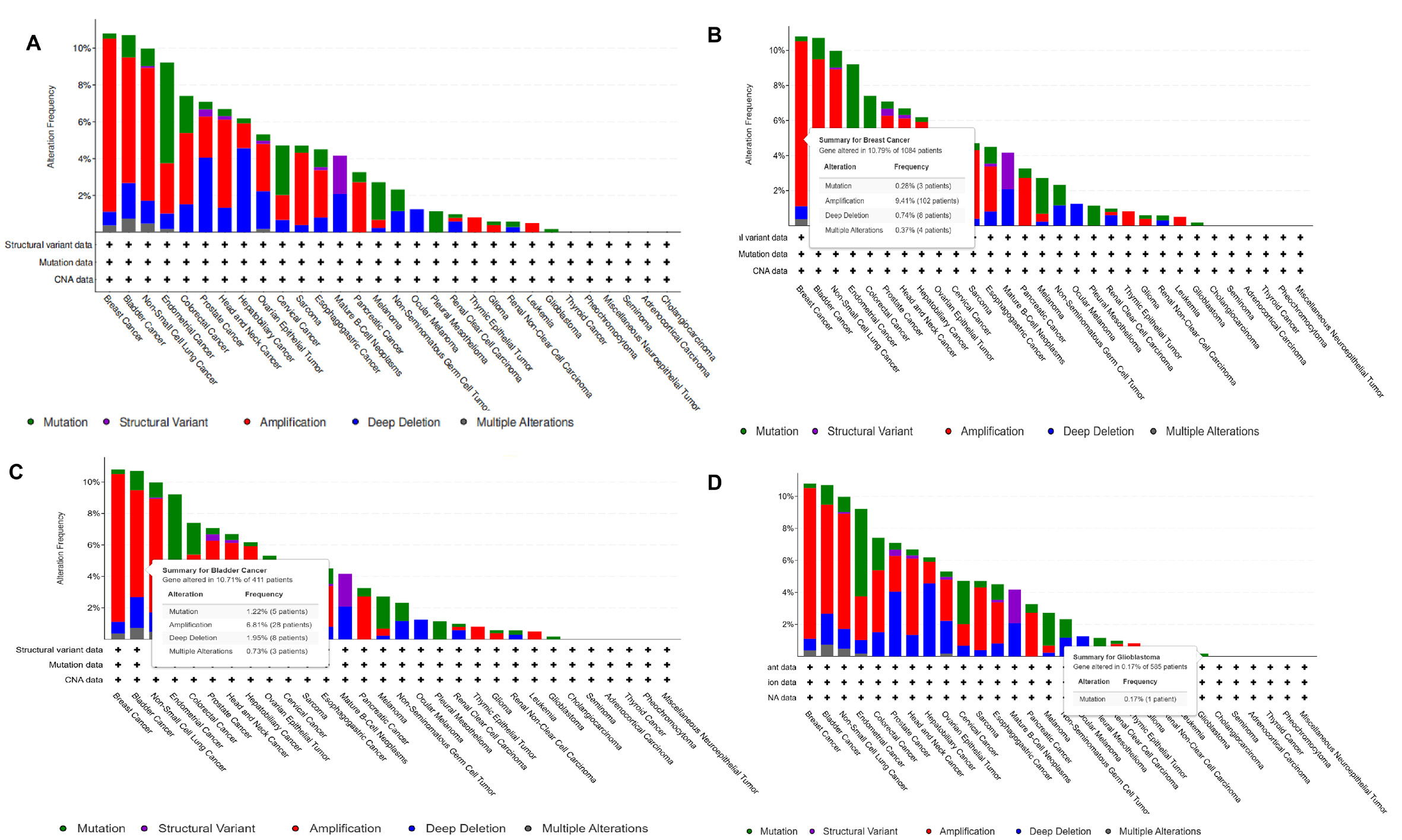
ADAM9 variants have been shown to be potential indicators of recurrence in prostate cancer patients after receiving a radical prostatectomy. The genetic variant of rs6474526 in ADAM9 affected its expressions, epidermal growth factor receptor (EGFR) mutation status, and disease progression in patients with LUAD. We thus explored ADAM9 mutations in different tumor tissues. In Figure 4, mutations are shown in green, amplifications in red, and deep deletions in blue. The detailed landscape of ADAM9 mutations in pan-cancers is shown in Figure 4A. Amplifications were predominant in most cancers. The highest alteration frequency was observed in BRCA (10.79% of 1084 cases; Figure 4B), followed by Bladder Carcinoma (BLCA) (10.71% of 411 cases; Figure 4C), while GBM showed the lowest frequency (0.57% of 348 cases; Figure 4D). No ADAM9 variants were found in CHOL, Seminoma, Pheochromocytoma and paraganglioma (PEPG), THCA, and Adrenocortical carcinoma (ACC).
Figure 4. ADAM9 deleterious variants in pan-cancers. (A) The detailed landscapes of ADAM9 mutations in pan-cancers; (B); The detailed alteration frequency of mutations in BRCA; (C) The detailed alteration frequency of mutations in BLCA; (D) The detailed alteration frequency of mutations in GBM. BRCA: Breast invasive carcinoma; BLCA: bladder carcinoma; GBM: glioblastoma multiforme; CNA: copy number alteration; NA: not available.
