Learning biological dynamics beyond pairwise networks
Abstract
Systems biology has traditionally relied on network abstractions and mechanistic models to study complex biological systems. However, advances in model expressiveness and data availability have not translated into proportional improvements in mechanistic understanding. We suggest that a key limitation may arise from a structural mismatch: prevailing pairwise interaction models fail to capture the inherently higher-order organization of biological systems. Across molecular, cellular, and ecological scales, system behavior is governed by cooperative, conditional, and context-dependent multi-body interactions that cannot be faithfully represented by pairwise projections alone. To address this challenge, we advocate a paradigm shift toward explicit higher-order structural representations combined with data-driven, learnable dynamical models. Within this framework, artificial intelligence enables the inference of governing dynamical rules and the discovery of mechanisms operating on higher-order structures, while large language models can accelerate hypothesis generation and the integration of prior knowledge. Together, these advances point toward a unified, generative approach to systems biology that moves beyond descriptive networks toward an interpretable, mechanism-driven understanding of the processes underlying biological function.
Keywords
INTRODUCTION
Over the past two decades, systems biology has established a powerful quantitative framework for studying biological complexity[1,2]. By integrating network representations, dynamical models, and multi-omics data, the field has enabled system-level analyses across molecular, cellular, and ecological scales[3]. Gene regulatory networks (GRNs), protein–protein interaction networks, metabolic networks, and ecological interaction networks have become canonical abstractions for organizing biological knowledge and guiding hypothesis-driven research[4]. These advances represent a major conceptual achievement, despite historical constraints in data resolution and computational capacity.
However, a growing gap has emerged between predictive performance and mechanistic understanding. Modern systems biology increasingly relies on high-dimensional data and expressive machine learning models; however, improvements in predictive accuracy have not proportionally translated into deeper biological insight, interpretability, or generalization[5,6]. For example, models that predict cell fate decisions from single-cell transcriptomics data or microbial community dynamics from metagenomics data can achieve high precision, but often do not reveal underlying causal mechanisms[7]. This gap highlights a structural limitation in current modeling frameworks rather than a purely algorithmic problem[8].
A key but often overlooked assumption underlying most systems biology approaches is that biological systems can be adequately represented as networks of pairwise interactions[9,10]. This assumption is embedded in both mechanistic models and data-driven approaches, ranging from ordinary differential equation models and stochastic reaction networks to graph-based machine learning methods[11]. Although pairwise networks are mathematically convenient and historically useful, they inherently constrain the types of mechanisms that models can capture[11,12]. In practice, however, biological systems exhibit rich higher-order interactions that involve three or more components simultaneously[13]. For instance, transcriptional regulation can depend on combinations of cofactors and chromatin states; metabolic fluxes often involve multi-enzyme complexes; microbial communities are shaped by context-dependent cooperation and competition; and ecological networks exhibit triadic and higher-order interactions that govern system stability[14]. Beyond representation, collapsing multi-component mechanisms into dyadic edges can yield wrong mechanistic conclusions even with high predictive fit. We highlight failures motivating higher-order models.
Combinatorial gene regulation. Many promoters are governed by logical AND/OR control: transcription occurs only when a specific set of TFs/cofactors co-occur (often under chromatin-modulated regulation). Pairwise GRNs encode this as multiple strong edges, falsely implying independent additive effects and hiding the minimal regulatory logic; a higher-order hyperedge captures the required combination as a single mechanistic unit, distinguishing conditional dependencies from marginal associations.
Metabolic flux control. Metabolic throughput may be governed by enzyme complexes or coupled reaction sets whose functional activity emerges only at the multi-enzyme level. Pairwise graphs can fit correlations yet misattribute control, leading to misleading causal interpretations of perturbation experiments such as knockouts or inhibition; higher-order interaction models allow dynamical systems to attribute influence to multi-enzyme units rather than spurious dyads.
Microbiome and ecological stability. Ecological outcomes often depend on triadic facilitation/inhibition (e.g., a third species mediates competition or supplies a shared resource). Pairwise reductions can misestimate stability margins and tipping points because effective interactions vary with context; higher-order interaction terms naturally capture this dependence and enable stability analyses conditioned on multi-species configurations.
