Artificial intelligence and multiomics for longevity across species
Abstract
Understanding the biological basis of extreme longevity and healthspan remains a fundamental challenge in aging research. Conventional model organisms fall short in explaining the exceptional cancer resistance and multi-decade lifespans of species such as bowhead whales and naked mole-rats, revealing critical gaps in current paradigms. We argue that the field now stands at a pivotal juncture, propelled by two converging advances: the systematic study of non-traditional, long-lived species and the emergence of high-resolution multi-omics technologies. To harness this synergy, we introduce Longevity Intelligence - an integrative framework that unites cross-species evolutionary principles, multi-layered omics data, and artificial intelligence (AI) to decode conserved mechanisms of extreme longevity. While traditional research has often relied on isolated biological models, this perspective proposes a conceptual closed-loop system where AI generates testable hypotheses from cross-species “big data”, which must subsequently be validated via rigorous wet-lab experimentation. By transcending taxonomic and molecular boundaries, Longevity Intelligence offers a powerful lens to uncover universal rules of aging and accelerate the translation of evolutionary insights into interventions that extend human healthspan.
Keywords: Longevity Intelligence, multiple model organisms, multi-omics, artificial intelligence, comparative biology, healthspan
EXPANDING THE HORIZON: A REPOSITORY OF MODEL ORGANISMS AS DATA GENERATORS
Natural selection has, over millions of years, curated a diverse portfolio of species with exceptional lifespans and pronounced resistance to age-related diseases. Moving beyond traditional models to incorporate these “non-model” organisms into our research arsenal is no longer optional but essential for a holistic understanding of longevity. Rather than viewing these species in isolated, descriptive contexts, the Longevity Intelligence framework treats them as fundamental data generators - naturally occurring experiments in delayed aging that provide the critical input data required for advanced computational modeling.
The bowhead whale: As the longest-lived mammal on Earth, with lifespans reliably estimated to exceed 200 years, the bowhead whale stands as a paramount example of negligible senescence in nature. This species exhibits a remarkable, near-complete resistance to cancer and other common age-related pathologies. Genomic comparative studies have pinpointed specific genes under significant positive selection in the bowhead whale, which are core components of DNA repair machinery, cell cycle regulation, and tumor suppression pathways - notably Excision Repair Cross-Complementing Group 1 (ERCC1) and Proliferating Cell Nuclear Antigen (PCNA)[1]. A pivotal recent study provides direct functional validation of these genomic insights[2], demonstrating that primary cells derived from bowhead whales, leveraging mechanisms such as the non-homologous end joining (NHEJ) pathway, achieve superior efficiency and fidelity in repairing DNA double-strand breaks. Thus, this species offers an unparalleled model for deciphering the synergistic mechanisms that maintain genomic integrity, suppress tumorigenesis, and preserve tissue function over a super-long lifespan. Given that the core DNA repair pathways are evolutionarily conserved in humans, their targeted activation represents a promising anti-aging strategy[3].
The naked mole-rat: Standing as a profound challenge to fundamental tenets of biogerontology, the naked mole-rat has a maximum lifespan exceeding 30 years. It displays exceptional resistance to a wide spectrum of age-associated pathologies, including cancer, neurodegeneration, and cardiovascular decline. The seminal discovery of high-molecular-weight hyaluronic acid (HMW-HA) in its extracellular matrix unveiled a primary mechanism for its exceptional cancer resistance via early contact inhibition[4]. Moving beyond structural mechanisms, a novel cyclic GMP-AMP synthase (cGAS)-mediated pathway has recently been identified as central to its enhanced DNA repair and delayed aging[5]. Notably, the cGAS-STING (Stimulator of Interferon Gene) pathway serves as a central sensor of genomic instability and a key driver of inflammation in humans; its dysregulation underpins age-related chronic inflammation (inflammaging) and degenerative diseases. The finely tuned cGAS activity of the naked mole-rat thus offers a compelling blueprint for therapeutic modulation of this pathway.