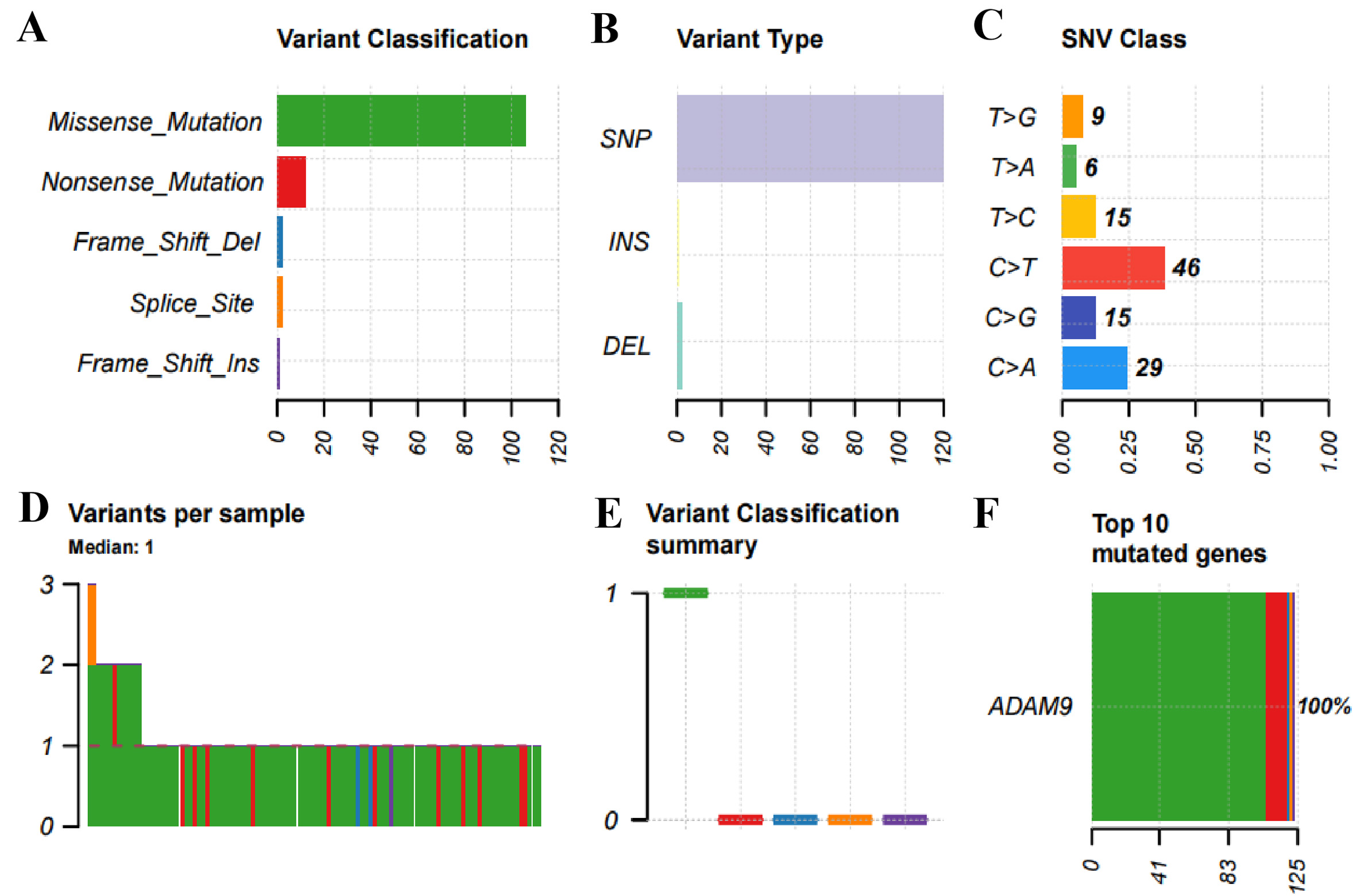
We also summarized the count and type of variants at the sample and gene levels of ADAM9. As shown in Figure 5, we systematically analyzed the mutation characteristics of the ADAM9 gene set across selected cancer types. Figure 5A presents the variant classification, showing the number of harmful mutations in the ADAM9 gene set for specific cancer types. Figure 5B displays the variant type, including the count of single nucleotide polymorphisms (SNPs), insertions (INS), and deletions (DELs) in the ADAM9 gene set. Figure 5C illustrates the single nucleotide variant (SNV) class, indicating the count of each SNV category. Figure 5D shows the number of variants per sample, in which each bar represents one sample, and the color corresponds to the variant classification. Figure 5E uses a box plot to exhibit the distribution of each variant classification count in the sample set. Figure 5F finally shows the number and proportion of variations in the top ten mutant genes. The results demonstrated that ADAM9 is highly mutated.
Figure 5. Plot summarizes the count and type of variants in the sample and gene levels of ADAM9. (A) Variant Classification: the number of harmful mutations in the ADAM9 gene set for specific cancer types; (B) Variant Type: the count of SNPs, INS and DELs in the ADAM9 gene set for selected cancer types; (C). SNV class: the count of each SNV class in the ADAM9 gene set for selected cancer types; (D) Variants per sample: the count of variants in each sample. One bar indicates a sample, and the color of the bar corresponds to the variant classification; (E) The distribution of each variant classification’s count in the sample set of chosen cancer types is displayed using a box plot; (F) The number and proportion of variations in the top ten mutant genes. SNPs: Single nucleotide polymorphisms; INS: insertions; DELs: deletions; SNV: single nucleotide variant.