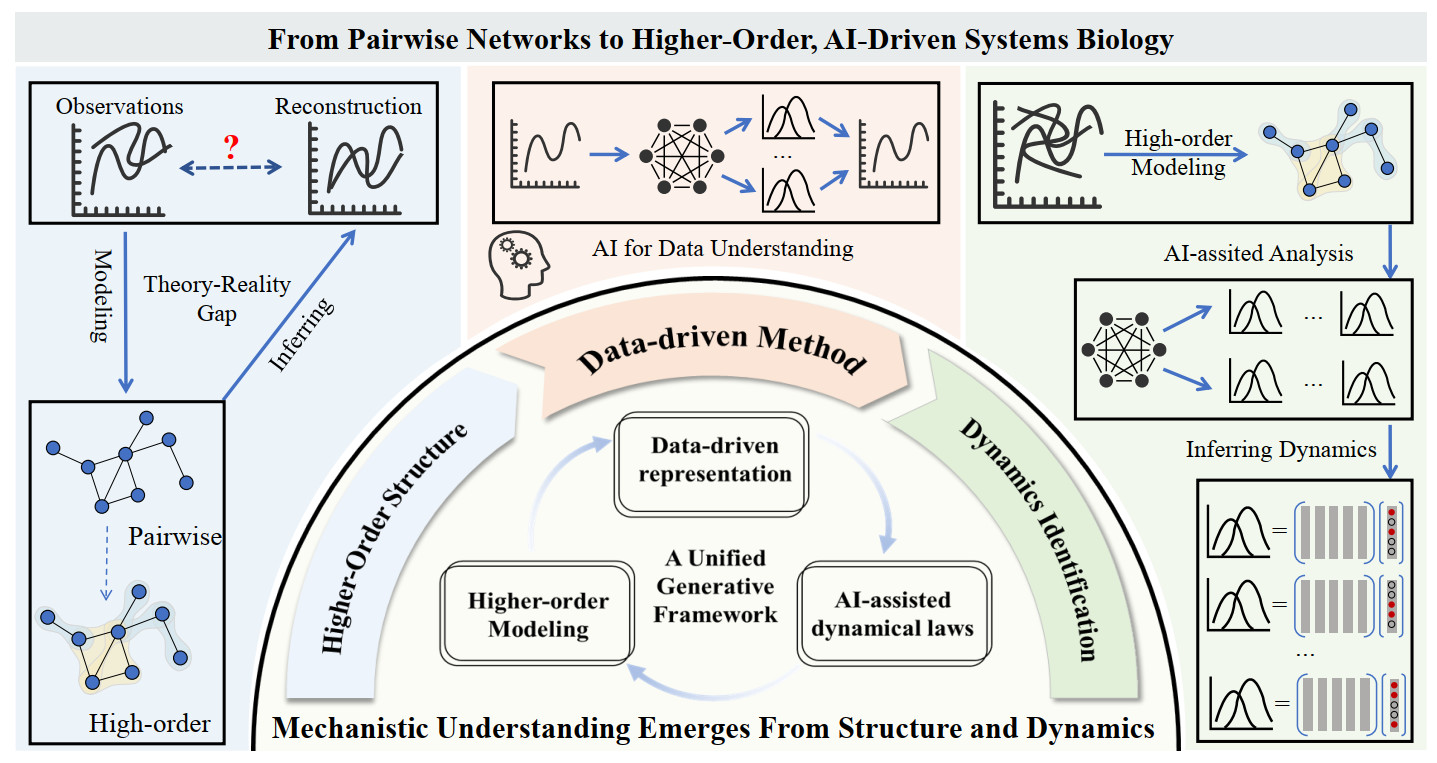
As a result, we suggest that some of the persistent challenges in systems biology may reflect structural limitations, rather than purely computational ones. Increasing the complexity of the model on pairwise networks cannot recover the lost higher-order structure[10]. As shown in Figure 1, the field is poised to transition toward explicit higher-order network representations, where nodes and hyperedges capture multi-component interactions, coupled with artificial intelligence (AI)-driven analysis tools capable of learning the dynamical rules governing these structures[15]. Specifically, the proposed framework consists of four parts: (1) Mismatch between observation and reconstruction: Time-resolved observations and their reconstructions may disagree. A key reason is that standard pairwise models can miss latent higher-order structures (e.g., multi-component constraints, combinatorial regulation, complexes, or context-dependent triads). Moving from dyadic graphs to hyperedges/simplices helps narrow this observation–reconstruction gap. (2) Data-driven methods implicitly capture dynamics: Modern AI and representation learning pipelines provide data-driven representations that can implicitly encode underlying dynamical processes from high-dimensional, noisy measurements, enabling prediction and interpretation even when explicit equations are unknown. (3) AI-assisted analysis enables explicit inference: By combining higher-order modeling with AI-assisted dynamical analysis, it becomes possible to move from implicit representations to explicit dynamical inference: learning governing laws/operators, attributing effects to specific edges or hyperedges, and quantifying uncertainty, ultimately yielding interpretable dynamics across conditions. (4) A closed-loop progression, The workflow advances through three coupled modules—structure discovery (pairwise to higher-order), data-driven methods (AI-based representation and generative modeling), and dynamics identification (system identification and attribution). These modules mutually reinforce one another, forming a feedback loop that progressively improves reconstruction and mechanistic understanding.