African elephant: Peto’s Paradox - the lack of correlation between body size (and thus cell number) and cancer incidence - is elegantly demonstrated in the African elephant. Despite having roughly 100 times the number of cells as a human, their cancer mortality rate is remarkably low. This is facilitated by the evolution of multiple copies of the tumor suppressor gene TP53 (p53), which orchestrates a robust apoptotic response to DNA damage[6]. Additionally, genomic and functional studies have identified a refunctionalized pseudogene leukemia inhibitory factor 6 (LIF6) as a key downstream effector of p53: LIF6 is transcriptionally activated by p53 upon DNA damage, translocates to mitochondria, and induces Bax/Bak-dependent apoptosis to eliminate damaged cells, thereby reinforcing cancer surveillance[7]. The p53-LIF6 axis, therefore, exemplifies a precise evolutionary adaptation that resolves Peto’s Paradox. Studying the elephant offers key insights into how fundamental cellular safeguards can be amplified and rewired to meet the unique demands of extreme longevity and body size. Consequently, the p53 pathway is frequently disrupted in human cancers as the most commonly mutated tumor suppressor network, representing a compelling therapeutic target. Strategies to enhance its activity or emulate the elephant’s evolutionarily refined, multi-copy p53 repertoire offer a promising frontier in both cancer prevention and aging biology[8].
Non-human primates: While distant models offer unique evolutionary insights, non-human primates (NHPs), such as the rhesus macaque and the cynomolgus monkey, provide an indispensable translational bridge to human aging. Sharing highly similar physiology, life history, and age-related pathologies, NHPs are the premier model for evaluating geroprotective interventions. Contemporary research leverages multi-omics to decode the molecular architecture of primate aging. A landmark 40-month study in male cynomolgus monkeys exemplifies this: the FDA-approved drug metformin systemically decelerates multi-dimensional biological aging across 79 tissues[9]. Integrating pan-tissue transcriptomics, DNA methylomics, plasma proteomics, and single-nucleus RNA sequencing, the work not only confirms a healthspan promoting effect but maps it at high resolution - delineating reversible cell-type-specific aging trajectories, pinpointing the nuclear factor erythroid 2–re-lated factor 2 (Nrf2)-mediated antioxidant pathway as a central mechanism, and quantifying a striking ~6-year reversal in brain epigenetic age with preserved cognitive function. This finding supports ongoing Phase III clinical trials of metformin for age-related cognitive decline. Empowered by deep multi-omics profiling, NHPs thus offer the most direct, mechanistic reference for understanding human organ-specific aging and accelerating the translation of geroscience discoveries into clinical applications.
Ectothermic centenarians: To ensure a comprehensive and diverse dataset for artificial intelligence (AI) training, this organismal repository must be systematically expanded to include long-lived ectotherms. Species such as certain testudines (turtles) and deep-sea fish offer unique evolutionary insights into metabolic rate regulation, hypoxia tolerance, and negligible senescence under extreme environmental pressures[10,11], preventing the framework from being biased solely toward mammalian physiology.
MULTI-OMICS AT THE FOREFRONT OF PRECISION GEROSCIENCE
The unique biological questions posed by these model organisms demand a sophisticated and multi-faceted analytical toolkit. Multi-omics technologies provide this, enabling a transition from descriptive observation to precise, mechanistic understanding.
Overcoming technical roadblocks in cross-species integration: Because cross-species data integration involves numerous technical challenges, rigorous data harmonization is paramount. Identifying conserved pathways requires sophisticated computational corrections for batch effects, genome annotation discrepancies across non-model species, and phylogenetic distance weighting to prevent evolutionary divergence from masking true functional similarities.
Functional genomic maps: A high-quality reference genome would provide a foundational map for all subsequent discoveries. The genomes of long-lived species are treasure troves of evolutionary innovation. However, traditional short-read sequencing technologies fail to accurately resolve complex repetitive regions, structural variations, and GC-rich areas, which are often hotspots for evolutionary change. The advent of long-read sequencing technologies, including BGI’s CycloneSEQ platforms and third-generation platforms such as PacBio’s HiFi and Oxford Nanopore ultralong-read sequencing (ONT), has revolutionized this field. These technologies enable the construction of truly complete, telomere-to-telomere (T2T) reference genomes[12]. This technological leap has directly addressed critical questions in comparative genomics of long-lived species. For instance, resolving the precise architecture of the p53 locus in the African elephant (Loxodonta africana) required long-read sequencing to definitively map and characterize its expanded repertoire of p53 retrogenes (including the functionally critical p53/LIF6), which underpin its exceptional cancer resistance and offer a solution to Peto’s Paradox[7]. This complete genomic picture is indispensable for functional studies.