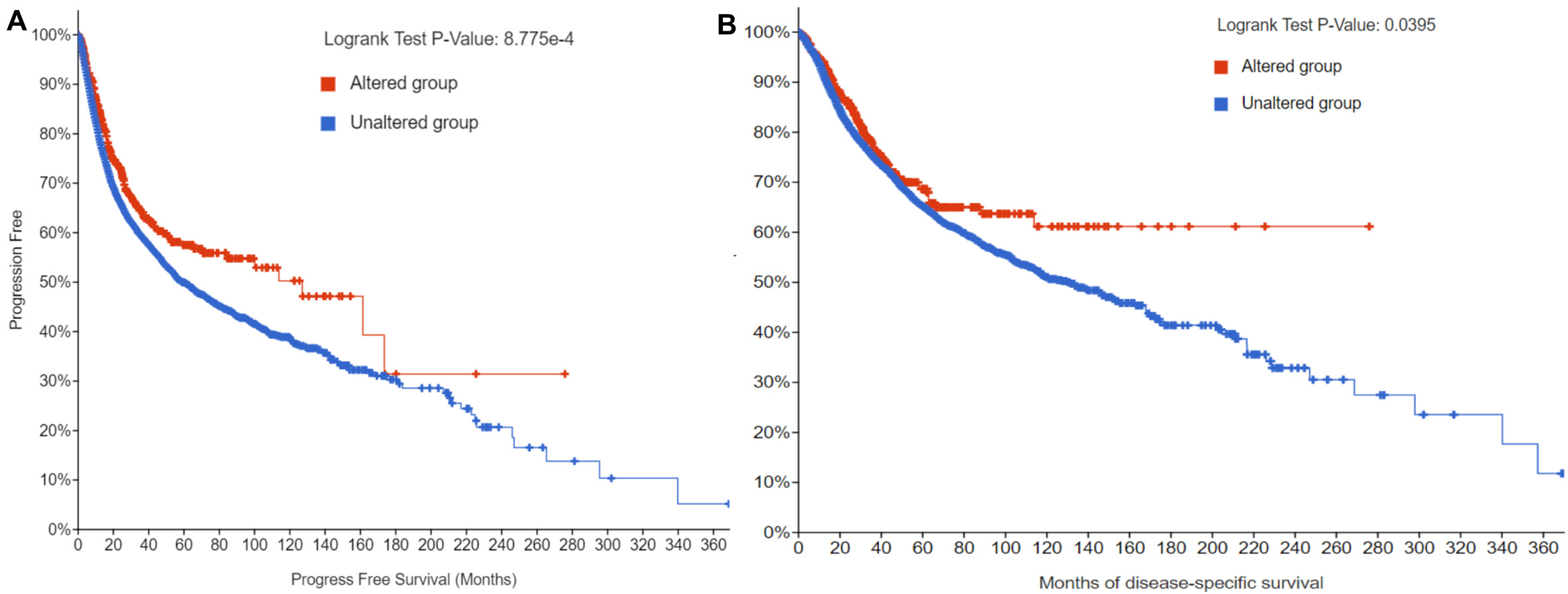
To further analyze the prognostic value of ADAM9 variants in cancers, survival differences between the ADAM9-altered and unaltered groups were assessed in pan-cancer cohorts. ADAM9 mutations were associated with significantly prolonged progression-free survival (PFS) and disease-specific survival (DSS) (Figure 6; P = 8.775e-4 and P = 0.0395, respectively; Table 1). For the group that remained unchanged, the median PFS months were 58.95 (55.10-65.88, 95%CI), while the altered group increased to 126.97 months (83.93-NA, 95%CI) [Figure 6A]. The median DSS months for the unchanged group were 131.51 months (116.84-152.02, 95%CI), but significantly increased for the altered group (NA, 95%CI) [Figure 6B]. Therefore, mutations of the ADAM9 gene might considerably influence disease prognosis in pan-cancers, including PFS and DSS, which indicates that ADAM9 variants could be a favorable biomarker in pan-cancer.
Figure 6. Effects of ADAM9 variants on prognosis of diseases across various cancers. (A) Progression-free survival for wild-type and mutant ADAM9; (B) Disease-specific survival for wild-type and mutant ADAM9. The data were analyzed by Cox regression.
Survival in pan-cancers with wild-type and mutant ADAM9
| Survival type | Number of patients | P-value | Q-value |
| Progression free | 10,613 | 8.78E-04 | 3.51E-03 |
| Disease-specific | 10,258 | 0.0395 | 0.079 |
| Disease-free | 5,383 | 0.152 | 0.203 |
| Overall | 10,803 | 0.518 | 0.518 |
TQFL12 inhibits ADAM9 expression in cancer cells
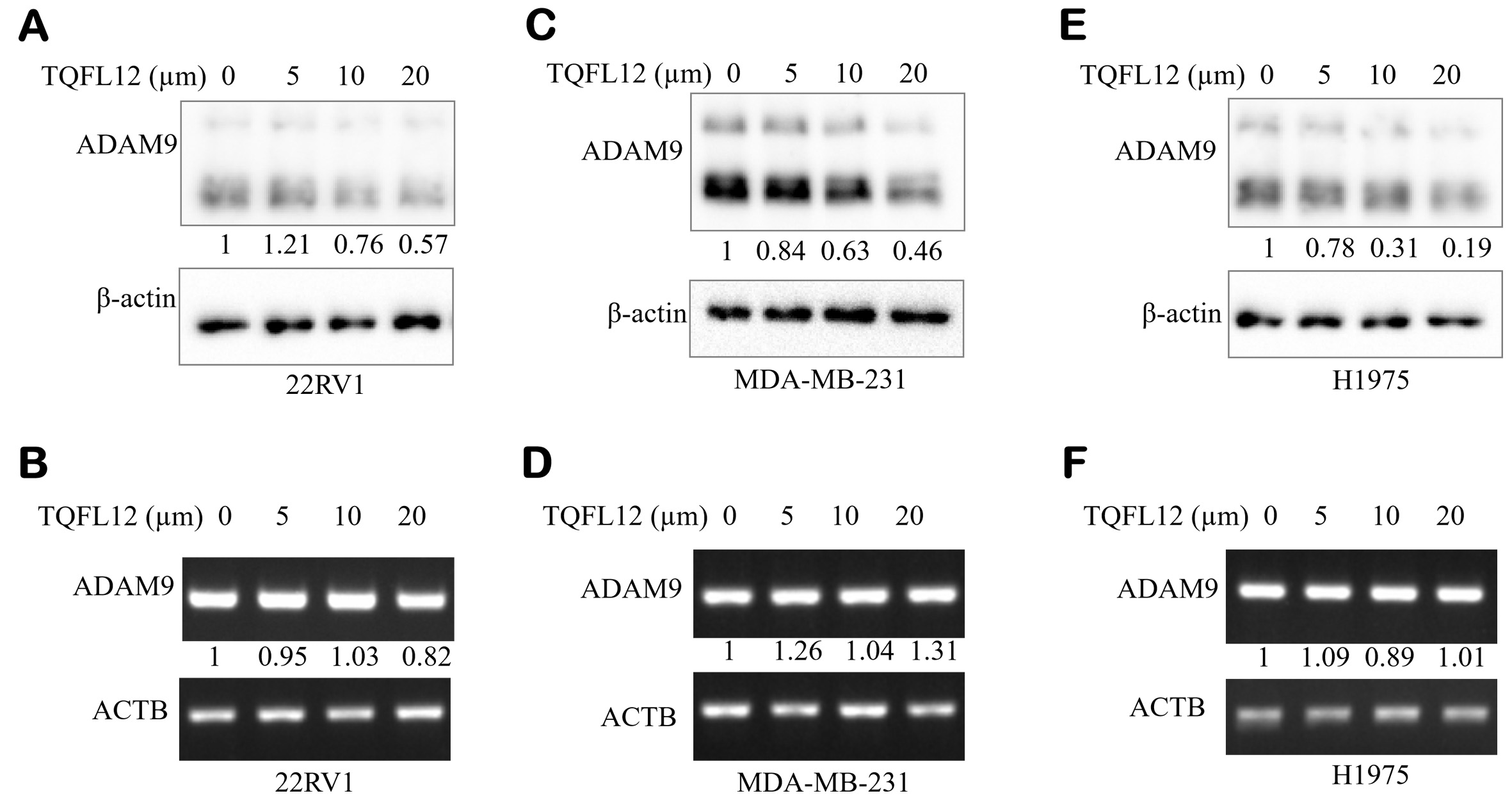