Figure 1. From pairwise networks to higher-order, AI-driven systems biology. Left : mismatch between observation and reconstruction. Center: unified data-driven framework. Right: higher-order modeling + AI-assisted analysis enables explicit inference. Outer arcs: a closed-loop progression
In practice, such an approach could enable, for instance, interpretable models of context-specific transcriptional regulation, multi-enzyme metabolic flux prediction, or ecosystem stability analysis under environmental perturbations. By integrating higher-order representations with data-driven and system identification methodologies, systems biology can go beyond prediction to achieve mechanistic understanding, causal insight, and principled discovery in complex biological systems.
HIGHER-ORDER STRUCTURE AS A FUNDAMENTAL FEATURE OF BIOLOGICAL SYSTEMS
Across biological scales, from molecular regulation to ecosystem dynamics, interactions among system components are frequently cooperative, conditional, and context-dependent, involving more than two entities simultaneously[9,16,17]. Transcriptional regulation often requires the joint presence of multiple transcription factors and cofactors; metabolic flux depends on multi-enzyme complexes and coupled reaction sets; cellular decision making integrates concurrent signaling pathways; and microbial and ecological communities display interaction patterns determined by community composition rather than isolated species pairs[18]. These multi-body dependencies are pervasive: system behavior generally cannot be decomposed into independent pairwise effects without loss of essential information[19]. This universality indicates that higher-order interactions are a fundamental organizing principle of living systems, rather than isolated exceptions[20]. For a broader systems-level view of how network-based data, models, and applications are used in computational biology, we refer readers to the comprehensive survey in[21].
Projecting inherently higher-order mechanisms onto pairwise network representations incurs structural costs[12]. Cooperative or conditional interactions are misrepresented as strong binary links, conflating multi-body causality with pairwise association. Such projections can distort stability properties, obscure critical transitions, and generate misleading dynamical predictions, particularly in nonlinear regimes dominated by collective effects[22]. In data-driven settings, machine learning models trained on pairwise networks may partially recover statistical patterns, but often at the expense of interpretability and mechanistic fidelity. Consequently, limitations commonly attributed to model complexity or data noise may instead originate from an overly restrictive structural hypothesis space[23].
Importantly, higher-order representations are not merely more detailed versions of traditional networks, but constitute a qualitatively different modeling language[24]. Structures such as hypergraphs and simplicial complexes encode multi-body constraints as first-class entities, allowing them to directly participate in system dynamics. This approach transforms the central challenge of systems biology: from approximating complex behavior on simplified structures to learning and discovering dynamical laws operating on native higher-order networks[9]. Moreover, higher-order dynamics has been investigated beyond biology in several canonical settings, including nonlinear consensus processes on temporal hypergraphs with noisy higher-order interactions[25], explicit three-body interaction models that reveal novel consensus and conservation laws[26], and connectivity/percolation phenomena in random hypergraphs (including variants with anchored nodes)[27]. These results highlight that qualitatively new dynamical regimes can emerge when interactions are irreducibly multi-body, motivating higher-order representations as more than a bookkeeping refinement.
Recent progress includes reconstructing higher-order structure from data, discovering higher-order motifs/ modules, and identifying nonlinear governing equations on networks/complexes. Rather than proposing a new estimator, we frame a unifying workflow where higher-order structure serves as a mechanistic hypothesis space, dynamical identification assigns interpretable roles to multi-body interactions, and AI accelerates iteration among prior knowledge, model selection, and experiment design—complementing existing reconstruction and system identification methods. By explicitly modeling higher-order dependencies, this framework integrates AI-based inference, interpretability, and mechanistic discovery, grounding next-generation systems biology[28].