Deconvoluting cellular heterogeneity: single-cell and spatial multi-omics: Aging is not a uniform process across all cells within an organism; it is driven by shifts in cellular composition and state-specific dysfunction. A core scientific problem is identifying which specific cell types and states are primary drivers of tissue aging and resilience. Single-cell RNA sequencing (scRNA-seq) and single-cell Assay for Transposase-Accessible Chromatin sequencing (scATAC-seq) directly address this by deconstructing tissues to identify rare cell types (e.g., stem and progenitor cells) and their state transitions during aging at unprecedented resolution. For instance, applying scRNA-seq to aged mammalian tissues has specifically resolved the problem of identifying the diverse and functionally distinct senescent cell subsets that accumulate with age, moving beyond bulk-tissue “senescence-associated” signatures to pinpoint precise cellular targets for senolytic therapies[13]. However, these techniques dissociate cells from their native spatial context, leaving the critical question of how cellular neighborhoods and communication networks break down during aging unanswered. Spatial omics technologies, such as the high-resolution BGI’s Stereo-seq platform, complement single-cell approaches by restoring the critical dimension of tissue architecture. These methods are uniquely positioned to dissect the functional organization of aging tissues and uncover the breakdown of intercellular communication networks. A recent spatial transcriptomic study exemplifies this power: it revealed that senescent cells in aged tissues form distinct spatial clusters that specifically upregulate immunoglobulin genes - a previously unrecognized “immunoglobin-associated senescence” phenotype that actively reshapes the local immune microenvironment[14]. Consequently, spatial omics elevates our understanding from how individual cells change to how cellular collectives drive the reorganization and functional decline of tissues during aging.
Capturing dynamic processes: Longevity is a dynamic, lifelong process. Cross-sectional studies provide snapshots, but longitudinal sampling is key to distinguishing cause from effect. Collecting tissues from the same animal or cohort at multiple time points - youth, middle age, and old age - and subjecting them to integrated genomic, epigenomic, transcriptomic, proteomic, and metabolomic analyses enables the construction of a “molecular movie” of the life course. This approach can identify early-warning biomarkers of aging, trace the origin of age-related functional decline, and uncover the temporal sequence of molecular events leading to pathology. For instance, longitudinal epigenomic studies in humans and model organisms have established the clock-like nature of DNA methylation changes, leading to the development of highly accurate “epigenetic clocks” that can serve as biomarkers of biological age[15].
INTEGRATION AND EMPOWERMENT: AN AI-DRIVEN PARADIGM
The scale and complexity of this “big data” challenge surpass the capabilities of conventional bioinformatics tools, creating a compelling opportunity for AI to drive discovery. However, it is highly unlikely that AI can unravel the biological complexity of aging without robust, standardized inputs. Recognizing that the primary hurdle for AI in geroscience is the small pool of available datasets, we propose a formalized, data-driven framework. A schematic overview of this integrated Longevity Intelligence framework is presented in Figure 1, which delineates the full closed-loop pipeline from evolutionary insights of longevity model organisms to experimental validation. Here, we elaborate on an “AI-Empowered Closed Loop Framework for Longevity Research”.