As described above, TQFL12 is a novel synthetic derivative of TQ that inhibits TNBC metastasis and invasion, and was synthesized in our laboratory. The small molecules may regulate ADAM9 expression; thus, we further investigate the regulation of ADAM9 expression in various cancer cells by TQFL12. As shown in Figure 7, TQFL12 inhibits ADAM9 protein expression but not mRNA expression in the prostate cancer cell line 22RV1 [Figure 7A and B], in the breast cancer cell line MDA-MB-231 [Figure 7C and D], and in the lung cancer cell line H1975 [Figure 7E and F] in a dosage-dependent manner.
Figure 7. TQFL12 inhibits the expression of ADAM9 in different cancer cell types. (A). TQFL12 inhibits ADAM9 protein in the 22RV1 cell line; (B) ADAM9 mRNA expression after TQFL12 treatment in the 22RV1 cell line; (C) TQFL12 inhibits ADAM9 protein in the MDA-MB-231 cell line; (D) ADAM9 mRNA expression after TQFL12 treatment in the MDA-MB-231 cell line; (E) TQFL12 inhibits ADAM9 protein in the H1975 cell line; (F) ADAM9 mRNA expression after TQFL12 treatment in the H1975 cell line. The β-actin protein or the ACTB gene were used as loading controls.
TQFL12 binds to ADAM9
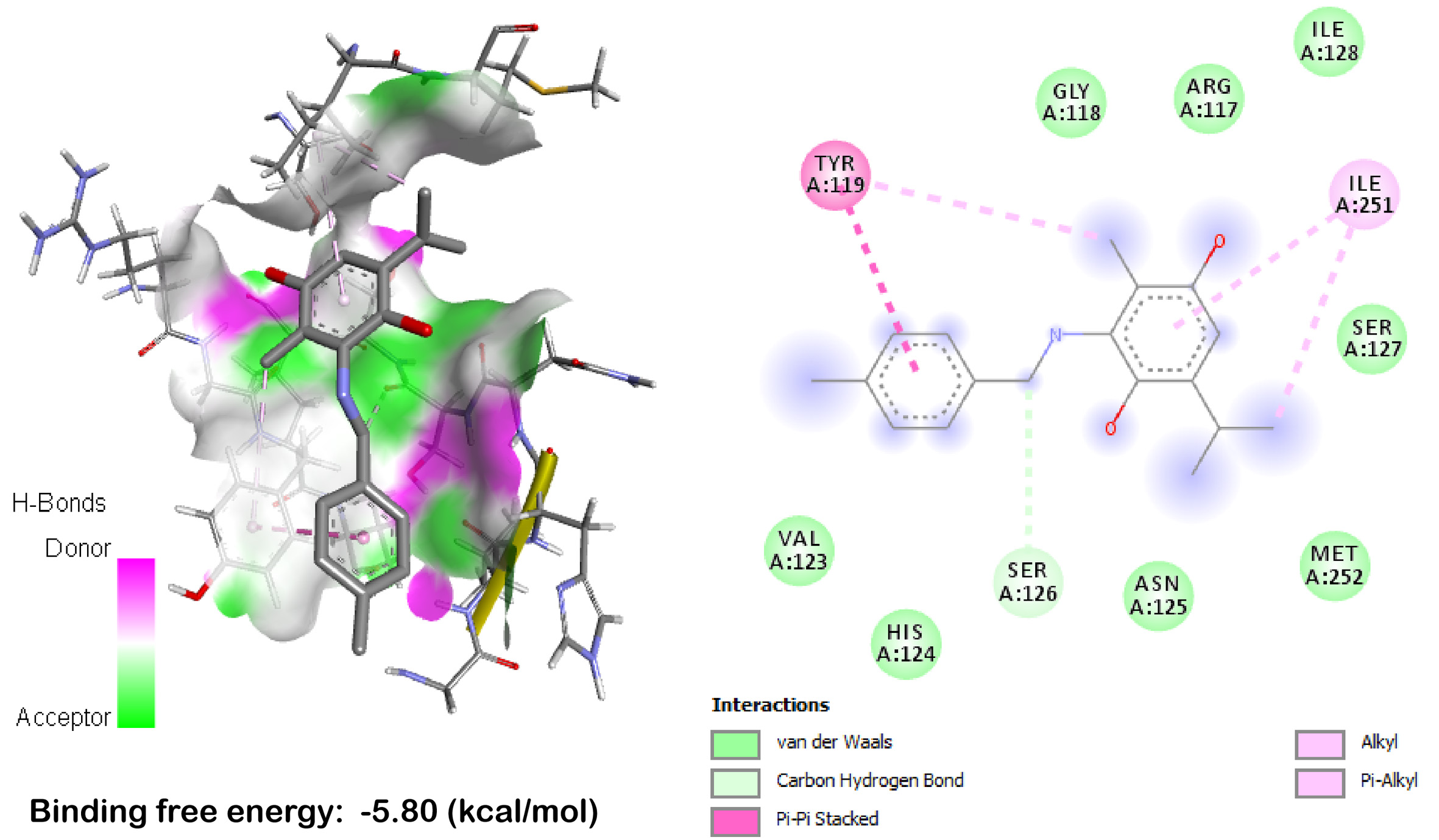
Molecular docking interaction of TQFL12 and ADAM9 was performed. The data in Table 2 and Figure 8 revealed the binding free energy of TQFL12 toward the binding site of ADAM9. TQFL12 interacted with TYR119 (hydrophobic interaction), SER126 (hydrogen bond), and ILE251 (hydrophobic interaction) residues in the binding site of ADAM9 with a binding free energy of -5.80 (kcal/mol).