LEARNING DYNAMICS AND DISCOVERING MECHANISMS ON HIGHER-ORDER BIOLOGICAL STRUCTURES
Given the pervasive higher-order dependencies in biological systems, traditional equation-centric modeling approaches face fundamental challenges. Classical frameworks rely on explicitly specified interaction terms, which become combinatorially intractable as the number of interacting components increases. As a result, many higher-order systems are either approximated with oversimplified dynamics or remain without mechanistic models, highlighting the limitations of pairwise and low-order modeling paradigms[29].
Data-driven dynamical modeling offers an alternative by treating governing laws as objects to be learned directly from observations. Rather than prescribing functional forms, these approaches infer state evolution rules from time-resolved measurements, capturing nonlinearities and multi-body dependencies that are difficult to anticipate a priori. Crucially, the goal is not limited to short-term prediction, but extends to the identification of dynamical principles that are consistent across conditions, thereby enabling mechanistic insight from the learned rules themselves[30].
Let
where
where
Higher-order structures such as hypergraphs and simplicial complexes provide a natural substrate for such learning. By explicitly representing multi-body interactions, they constrain the hypothesis space of possible dynamics, reducing the combinatorial burden and promoting sparsity, stability, and interpretability[32]. Within this framework, learning algorithms can identify which higher-order interactions are functionally relevant and how their collective influence shapes system behavior. Higher-order representations thus provide both accuracy and inductive bias, facilitating reliable inference of complex dynamical patterns[33].
Integrating higher-order structure with learnable dynamics opens new avenues for mechanistic discovery. Mechanisms emerge as identifiable dynamical patterns linked to specific multi-body interactions, enabling the detection of context-dependent interaction regimes, regime shifts, and drivers of emergent behavior. Large language models (LLMs) can further assist this process by generating structured hypotheses, synthesizing prior biological knowledge, and constraining the search space for data-driven inference[34,35]. Combined with empirical data and higher-order learning, such hybrid workflows offer a scalable path toward systematic, interpretable, and mechanism-driven modeling of complex biological systems[36].
DISCUSSION
The developments described above illustrate a broader conceptual shift in systems biology: from descriptive pairwise network analyses toward a framework that integrates higher-order structures, learnable dynamics, and AI-driven inference to explain how system-level behaviors arise. Although traditional network metrics such as connectivity, modularity, and centrality provide valuable insights, they are insufficient to capture the dynamic and context-dependent nature of biological function. Biological outcomes emerge from the interplay between structural constraints and the governing dynamical laws, highlighting the need for a generative and mechanism-focused perspective[37].
Within this emerging paradigm, three interdependent components form a coherent framework. Higher-order structural representations encode biologically grounded multi-body constraints; learnable dynamical laws extract governing rules directly from time-resolved data; and AI tools, including LLMs, facilitate the exploration of complex hypothesis spaces and the integration of prior knowledge. Together, these components establish a closed loop: structure constrains dynamics, dynamics reveal mechanisms, and AI accelerates the cycle of discovery and hypothesis testing.
This framework has concrete implications for the practice of systems biology. Experimental design may prioritize temporally resolved, perturbation-based, or combinatorial data over static measurements. Modeling strategies may shift from fitting predefined equations to learning dynamical rules under explicit higher-order constraints. Model evaluation may emphasize mechanistic consistency, identification of dynamical regimes, and generalization across conditions, rather than predictive accuracy alone. Collectively, these changes align computational modeling more closely with the causal processes underlying biological phenomena.
Learning higher-order dynamics brings key technical hurdles for scientific reliability. Data/observability: effects are context-activated and hidden in cross-section data; therefore, time series, perturbation–response, combinatorial interventions, and latent-state models should be employed. Identifiability/overfitting: hyperedges grow combinatorially; thus, sparsity, cap order, mechanistic priors, and cross-condition generalization tests should be applied. Validation: prediction
LLMs are most useful when treated as hypothesis generators and translators between unstructured knowledge and structured model components, rather than as sources of evidence. Concretely, LLMs can (i) extract candidate multi-component relations from literature and databases to propose a structured prior over plausible hyperedges/motifs, (ii) map learned higher-order dynamical terms into human-interpretable mechanistic narratives and testable predictions, and (iii) assist in designing perturbation panels that are maximally diagnostic for distinguishing among competing interaction hypotheses. However, because LLM outputs may be incomplete or biased, safeguards are essential: candidate outputs should be citation-grounded where possible, cross-checked for consistency across sources, and always treated as provisional hypotheses to be filtered through data-driven identification, uncertainty quantification, and experimental validation.
Even so, by explicitly embracing higher-order organization and data-driven discovery within a unified, AI-augmented framework, systems biology is positioned to move beyond descriptive modeling toward a mechanistic, interpretable, and generative understanding of complex living systems.