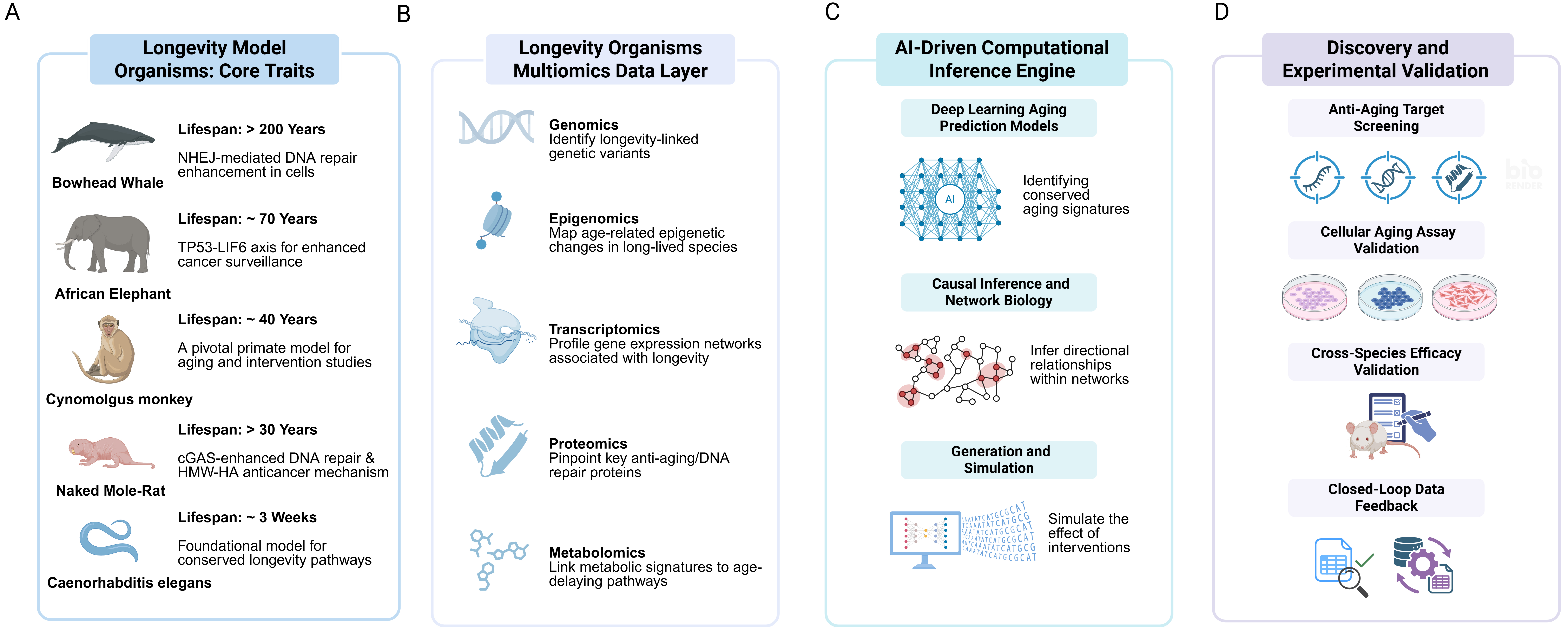
Figure 1. AI-empowered closed loop framework for longevity research.
This schematic outlines the integrated pipeline of Longevity Intelligence, a unified discovery framework that leverages evolutionary insights from non-traditional longevity model organisms and multi-omics data to uncover fundamental mechanisms of longevity. (A) Longevity Model Organisms: Core Traits - species with exceptional lifespan and resistance to age-related diseases (e.g., bowhead whale, naked mole-rat, certain testudines and deep-sea fish) serve as biological anchors for comparative analysis; (B) Longevity Organisms Multiomics Data Layer - high-resolution genomic, transcriptomic, epigenomic, proteomic, and metabolomic profiles are collected across species and life stages to build comprehensive datasets; (C) AI-Driven Computational Inference Engine - deep learning models, causal inference networks, and simulation tools integrate multi-omics data to predict anti-aging pathways, identify conserved molecular signatures, and generate testable hypotheses; (D) Discovery and Experimental Validation - predicted targets undergo high-throughput screening, cellular aging assays, cross-species functional validation, and in vivo testing, with closed-loop feedback refining computational models. This iterative cycle enables the translation of evolutionary insights into actionable strategies for extending healthspan. [This figure was created in BioRender. Shi, Z. (2026) https://BioRender.com/nuwpcd3.] AI: Artificial intelligence.
Data layer: the longevity multidimensional databank. The first step is the creation of a centralized, standardized, and FAIR (Findable, Accessible, Interoperable, Reusable)[16] databank. This would integrate genomic, transcriptomic, epigenomic, proteomic, metabolomic, and imaging data from all the aforementioned model organisms, across their entire lifespans, and in response to various interventions[17]. Massive, scalable computational infrastructure is needed here.
Algorithm and model layer: the AI engine. This layer converts raw multi-omics data into actionable biological insights through a three-stage analytical pipeline
Predictive models: Supervised deep learning models can be trained on multi-omics data to predict biological age with higher accuracy than any single biomarker. These “holistic aging clocks” can be species-specific or, more powerfully, cross-species, identifying conserved aging signatures[18]. Beyond clocks, AI models are directly identifying lifespan-extending compounds by addressing the limitations of traditional drug discovery. For instance, the AI model ElixirSeeker identified potential lifespan-extending compounds, and 4 out of the tested candidates were validated to extend lifespan in Caenorhabditis elegans[19]. Using interpretability techniques, these predictions can be anchored to specific pathways, offering testable biological hypotheses for validation.
Causal inference and network biology: Correlation does not imply causation. Advanced AI methods, such as graph neural networks and causal machine learning, can be applied to infer directional relationships within molecular interaction networks. This helps identify “master regulator” genes and pathways that are upstream drivers of aging, rather than downstream consequences, thereby pinpointing optimal intervention targets[20]. The resulting causal graphs provide an interpretable and testable mechanistic framework for predictions.