Figure 8. TQFL12 binds to ADAM9 by molecular docking. TYR: Tyrosine; GLY: glycine; ARG: arginine; ILE: isoleucine; SER: serine; VAL: valine; HIS: histidine; SER: serine; ASN: asparagine; MET: methionine.
Molecular docking interaction of TQFL12 and ADAM9
| Residues | Hydrogen bond | Charge | Hydrophobic interaction | Halogen |
| TYR119 | 0 | 0 | 1 | 0 |
| SER126 | 1 | 0 | 0 | 0 |
| ILE251 | 0 | 0 | 1 | 0 |
DISCUSSION
ADAM9 gene, located on human chromosome 8p11.23, encodes a type I transmembrane glycoprotein[27]. As a crucial member of the ADAM family, ADAM9 is significantly overexpressed in various malignant tumors, including lung, prostate, pancreatic, breast, and colorectal cancers[1,11]. Research demonstrates that ADAM9 promotes tumor progression through dual mechanisms involving both its metalloproteinase domain and non-proteolytic pathways. On one hand, ADAM9 cleaves and releases multiple key molecules associated with tumorigenesis and angiogenesis [such as Heparin binding epidermal-like growth factor (HB-EGF), Vascular Endothelial Growth Factor A (VEGFA), and Ephrin Receptor B4 (EPHB4)], thereby activating EGFR signaling pathways and promoting tumor proliferation and vascular remodeling[28,29]. On the other hand, ADAM9 induces EMT and significantly enhances tumor cell invasion and metastasis by regulating microRNAs (miRNAs) (e.g., miR-1, miR-218) and stabilizing N-cadherin[5,30]. A recent groundbreaking study revealed that inhibiting ADAM9 promotes selective degradation of Kirsten rat sarcoma viral oncogene (KRAS), sensitizing pancreatic cancers to chemotherapy[6]. Additionally, ADAM9 genetic polymorphisms have been validated as independent predictors for biochemical recurrence in prostate cancer patients after radical prostatectomy[31]. These findings position ADAM9 as an important potential target for precision tumor diagnosis and targeted therapy.
Our comprehensive analysis revealed a complex and context-dependent role of ADAM9 in tumor biology and pan-cancer. Firstly, ADAM9 mRNA was remarkably upregulated in nine cancer types (CHOL, COAD, GBM, HNSC, KIRC, KIRP, LIHC, LUAD, and STAD) but downregulated in KICH and THCA, demonstrating significant tumor heterogeneity in ADAM9 expression patterns[32,33]. This differential expression likely reflects distinct activation states of signaling pathways in various tumor microenvironments; for instance, the frequent activation of EGFR pathway in lung and head and neck cancers can induce ADAM9 transcription[28,29]. Mechanistically, ADAM9 overexpression promotes tumor cell proliferation, angiogenesis, and metastasis through dual pathways: its metalloproteinase activity releases pro-growth factors (e.g., HB-EGF and VEGFA) and non-proteolytically stabilizes adhesion molecules such as N-cadherin[5,28,30], which aligns with the pro-tumorigenic phenotype observed in most cancer types in our study.
Notably, the association between ADAM9 expression levels and patient prognosis also exhibited cancer-type specificity. High ADAM9 expression correlated with longer OS only in COAD, whereas it predicted shorter OS in BRCA, CESC, KICH, LIHC, LGG, MESO, PAAD, and UVM, suggesting its utility as a biomarker of unfavorable outcomes in these cancers[8,31]. This prognostic discrepancy may reflect the functional plasticity of ADAM9 in different tumor contexts. For example, in colorectal cancer, ADAM9 might participate in tissue repair or immune modulation, whereas in pancreatic and liver cancers, it primarily exerts pro-invasive effects[6,12].
In this study, high ADAM9 expression was associated with better OS in COAD, which contrasts with reports in most cancer types where ADAM9 serves as a poor prognostic factor[9,34,35]. This seemingly paradoxical phenomenon may be explained by the following mechanisms: ADAM9 functions in a highly context-dependent manner. First, its cellular source is crucial - in colorectal cancer, the pro-tumorigenic effects of ADAM9 primarily derive from cancer-associated fibroblasts (CAFs) rather than tumor cells themselves[9,36]. When high ADAM9 expression reflects stromal components, its prognostic significance may differ. Second, ADAM9’s immunomodulatory functions are cancer type-specific; mechanisms promoting immune evasion in hepatocellular carcinoma[2,35] may not dominate in the COAD microenvironment. Third, downstream pathways activated by ADAM9 vary by cell type - in colon cancer, ADAM9 primarily promotes invasion rather than proliferation[36], which may paradoxically associate with longer survival.