DECLARATIONS
Authors' contributions
Conceived and designed the study, and wrote the manuscript: Shao, Q.
Contributed to the discussion and provided significant feedback on the manuscript: Zhang, X.; Liu, X.
Supervised the research, provided overall guidance, and revised the manuscript: Dong, G.; Chen, D.
Availability of data and materials
Not applicable.
AI and AI-assisted tools Statement
Artificial Intelligence (AI) (specifically ChatGPT version 5.2) was used for language polishing and for generating elements for schematic figures. All ideas and conclusions are those of the authors; AI tools did not generate any scientific claims or original research content. The authors reviewed and edited all AI-generated content and accept full responsibility for the published material.
Financial support and sponsorship
This work was supported by the National Key Research and Development Program of China (Grant No. 2022ZD0120004), the Zhishan Youth Scholar Program, the National Natural Science Foundation of China (Grant Nos. 62233004, 62273090, and 62073076), and the Jiangsu Provincial Scientific Research Center of Applied Mathematics (Grant BK20233002).
Conflicts of interest
Chen, D. is an Associate Editor of the journal Complex Engineering Systems. Chen, D. was not involved in any stage of the editorial process, notably including reviewer selection, manuscript handling, or decision making, while the other authors have declared that they have no conflicts of interest.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Copyright
© The Author(s) 2026.
REFERENCES
1. Yan, J.; Risacher, S. L.; Shen, L.; Saykin, A. J. Network approaches to systems biology analysis of complex disease: integrative methods for multi-omics data. Brief. Bioinform. 2018, 19, 1370-81.
2. Baião, A. R.; Cai, Z.; Poulos, R. C.; et al. A technical review of multi-omics data integration methods: from classical statistical to deep generative approaches. Brief. Bioinform. 2025, 26, bbaf355.
3. Li, J.; Liao, Q.; Zhou, H.; et al. Multi-omics analyses reveal regulatory networks underpinning metabolite biosynthesis in Nicotiana tabacum. Nat. Commun. 2025, 16, 10339.
4. Wright, S. N.; Colton, S.; Schaffer, L. V.; et al. State of the interactomes: an evaluation of molecular networks for generating biological insights. Mol. Syst. Biol. 2025, 21, 1-29.
5. Köhler, D.; Rügamer, D.; Boyle, L. J.; Maloney, K. O.; Schmid, M. Achieving interpretable machine learning by functional decomposition of black-box models into explainable predictor effects. npj. Artif. Intell. 2025, 1, 34.
6. Murdoch, W. J.; Singh, C.; Kumbier, K.; Abbasi-Asl, R.; Yu, B. Interpretable machine learning: definitions, methods, and applications. arXiv 2019, arXiv: 190104592.
7. Lagemann, K.; Lagemann, C.; Taschler, B.; Mukherjee, S. Deep learning of causal structures in high dimensions under data limitations. Nat. Mach. Intell. 2023, 5, 1306-16.
8. Rudin, C.; Chen, C.; Chen, Z.; Huang, H.; Semenova, L.; Zhong, C. Interpretable machine learning: fundamental principles and 10 grand challenges. Statist. Surv. 2022, 16.
9. Bick, C.; Gross, E.; Harrington, H. A.; Schaub, M. T. What are higher-order networks? SIAM. Rev. 2023, 65, 686-731.
10. Majhi, S.; Perc, M.; Ghosh, D. Dynamics on higher-order networks: a review. J. R. Soc. Interface. 2022, 19, 20220043.
11. Zhang, Y.; Lucas, M.; Battiston, F. Higher-order interactions shape collective dynamics differently in hypergraphs and simplicial complexes. Nat. Commun. 2023, 14, 1605.
12. Bian, J.; Zhou, T.; Bi, Y. Unveiling the role of higher-order interactions via stepwise reduction. Commun. Phys. 2025, 8, 228.
13. Terry, J. C. D.; Bonsall, M. B.; Morris, R. J. The impact of structured higher-order interactions on ecological network stability. Theor. Ecol. 2025, 18, 9.
14. Lucas, M.; Gallo, L.; Ghavasieh, A.; Battiston, F.; De Domenico, M. Reducibility of higher-order networks via information flow. arXiv 2024, arXiv: 240408547.
15. Kim, S.; Lee, S. Y.; Gao, Y.; Antelmi, A.; Polato, M.; Shin, K. A survey on hypergraph neural networks: an in-depth and step-by-step guide. In Proceedings of the 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining; 2024. pp. 6534-44.