Generation and biological foundation models: Generative AI models, including generative adversarial networks (GANs) and diffusion models, can be trained on the molecular states of young, healthy tissues. They can then be used to simulate the effects of genetic or pharmacological interventions, predicting how they might shift an aged molecular profile towards a more youthful one. This allows for the in silico screening and prioritization of thousands of potential anti-aging strategies, focusing experimental resources on the most promising candidates. For example, Munson et al.[21] demonstrated that generative AI models can design novel drug candidates by concurrently modulating functionally linked protein targets - such as mammalian target of rapamycin, a key node in aging-related signaling pathways - highlighting their potential for application in Geroscience-driven drug discovery. Furthermore, emerging Biological Foundation Models, pre-trained on massive datasets of multi-species multi-omics, offer the unprecedented capability to capture the fundamental “syntax” of biology, enabling zero-shot predictions of cross-species aging trajectories and regulatory mechanisms[22].
Discovery and validation layer: closing the loop. A computational framework remains purely conceptual without empirical proof. Therefore, predictions and targets generated by the AI models must be rigorously validated in vitro and in vivo. The first step: In Vitro Validation. Where feasible, primary cells or established cell lines derived from long-lived species serve as the first testbed. Functional assays are used to validate AI-predicted mechanisms within the relevant cellular context. The second step: In Vivo Validation. The most critical step involves testing the conserved role of AI-identified targets - such as the differential regulatory mechanics of the elephant p53-LIF6 axis or the whale ERCC1/PCNA clusters - in tractable laboratory models. Through targeted genetic manipulation (e.g., CRISPR-Cas9 engineering) in short-lived mammals, we can definitively assess the causal geroprotective efficacy of these pathways. The wet-lab results feed back into the databank, creating a virtuous cycle that refines the AI models.
PAST PROGRESS AND FUTURE PERSPECTIVES
Historically, the field of aging research relied heavily on isolated, single-model organisms to identify basic, conserved longevity pathways. Today, propelled by the convergence of comparative biology and advanced computational science, the field stands at an inflection point. The path forward requires moving from descriptive multi-omics toward predictive and generative biology. Emerging opportunities are poised to redefine longevity research:
(1) Digital twins and virtual interventions: The ultimate ambition is to construct high-fidelity, dynamic “digital twins” of biological systems - from single cells to whole organisms. These in silico models would enable non-invasive, large-scale testing of personalized anti-aging interventions, dramatically accelerating discovery while reducing reliance on animal experimentation.
(2) Cross-species principles of longevity: By leveraging AI to cut through taxonomic noise, this integrative framework aims to uncover deeply conserved molecular networks, regulatory logic, and pathway architectures that govern longevity across the tree of life.
(3) Next-generation of biomarkers and therapeutic targets: Systematic AI-driven mining of multi-species, multi-omics datasets will yield robust biomarkers of biological age and reveal a pipeline of evolutionarily validated, high-specificity targets for gerotherapeutic development.
The path ahead holds transformative potential, yet significant hurdles remain - particularly in data standardization and cross-platform, cross-species interoperability; the substantial cost of generating and storing multi-omics datasets; the urgent need for biologically interpretable (explainable) AI models; and the ethical implications of large-scale animal studies and data-intensive research.
To address these barriers, we issue “a call to action: establish a Global Longevity Data Commons (GLDC)”. We propose an international, cross-disciplinary consortium - uniting biogerontologists, computational biologists, AI scientists, and ethics and policy experts - with a unified mission: (1) to develop and deploy standardized protocols for generating, annotating, and sharing multi-omics data across diverse longevity model organisms and human aging cohorts; (2) to build a federated, cloud-based platform that adheres to FAIR principles while safeguarding data privacy, equity, and national sovereignty; and (3) to create a shared infrastructure for training, benchmarking, and validating next-generation AI models aimed at uncovering evolutionarily conserved mechanisms of aging. By dismantling data silos and establishing a common foundation for collaborative discovery, the GLDC would dramatically accelerate progress, amplify the return on global research investments, and provide the coordinated infrastructure essential to realizing the transformative vision outlined above.
CONCLUSIONS
Longevity research is entering a pivotal phase driven by the deepening convergence of comparative biology, advanced biotechnology, and computational science. We possess a diverse array of naturally long-lived model organisms and a sophisticated multi-omics toolbox capable of multi-dimensional measurement. To unlock these resources, AI will serve as an essential analytical engine. By deeply integrating multiple model organisms, multi-level omics data, and AI, we can systematically decode the biological foundations of longevity and open new pathways for extending human healthspan. This perspective serves as a call to action to collectively advance the field toward deeper and more impactful frontiers.