Furthermore, our study is the first to systematically unveil the prognostic value of ADAM9 mutations across pan-cancers: patients with ADAM9 mutations exhibited significantly superior PFS and DSS compared to wild-type carriers, with mutations distributed across the entire gene[32,37]. This intriguing finding implies that loss-of-function mutations may reduce the pro-tumorigenic activity of ADAM9 and thereby improve clinical outcomes, providing a genetic rationale for ADAM9-targeted therapy. It further suggests that inhibiting ADAM9 function could replicate the survival benefit conferred by mutations in certain cancer patients[6]. The unexpected association between ADAM9 mutations and favorable clinical outcomes (longer progression-free and disease-specific survival) can be attributed to the subtype-specific functional effects of ADAM9 mutations and their context-dependent regulatory roles in tumor progression and anti-tumor immunity, rather than a simple loss-of-function mode[2,35]. As a multidomain metalloprotease, ADAM9 mutations identified in this study are predominantly missense variants that do not abrogate protein function but alter its stability, substrate specificity or proteolytic activity[2,38]. Unlike truncating mutations that eliminate metalloprotease activity, these missense mutations reduce ADAM9-mediated shedding of major histocompatibility complex class I-related chain A (MICA) - a key ligand for natural killer (NK) cells - thereby enhancing NK cell-mediated tumor surveillance and inhibiting immune escape. This represents a gain-of-function effect in anti-tumor immunity that contributes to improved survival[35]. Additionally, ADAM9 missense mutations can disrupt its pro-oncogenic signaling crosstalk with EGFR and integrin pathways[2], attenuating tumor cell migration and invasion without impairing normal physiological functions. This context-dependent functional divergence of ADAM9 mutations, which reshapes tumor-immune microenvironment interactions and blunts oncogenic signaling, explains the paradoxical prognostic association and highlights the heterogeneous biological roles of ADAM9 in cancer[2,35].
Natural product-derived small molecules play significant roles in anti-tumor and antiviral activities, possibly through regulating ADAM family protein expression[39]. Given their potential role in modulating ADAM9, we further investigated the effect of TQFL12 on ADAM9 expression. TQFL12 (molecular formula: C17H16ClNO2; molecular weight: 302.09) is a novel synthetic derivative of TQ that inhibits TNBC metastasis and invasion and was synthesized in our laboratory[31]. The results showed that TQFL12 inhibits ADAM9 protein expression, but not mRNA expression, in prostate cancer and breast cancer cell lines in a dose-dependent manner. In contrast, TQ has been shown to inhibit protein expression of ADAM17, a family member of ADAM9, in the breast cancer cell line MCF7 and lung cancer cell line H460[39]. Beyond natural product-derived compounds, emerging gene and cell-based therapeutic strategies, including Chimeric antigen receptor T (CAR-T) and Clustered Regularly Interspaced Short Palindromic Repeats – CRISPR associated protein 9 (CRISPR-Cas9) approaches, are being actively explored for prostate cancer treatment[40].
TQFL12 is a novel synthetic derivative optimized from the natural product TQ that demonstrates significant anti-tumor activity across multiple cancer models[4,31]. As a structural optimization product of TQ, TQFL12 overcomes the limitation of weak anti-tumor effects of the parent compound, showing enhanced efficacy and reduced toxicity in both in vitro and in vivo studies[4,31]. Research indicates that TQFL12 exerts its anti-cancer effects primarily through activation of the AMP-activated protein kinase (AMPK)/ACC signaling pathway, which inhibits proliferation, migration, and invasion of TNBC cells and induces apoptosis[31]. In TNBC cell line-derived xenograft mouse models, TQFL12 effectively suppressed tumor growth and distant metastasis with markedly lower toxicity compared to TQ[4,31]. Notably, a pan-cancer study further demonstrated that TQFL12, as a small-molecule inhibitor, exhibits broad-spectrum anti-tumor effects across multiple cancer types, and its inhibitory effect is associated with TMPRSS2 expression levels and mutational status, suggesting that TQFL12 may influence tumor progression[18]. Moreover, molecular docking experiments confirmed that TQFL12 can directly interact with and stabilize AMP-activated protein kinase α (AMPKα) protein, leading to sustained activation of the AMPK/ACC pathway[31]. These findings suggest that TQFL12, as a highly effective and low-toxicity drug candidate, holds significant therapeutic potential for the treatment of malignant tumors including breast cancer.
TQFL12 significantly downregulated ADAM9 protein expression but showed no obvious effect on ADAM9 mRNA levels, indicating that the regulatory effect of TQFL12 on ADAM9 may occur at the post-translational or translational level rather than at the transcriptional level. Previous studies have demonstrated that ADAM9 protein can be regulated by phosphorylation-dependent protein stability[41], histone methylation-mediated epigenetic control[42], and N6 -Methyladenosine (m6A) RNA methylation-modulated translation[43], which may represent potential mechanisms underlying the regulation of ADAM9 by TQFL12.
Although our work has yielded a highly promising discovery, several limitations remain. Our study demonstrated that TQFL12 effectively downregulates ADAM9 protein expression and inhibits the malignant phenotypes of cancer cells in vitro. Although in vivo validation has not been performed in the current work, the therapeutic potential of targeting ADAM9 has been extensively validated in preclinical models. Specifically, genetic silencing, neutralizing antibody treatment, or pharmacological inhibition of ADAM9 has been shown to significantly suppress tumor growth, invasion, and metastasis in nude mouse xenograft models of melanoma, prostate cancer, and lung cancer[44-46]. These findings provide a solid foundation for the hypothesis that TQFL12 may serve as a promising candidate for ADAM9-targeted cancer therapy. The in vivo anti-tumor efficacy and detailed mechanism of TQFL12 will be further investigated in future research. However, nude mouse tumor models or other animal experiments should be included to demonstrate that TQFL12 can downregulate ADAM9 protein expression in tumor tissues in vivo and inhibit tumor growth or metastasis.
Although we demonstrated that TQFL12 inhibits ADAM9 protein expression without affecting its mRNA levels, the precise post-transcriptional regulatory mechanism remains to be elucidated, including its potential effects on protein translation, stability, or degradation pathways[27]. Third, the clinical value of ADAM9 as a prognostic biomarker and TQFL12 as a therapeutic agent requires validation through prospective clinical trials, which are currently lacking[31].