16. Boccaletti, S.; De Lellis, P.; Del Genio, C.; et al. The structure and dynamics of networks with higher order interactions. Phys. Rep. 2023, 1018, 1-64.
17. Lambiotte, R.; Rosvall, M.; Scholtes, I. From networks to optimal higher-order models of complex systems. Nat. Phys. 2019, 15, 313-20.
18. Coyte, K. Z.; Schluter, J.; Foster, K. R. The ecology of the microbiome: networks, competition, and stability. Science 2015, 350, 663-6.
19. Grilli, J.; Barabás, G.; Michalska-Smith, M. J.; Allesina, S. Higher-order interactions stabilize dynamics in competitive network models. Nature 2017, 548, 210-3.
20. Battiston, F.; Amico, E.; Barrat, A.; et al. The physics of higher-order interactions in complex systems. Nat. Phys. 2021, 17, 1093-8.
21. Liu, C.; Ma, Y.; Zhao, J.; et al. Computational network biology: data, models, and applications. Phys. Rep. 2020, 846, 1-66.
22. Wang, Y.; Li, A.; Wang, L. Networked dynamic systems with higher-order interactions: stability versus complexity. Natl. Sci. Rev. 2024, 11, nwae103.
23. Wang, Y.; Kleinberg, J. From graphs to hypergraphs: hypergraph projection and its remediation. arXiv 2024, arXiv: 240108519.
24. Tian, H.; Zafarani, R. Higher-order networks representation and learning: a survey. ACM. SIGKDD. Explor. Newsl. 2024, 26, 1-18.
25. Shang, Y. Non-linear consensus dynamics on temporal hypergraphs with random noisy higher-order interactions. J. Complex. Netw. 2023, 11, cnad009.
26. Shang, Y. A system model of three-body interactions in complex networks: consensus and conservation. Proc. R. Soc. A. 2022, 478, 20210564.
28. Skardal, P.S.; Arenas, A. Higher order interactions in complex networks of phase oscillators promote abrupt synchronization switching. Commun. Phys. 2020, 3, 218.
29. Champion, K.; Lusch, B.; Kutz, J. N.; Brunton, S. L. Data-driven discovery of coordinates and governing equations. Proc. Natl. Acad. Sci. USA. 2019, 116, 22445-51.
30. Brunton, S. L.; Proctor, J. L.; Kutz, J. N. Discovering governing equations from data by sparse identification of nonlinear dynamical systems. Proc. Natl. Acad. Sci. USA. 2016, 113, 3932-7.
31. Battiston, F.; Bick, C.; Lucas, M.; Millán, A. P.; Skardal, P. S.; Zhang, Y. Collective dynamics on higher-order networks. Nat. Rev. Phys. 2026, 8, 146-59.
32. Delabays, R.; De Pasquale, G.; Dorfler, F.; Zhang, Y. Hypergraph reconstruction from dynamics. Nat. Commun. 2025, 16, 2691.
33. Antelmi, A.; Cordasco, G.; Polato, M.; Scarano, V.; Spagnuolo, C.; Yang, D. A survey on hypergraph representation learning. ACM. Comput. Surv. 2023, 56, 1-38.
34. Zhang, Y.; Khan, S. A.; Mahmud, A.; Yang, H.; Lavin, A.; et al. Exploring the role of large language models in the scientific method: from hypothesis to discovery. NPJ. Artif. Intell. 2025, 1, 14.
35. Zheng, T.; Deng, Z.; Tsang, H. T.; et al. From automation to autonomy: a survey on large language models in scientific discovery. In Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing; 2025. pp. 17744-61.
36. Alkan, A. K.; Sourav, S.; Jablonska, M, et al. A survey on hypothesis generation for scientific discovery in the era of large language models. arXiv 2025, arXiv: 250405496.
Cite This Article
How to Cite
Download Citation
Export Citation File:
Type of Import
Tips on Downloading Citation
Citation Manager File Format
Type of Import
Direct Import: When the Direct Import option is selected (the default state), a dialogue box will give you the option to Save or Open the downloaded citation data. Choosing Open will either launch your citation manager or give you a choice of applications with which to use the metadata. The Save option saves the file locally for later use.
Indirect Import: When the Indirect Import option is selected, the metadata is displayed and may be copied and pasted as needed.
About This Article
Copyright
Data & Comments
Data
















Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at [email protected].