DECLARATIONS
Acknowledgments
We thank the global research community for foundational contributions to longevity research, multi-omics technologies, and AI-driven discovery. Gratitude extends to teams generating and sharing datasets of long-lived species, and developers of key sequencing and computational tools that enabled this work. We apologize for not citing all impactful studies due to space limitations.
Authors’ contributions
Writing - original draft: Nie, C.; Shi, Z.
Writing - review and editing: Wang, F.; Min, J.; Song, Z.
Conceptualization: Wang, F.; Min, J.
Nie, C. and Shi, Z. contributed equally to the article.
Availability of data and materials
Not applicable.
AI and AI-assisted tools statement
During the preparation of this manuscript, the AI tool DeepSeek (version 3.2, released 2025-12-1) was used solely for language editing. The tool did not influence the study design, data collection, analysis, interpretation, or the scientific content of the work. All authors take full responsibility for the accuracy, integrity, and final content of the manuscript.
Financial support and sponsorship
This work was made possible through the generous support of the National Key R&D Program of China (2023YFC3603300), the Ministry of Science and Technology of China (2022ZD0211600), and the Shenzhen Science and Technology Program (SYSPG20241211173845012).
Conflicts of interest
Wang, F. is the Editor-in-Chief of the journal Element, and Min, J. is a Deputy Editor of the journal Element. Wang, F. and Min, J. were not involved in any steps of the editorial process, including reviewer selection, manuscript handling, or decision-making. The other authors declare that there are no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Keane, M. et al. Insights into the evolution of longevity from the bowhead whale genome. Cell Rep. 10, 112-22 (2015).
2. Firsanov, D. et al. Evidence for improved DNA repair in the long-lived bowhead whale. Nature 648, 717-25 (2025).
3. López-Otín, C.; Blasco, M. A.; Partridge, L.; Serrano, M.; Kroemer, G. Hallmarks of aging: an expanding universe. Cell 186, 243-78 (2023).
4. Tian, X. et al. High-molecular-mass hyaluronan mediates the cancer resistance of the naked mole rat. Nature 499, 346-9 (2013).
5. Chen, Y. et al. A cGAS-mediated mechanism in naked mole-rats potentiates DNA repair and delays aging. Science 390, eadp5056 (2025).
6. Abegglen, L. M. et al. Potential mechanisms for cancer resistance in elephants and comparative cellular response to DNA damage in humans. JAMA 314, 1850-60 (2015).
7. Vazquez, J. M.; Sulak, M.; Chigurupati, S.; Lynch, V. J. A zombie LIF gene in elephants is upregulated by TP53 to induce apoptosis in response to DNA damage. Cell Rep. 24, 1765-76 (2018).
8. Levine, A. J. p53: 800 million years of evolution and 40 years of discovery. Nat. Rev. Cancer 20, 471-80 (2020).
10. Glen, C. G. et al. Aging dynamics in captive sea turtles reflect conserved life-history patterns across the testudine phylogeny. Commun. Biol. 8, 1297 (2025).
11. Muradian, K. K.; Fraifeld, V. E. Metazoans who live longer than jeanne calment. In: Rejuvenation and Longevity; Healthy Ageing and Longevity, Vol. 20; Springer, Cham, 2024.
13. Tabula Muris Consortium. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590-5 (2020).
14. Ma, S. et al. Spatial transcriptomic landscape unveils immunoglobin-associated senescence as a hallmark of aging. Cell 187, 7025-44.e34 (2024).
15. Horvath, S.; Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 19, 371-84 (2018).
16. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
18. Lu, A. T. et al. Universal DNA methylation age across mammalian tissues. Nat. Aging 3, 1144-66 (2023).
19. Pan, Y. et al. ElixirSeeker: a machine learning framework utilizing fusion molecular fingerprints for the discovery of lifespan-extending compounds. Aging Cell 24, e70116 (2025).
20. Li, H. et al. Inferring transcription factor regulatory networks from single-cell ATAC-seq data based on graph neural networks. Nat. Mach. Intell. 4, 389-400 (2022).
21. Munson, B. P. et al. De novo generation of multi-target compounds using deep generative chemistry. Nat. Commun. 15, 3636 (2024).
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
Article Notes
About This Article
Copyright




Data & Comments
Data












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].