CONCLUSION
Altogether, our studies provide an understanding of ADAM9 expression, mutation, prognosis, and its significance in different cancer types. Targeting ADAM9 with the small-molecule TQFL12 might provide a potential therapeutic strategy for various cancers, such as prostate and breast cancer. Future research may focus on further exploring the underlying mechanisms and conducting preclinical and clinical trials to validate the therapeutic potential of TQFL12 in cancer treatment.
DECLARATIONS
Acknowledgments
The authors thank the colleagues from the Research Center for Preclinical Medicine, Southwest Medical University.
Authors’ contributions
Performed experimental studies, data analysis, tissue collection, and literature searches: Zhang F, Cheng J, Fu J (Jiewen Fu), Tan Q, Du J, Zhang L, Liu Z, Dong Y
Conducted data analysis and data acquisition: Zhang F, Cheng J, Fu J (Junjiang Fu), Maghsoudloo M, El-Far A, Guo K
Wrote the manuscript: Maghsoudloo M, Zhang F
Revised the manuscript: Zhang F, Cheng J, Maghsoudloo M, Fu J (Junjiang Fu)
Supervised the project: Fu J (Junjiang Fu), Maghsoudloo M, Cheng J
All authors reviewed and approved the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
AI and AI-assisted tools statement
During the preparation of this manuscript, the AI tool Doubao (version Doubao-Seed-2.0, released 2026-02-14) was used solely for language editing. The tool did not influence the study design, data collection, analysis, interpretation, or the scientific content of the work. All authors take full responsibility for the accuracy, integrity, and final content of the manuscript.
Financial support and sponsorship
This work was supported by the Key Project of Applied Basic Research of Southwest Medical University (Grant No. 2023ZD010), the Foundation of Science and Technology Department of Sichuan Province (Grant No. 2025ZNSFSC0986), and in part by Sichuan Provincial Administration of Traditional Chinese Medicine research project (Grant No.: 2023MS235), the Huaian Science and Technology Plan Project (Grant No. HAB2024004) and the Zigong Key Science and Technology Plan (Collaborative Innovation Project of Zigong Academy of Medical Sciences) (Grant No.2023YKYXT09) and Project of Medical and Health Communication Research Center of Zigong Social Sciences Association in 2025 (Project No. YXJKCB-2025-10).
Conflicts of interest
Fu J is the Guest Editor of the Special Issue “Power of Epigenetics in Tumor Detection, Treatment, and Prognosis” of Journal of Translational Genetics and Genomics and the Section Editor of Journal of Translational Genetics and Genomics. He was not involved in any steps of the editorial processing, notably including reviewer selection, manuscript handling, or decision-making. The other authors declared that there are no conflicts of interest.
Ethical approval and consent to participate
The study was approved by the Affiliated Huaian No. 1 People’s Hospital of Nanjing Medical University (NO. YX-2021-090-01, November 19, 2021). Informed consent was obtained from patients for tissue samples. We confirmed that all methods were carried out in accordance with relevant guidelines and regulations.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Arora S, Scott AM, Janes PW. ADAM proteases in cancer: biological roles, therapeutic challenges, and emerging opportunities. Cancers. 2025;17:1703.
2. Chou CW, Huang YK, Kuo TT, Liu JP, Sher YP. An overview of ADAM9: structure, activation, and regulation in human diseases. Int J Mol Sci. 2020;21:7790.
3. Parry DA, Toomes C, Bida L, et al. Loss of the metalloprotease ADAM9 leads to cone-rod dystrophy in humans and retinal degeneration in mice. Am J Hum Genet. 2009;84:683-91.
4. Liu JP, Shen KY, Cheng WC, et al. ADAM9 drives the immunosuppressive microenvironment by cholesterol biosynthesis-mediated activation of IL6-STAT3 signaling for lung tumor progression. Am J Cancer Res. 2024;14:1850-65.
5. Slapak EJ, El Mandili M, Brink MST, Kros A, Bijlsma MF, Spek CA. Preclinical assessment of ADAM9-responsive mesoporous silica nanoparticles for the treatment of pancreatic cancer. Int J Mol Sci. 2023;24.
6. Huang YK, Cheng WC, Kuo TT, et al. Inhibition of ADAM9 promotes the selective degradation of KRAS and sensitizes pancreatic cancers to chemotherapy. Nat Cancer. 2024;5:400-19.
7. Le TT, Hsieh CL, Lin IH, et al. The ADAM9/UBN2/AKR1C3 axis promotes resistance to androgen-deprivation in prostate cancer. Am J Cancer Res. 2022;12:176-97.
8. Wu S, Cheng L, Luo T, Makeudom A, Wang L, Krisanaprakornkit S. Overexpression of a disintegrin and metalloproteinase 9 (ADAM9) in relation to poor prognosis of patients with oral squamous cell carcinoma. Discov Oncol. 2024;15:582.
9. Ao T, Mochizuki S, Kajiwara Y, et al. Cancer-associated fibroblasts at the unfavorable desmoplastic stroma promote colorectal cancer aggressiveness: potential role of ADAM9. Int J Cancer. 2022;150:1706-21.
10. Romanowicz A, Lukaszewicz-Zajac M, Mroczko B. Exploring potential biomarkers in oesophageal cancer: a comprehensive analysis. Int J Mol Sci. 2024;25:4253.
11. Micocci KC, Moritz MN, Lino RL, et al. ADAM9 silencing inhibits breast tumor cells transmigration through blood and lymphatic endothelial cells. Biochimie. 2016;128-129:174-82.
12. Zhang XY, Zhao CY, Dong JM, et al. ADAM9 mediates Cisplatin resistance in gastric cancer cells through DNA damage response pathway. Med Oncol. 2025;42:122.
13. Jotatsu Y, Sung SY, Wu MH, et al. An antibody of the secreted isoform of disintegrin and metalloprotease 9 (sADAM9) inhibits epithelial-mesenchymal transition and migration of prostate cancer cell lines. Int J Mol Sci. 2024;25:6646.
14. Ding YF, Ho KH, Lee WJ, et al. Cyclic increase in the histamine receptor H1-ADAM9-Snail/Slug axis as a potential therapeutic target for EMT-mediated progression of oral squamous cell carcinoma. Cell Death Dis. 2025;16:191.
15. Banerjee S, Padhye S, Azmi A, et al. Review on molecular and therapeutic potential of thymoquinone in cancer. Nutr Cancer. 2010;62:938-46.
16. Ahmad A, Mishra RK, Vyawahare A, et al. Thymoquinone (2-Isoprpyl-5-methyl-1, 4-benzoquinone) as a chemopreventive/anticancer agent: chemistry and biological effects. Saudi Pharm J. 2019;27:1113-26.
17. Wei C, Zou H, Xiao T, et al. TQFL12, a novel synthetic derivative of TQ, inhibits triple-negative breast cancer metastasis and invasion through activating AMPK/ACC pathway. J Cell Mol Med. 2021;25:10101-10.
18. Mollica V, Rizzo A, Massari F. The pivotal role of TMPRSS2 in coronavirus disease 2019 and prostate cancer. Future Oncol. 2020;16:2029-33.
19. Uhlén M, Fagerberg L, Hallström BM, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347:1260419.
20. Uhlen M, Zhang C, Lee S, et al. A pathology atlas of the human cancer transcriptome. Science. 2017;357:eaan2507.
21. Liao C, Wang X. TCGAplot: an R package for integrative pan-cancer analysis and visualization of TCGA multi-omics data. BMC Bioinformatics. 2023;24:483.
22. Chandrashekar DS, Karthikeyan SK, Korla PK, et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18-27.
23. Zhang L, Wei C, Li D, et al. COVID-19 receptor and malignant cancers: association of CTSL expression with susceptibility to SARS-CoV-2. Int J Biol Sci. 2022;18:2362-71.
24. Jiang L, Huang W, Cao M, et al. Deciphering the oncogenic potential of ADAM9 in hepatocellular carcinoma through bioinformatics and experimental approaches. Sci Rep. 2024;14:26432.
25. Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605-12.
26. O'Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open babel: an open chemical toolbox. J Cheminform. 2011;3:33.
27. Zheng R, Wan C, Mei S, et al. Cistrome data browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res. 2019;47:D729-35.
28. Lin CY, Chen HJ, Huang CC, et al. ADAM9 promotes lung cancer metastases to brain by a plasminogen activator-based pathway. Cancer Res. 2014;74:5229-43.
29. Lin CY, Cho CF, Bai ST, et al. ADAM9 promotes lung cancer progression through vascular remodeling by VEGFA, ANGPT2, and PLAT. Sci Rep. 2017;7:15108.
30. Sher YP, Wang LJ, Chuang LL, et al. ADAM9 up-regulates N-cadherin via miR-218 suppression in lung adenocarcinoma cells. PLoS ONE. 2014;9:e94065.
31. Lin YW, Wen YC, Lin CY, et al. Genetic variants of ADAM9 as potential predictors for biochemical recurrence in prostate cancer patients after receiving a radical prostatectomy. Int J Med Sci. 2024;21:2934-42.
32. Gao J, Aksoy BA, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
33. Cerami E, Gao J, Dogrusoz U, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401-4.
34. Pan C, Chen H, Yang B. Licochalcone a inhibits proliferation and metastasis of colon cancer by regulating miR-1270/ADAM9/Akt/NF-κB axis. Iran J Public Health. 2023;52:1962-72.
35. AmeliMojarad M, AmeliMojarad M, Wang J, Tavakolpour V, Shariati P. A pan-cancer study of ADAM9’s immunological function and prognostic value particularly in liver cancer. Sci Rep. 2024;14:26862.
36. Li J, Ji Z, Qiao C, Qi Y, Shi W. Overexpression of ADAM9 promotes colon cancer cells invasion. J Invest Surg. 2013;26:127-33.
37. Forbes SA, Beare D, Boutselakis H, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017;45:D777-83.
38. Drobiova H, Al-Mulla F, Al-Temaimi R. ADAM9 genetic variants and their role in modulating enzyme activity in diabetes and metabolic traits. J Diabetes Res. 2025;2025:5519447.
39. Ilieva M, Tschaikowski M, Vandin A, Uchida S. The current status of gene expression profilings in COVID-19 patients. Clin Transl Discov. 2022;2:e104.
40. Ibor OY, Ekpenyong B, Charles M. A, Kenneth E. M. Review on gene and cell therapies in prostate cancer treatment: prospects and challenges. J Transl Genet Genom. 2025;9:62-75.
41. Chen CM, Hsieh YH, Hwang JM, et al. Fisetin suppresses ADAM9 expression and inhibits invasion of glioma cancer cells through increased phosphorylation of ERK1/2. Tumour Biol. 2015;36:3407-15.
42. Wang JJ, Zou JX, Wang H, et al. Histone methyltransferase NSD2 mediates the survival and invasion of triple-negative breast cancer cells via stimulating ADAM9-EGFR-AKT signaling. Acta Pharmacol Sin. 2019;40:1067-75.
43. Liu R, Gu J, Jiang P, et al. DNMT1-microRNA126 epigenetic circuit contributes to esophageal squamous cell carcinoma growth via ADAM9-EGFR-AKT signaling. Clin Cancer Res. 2015;21:854-63.
44. Liu CM, Hsieh CL, He YC, et al. In vivo targeting of ADAM9 gene expression using lentivirus-delivered shRNA suppresses prostate cancer growth by regulating REG4 dependent cell cycle progression. PLoS ONE. 2013;8:e53795.
45. Giebeler N, Schönefuß A, Landsberg J, Tüting T, Mauch C, Zigrino P. Deletion of ADAM-9 in HGF/CDK4 mice impairs melanoma development and metastasis. Oncogene. 2017;36:5058-67.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Special Topic
Copyright
Data & Comments
Data

















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].